Research Article
Heterospecific transcription of the Escherichia coli rpoB-3 allele in Gram-negative bacteria
-
- Published online by Cambridge University Press:
- 14 April 2009, pp. 87-90
-
- Article
-
- You have access
- Export citation
An assessment of the homoeology of six Agropyron intermedium chromosomes added to wheat
-
- Published online by Cambridge University Press:
- 14 April 2009, pp. 91-97
-
- Article
-
- You have access
- Export citation
Temporal stability of P–M cytotype polymorphism in a natural population of Drosophila melanogaster
-
- Published online by Cambridge University Press:
- 14 April 2009, pp. 99-103
-
- Article
-
- You have access
- Export citation
Hybrid dysgenesis in natural populations of Drosophila melanogaster in Japan. II. Strains which cannot induce P-M dysgenesis may completely suppress functional P element activity
-
- Published online by Cambridge University Press:
- 14 April 2009, pp. 105-111
-
- Article
-
- You have access
- Export citation
Genetic and developmental analysis of the sex-determining gene ‘double sex’ (dsx) of Drosophila melanogaster
-
- Published online by Cambridge University Press:
- 14 April 2009, pp. 113-123
-
- Article
-
- You have access
- Export citation
Correlation of compound autosome segregation and sex-chromosome disjunction in female Lucilia cuprina (Wied.) (Diptera: Calliphoridae)
-
- Published online by Cambridge University Press:
- 14 April 2009, pp. 125-130
-
- Article
-
- You have access
- Export citation
Autosomal assignment of OTC in marsupials and monotremes: implications for the evolution of sex chromosomes
-
- Published online by Cambridge University Press:
- 14 April 2009, pp. 131-136
-
- Article
-
- You have access
- Export citation
Nucleotide sequence analysis of class II genes borne by mouse t chromosomes
-
- Published online by Cambridge University Press:
- 14 April 2009, pp. 137-146
-
- Article
-
- You have access
- Export citation
Genetic effects on susceptibility to histidine induced teratogenesis in the mouse
-
- Published online by Cambridge University Press:
- 14 April 2009, pp. 147-153
-
- Article
-
- You have access
- Export citation
Estimating the proportion of neutral mutants
-
- Published online by Cambridge University Press:
- 14 April 2009, pp. 155-163
-
- Article
-
- You have access
- Export citation
Erratum
Erratum
-
- Published online by Cambridge University Press:
- 14 April 2009, p. 165
-
- Article
-
- You have access
- Export citation
Erratum
-
- Published online by Cambridge University Press:
- 14 April 2009, p. 166
-
- Article
-
- You have access
- Export citation
Book Reviews
Human Genetics: Problems and Approaches. Second edition. By F. Vogel and A. G. Motulsky. Berlin, Heidelberg, New York, Tokyo: Springer-Verlag1986, 807 pages. Hard cover DM 148, £52.25. ISBN 3 540 16411 1.
-
- Published online by Cambridge University Press:
- 14 April 2009, p. 167
-
- Article
-
- You have access
- Export citation
Natural History of the Major Histocompatability Complex. By Jan Klein. New York, Chichester: John Wiley & Sons. 1986. 775 pages. £90.75. ISBN 0 471 80953 5.
-
- Published online by Cambridge University Press:
- 14 April 2009, pp. 167-168
-
- Article
-
- You have access
- Export citation
Electron Microscopy in Molecular Biology: A Practical Approach. Edited by J. Sommerville and U. Scheer. Oxford: IRL Press Ltd.1987. 248 pages. £16.00, US $29.00. ISBN 0 947946 54 3.
-
- Published online by Cambridge University Press:
- 14 April 2009, pp. 168-169
-
- Article
-
- You have access
- Export citation
Build Your Own DNA Kit. By William G. Thilly and Alexander Varshavsky. London: Butterworths. £17.25. ISBN 0 409 90094 X.
-
- Published online by Cambridge University Press:
- 14 April 2009, pp. 169-170
-
- Article
-
- You have access
- Export citation
Books Received
Books Received
-
- Published online by Cambridge University Press:
- 14 April 2009, p. 171
-
- Article
-
- You have access
- Export citation
Front matter
GRH volume 50 issue 2 Cover and Front matter
-
- Published online by Cambridge University Press:
- 14 April 2009, pp. f1-f2
-
- Article
-
- You have access
- Export citation
Back matter
GRH volume 50 issue 2 Cover and Back matter
-
- Published online by Cambridge University Press:
- 14 April 2009, pp. b1-b3
-
- Article
-
- You have access
- Export citation



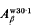 borne by the
borne by the 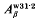 and
and 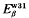 borne by the
borne by the 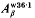 and
and 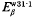 borne by the
borne by the 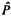 , of the proportion of neutral alleles is positively biased if (roughly) 4
, of the proportion of neutral alleles is positively biased if (roughly) 4