24 results
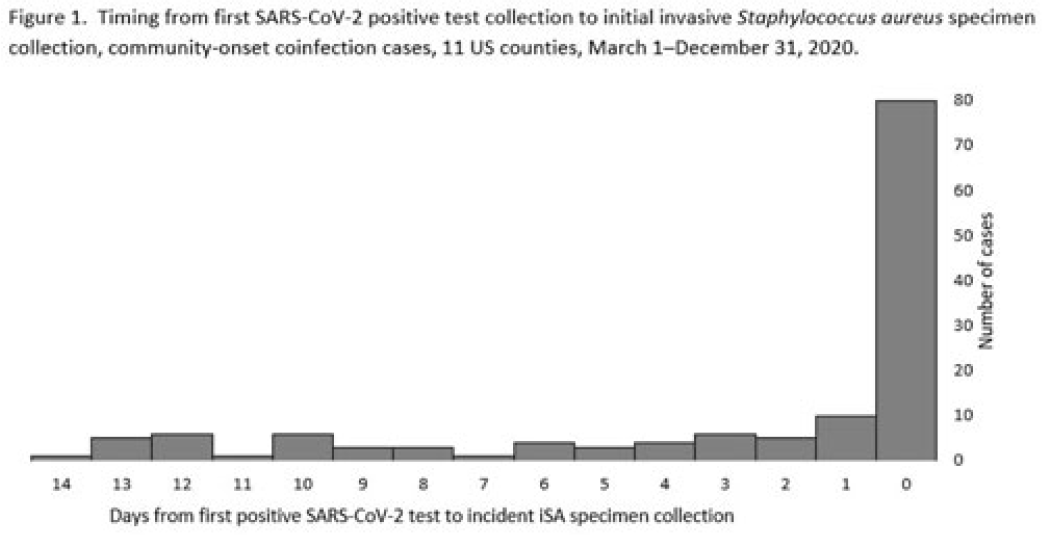
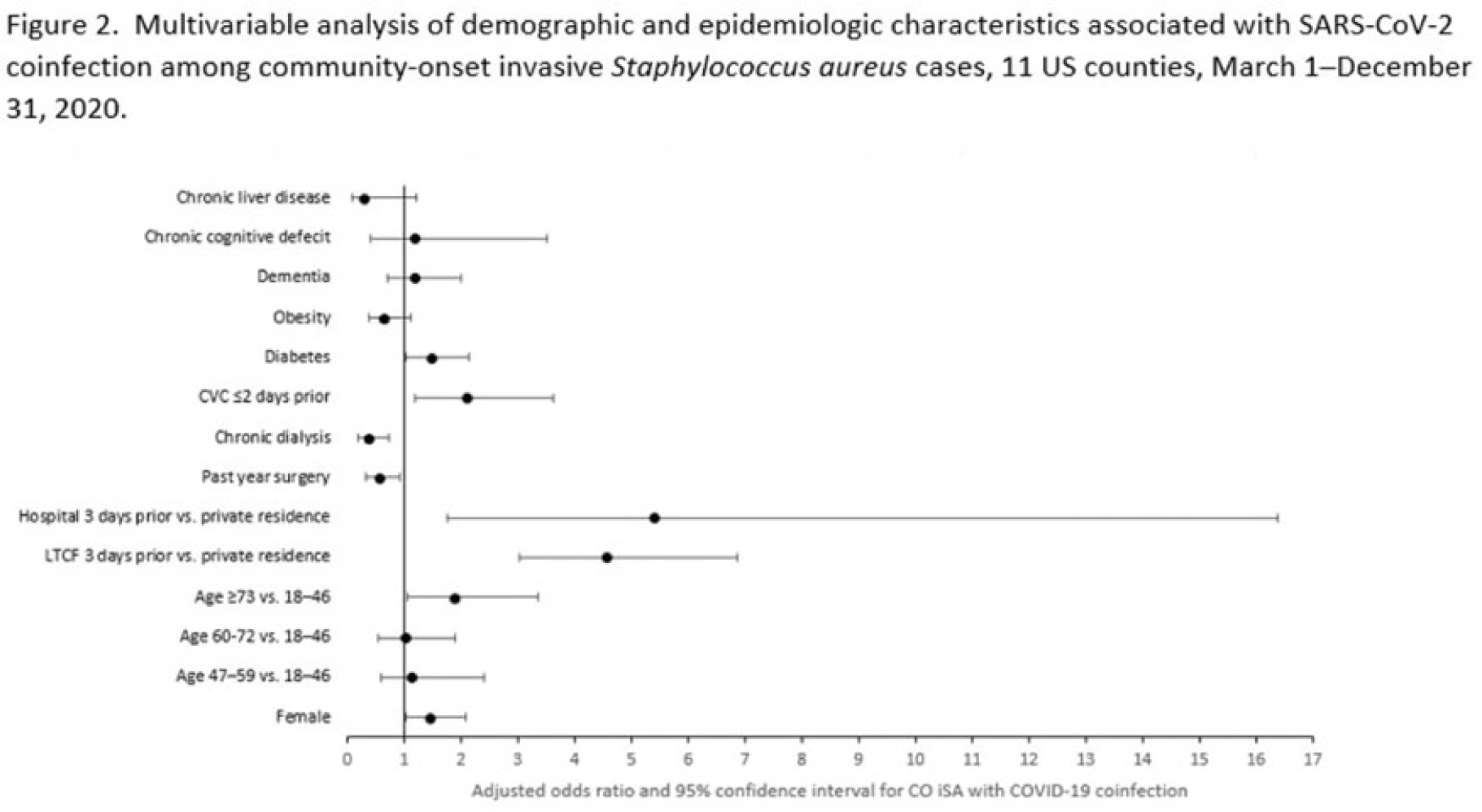
Factors associated with SARS-CoV-2 and community-onset invasive Staphylococcus aureus coinfection, 2020
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 3 / Issue S2 / June 2023
- Published online by Cambridge University Press:
- 29 September 2023, pp. s84-s85
-
- Article
-
- You have access
- Open access
- Export citation
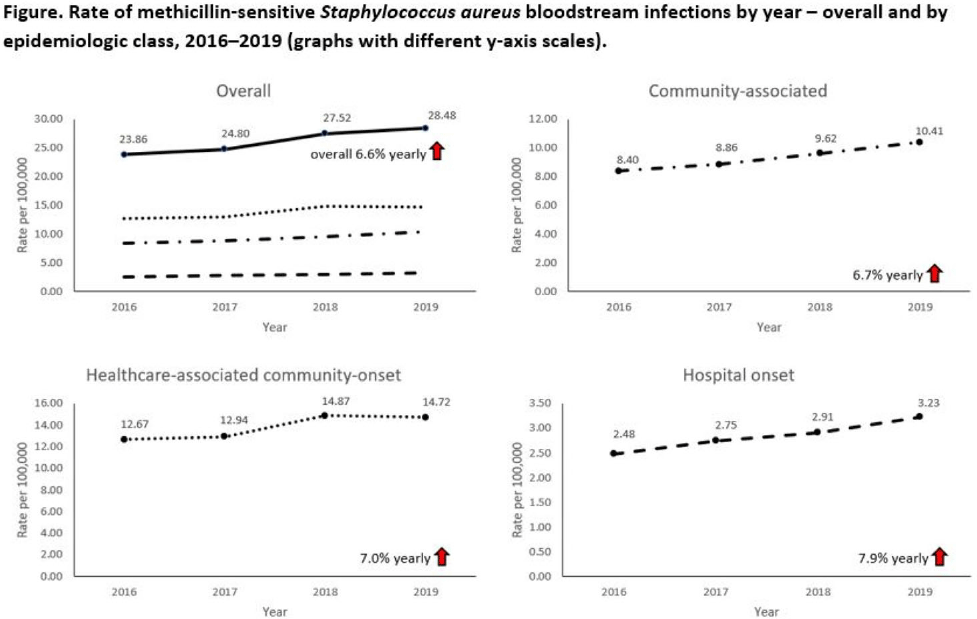
Increases in methicillin-sensitive Staphylococcus aureus bloodstream infection incidence, 2016–2019
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 2 / Issue S1 / July 2022
- Published online by Cambridge University Press:
- 16 May 2022, pp. s63-s64
-
- Article
-
- You have access
- Open access
- Export citation
Epidemiology of extended-spectrum β-lactamase–producing Enterobacterales in five US sites participating in the Emerging Infections Program, 2017
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 43 / Issue 11 / November 2022
- Published online by Cambridge University Press:
- 14 February 2022, pp. 1586-1594
- Print publication:
- November 2022
-
- Article
- Export citation
Evaluation of Discrepancies in Carbapenem Minimum Inhibitory Concentrations Obtained at Clinical Laboratories Compared to a Public Health Laboratory
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s474-s476
- Print publication:
- October 2020
-
- Article
-
- You have access
- Export citation
Characterization of Ceftazidime-Avibactam-Resistant Carbapenem-Resistant Enterobacteriaceae, United States, 2015–2017
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s465-s466
- Print publication:
- October 2020
-
- Article
-
- You have access
- Export citation
Substance Use Diagnoses Among Persons with Community-Onset Methicillin-Resistant Staphylococcus aureus Bloodstream Infections
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s392-s393
- Print publication:
- October 2020
-
- Article
-
- You have access
- Export citation
Molecular Typing of Invasive Staphylococcus aureus from the Emerging Infections Program (EIP) Using Whole-Genome Sequencing
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s71-s72
- Print publication:
- October 2020
-
- Article
-
- You have access
- Export citation
Characteristics Associated With Invasive Staphylococcus aureus Infection Rates in Nursing Homes, Emerging Infections Program
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s60-s61
- Print publication:
- October 2020
-
- Article
-
- You have access
- Export citation
Validation of Administrative Codes for Identification of Staphylococcus aureus Infections Among Electronic Health Data
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s507-s509
- Print publication:
- October 2020
-
- Article
-
- You have access
- Export citation
Epidemiologic Characteristics of ESBL-Producing ST131 E. coli Identified Through the Emerging Infections Program, 2017
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s214-s215
- Print publication:
- October 2020
-
- Article
-
- You have access
- Export citation
Antimicrobial Nonsusceptibility Among Invasive MRSA USA300 Strains by Healthcare Exposure, Three Sites, 2005–2016
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s120-s121
- Print publication:
- October 2020
-
- Article
-
- You have access
- Export citation
Area-Based Socioeconomic Status Measures and Incidence of Community-Associated ESBL-Producing Enterobacteriaceae, 2017
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s128-s129
- Print publication:
- October 2020
-
- Article
-
- You have access
- Export citation
P-495 - Gender Difference in Antidepressant-related Sexual Dysfunction in Patients With Major Depressive Disorder (mdd)
-
- Journal:
- European Psychiatry / Volume 27 / Issue S1 / 2012
- Published online by Cambridge University Press:
- 15 April 2020, p. 1
-
- Article
-
- You have access
- Export citation
Trends in methicillin-resistant Staphylococcus aureus bloodstream infections using statewide population-based surveillance and hospital discharge data, Connecticut, 2010–2018
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue 6 / June 2020
- Published online by Cambridge University Press:
- 13 April 2020, pp. 734-736
- Print publication:
- June 2020
-
- Article
- Export citation
Antimicrobial-resistant pathogens associated with adult healthcare-associated infections: Summary of data reported to the National Healthcare Safety Network, 2015–2017
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue 1 / January 2020
- Published online by Cambridge University Press:
- 26 November 2019, pp. 1-18
- Print publication:
- January 2020
-
- Article
-
- You have access
- HTML
- Export citation
Antimicrobial-resistant pathogens associated with pediatric healthcare-associated infections: Summary of data reported to the National Healthcare Safety Network, 2015–2017
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue 1 / January 2020
- Published online by Cambridge University Press:
- 25 November 2019, pp. 19-30
- Print publication:
- January 2020
-
- Article
-
- You have access
- HTML
- Export citation
Pathogen Distribution and Antimicrobial Resistance Among Pediatric Healthcare-Associated Infections Reported to the National Healthcare Safety Network, 2011–2014
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 39 / Issue 1 / January 2018
- Published online by Cambridge University Press:
- 18 December 2017, pp. 1-11
- Print publication:
- January 2018
-
- Article
- Export citation
Clinical Correlates of Surveillance Events Detected by National Healthcare Safety Network Pneumonia and Lower Respiratory Infection Definitions—Pennsylvania, 2011–2012
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 37 / Issue 7 / July 2016
- Published online by Cambridge University Press:
- 13 April 2016, pp. 818-824
- Print publication:
- July 2016
-
- Article
- Export citation
Completeness of Methicillin-Resistant Staphylococcus aureus Bloodstream Infection Reporting From Outpatient Hemodialysis Facilities to the National Healthcare Safety Network, 2013
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 37 / Issue 2 / February 2016
- Published online by Cambridge University Press:
- 11 November 2015, pp. 205-207
- Print publication:
- February 2016
-
- Article
- Export citation
Mucosal Barrier Injury Laboratory-Confirmed Bloodstream Infections (MBI-LCBI): Descriptive Analysis of Data Reported to National Healthcare Safety Network (NHSN), 2013
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 37 / Issue 1 / January 2016
- Published online by Cambridge University Press:
- 12 October 2015, pp. 2-7
- Print publication:
- January 2016
-
- Article
- Export citation