52 results
Large-scale circulation reversals explained by pendulum correspondence
-
- Journal:
- Journal of Fluid Mechanics / Volume 993 / 25 August 2024
- Published online by Cambridge University Press:
- 13 September 2024, A3
-
- Article
-
- You have access
- Open access
- HTML
- Export citation
Predicting convection configurations in coupled fluid–porous systems
-
- Journal:
- Journal of Fluid Mechanics / Volume 953 / 25 December 2022
- Published online by Cambridge University Press:
- 09 December 2022, A23
-
- Article
-
- You have access
- Open access
- HTML
- Export citation
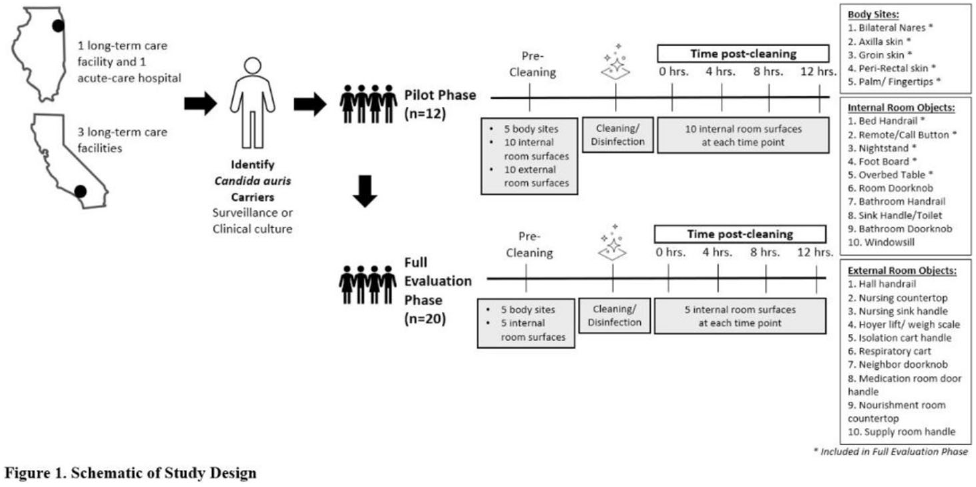
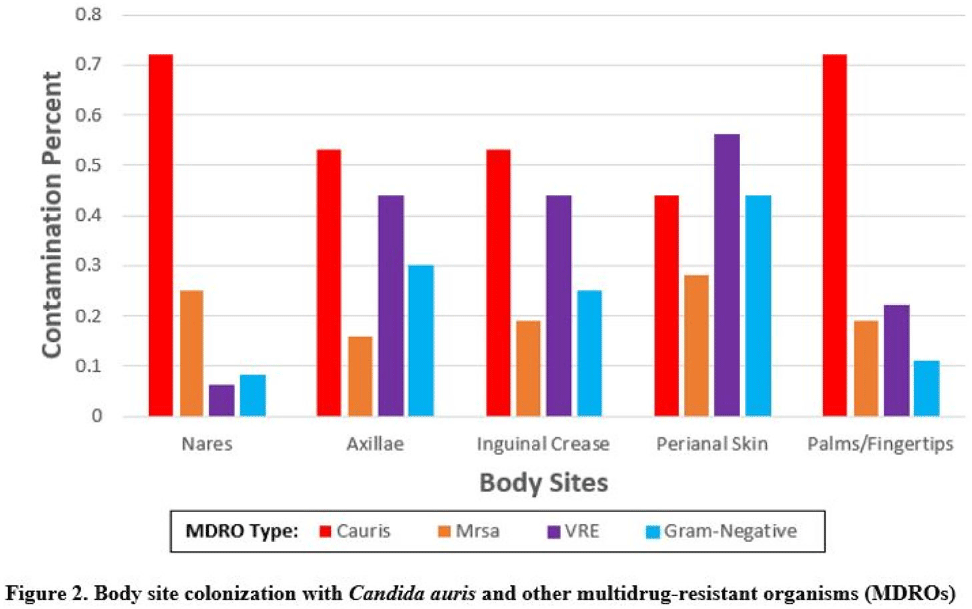
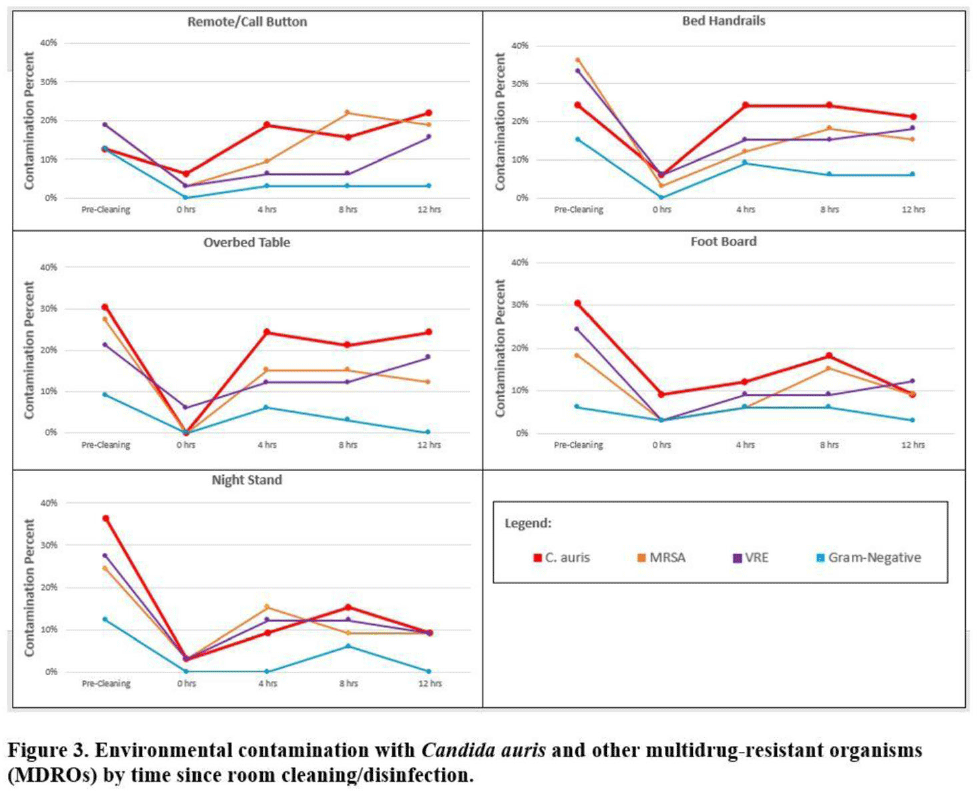
Multicenter evaluation of contamination of the healthcare environment near patients with Candida auris skin colonization – ERRATUM
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 2 / Issue 1 / 2022
- Published online by Cambridge University Press:
- 07 October 2022, e166
-
- Article
-
- You have access
- Open access
- HTML
- Export citation
Multicenter evaluation of contamination of the healthcare environment near patients with Candida auris skin colonization
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 2 / Issue S1 / July 2022
- Published online by Cambridge University Press:
- 16 May 2022, pp. s78-s79
-
- Article
-
- You have access
- Open access
- Export citation
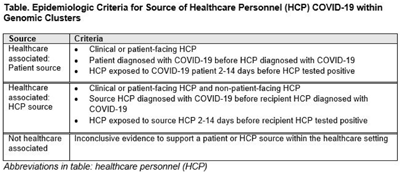
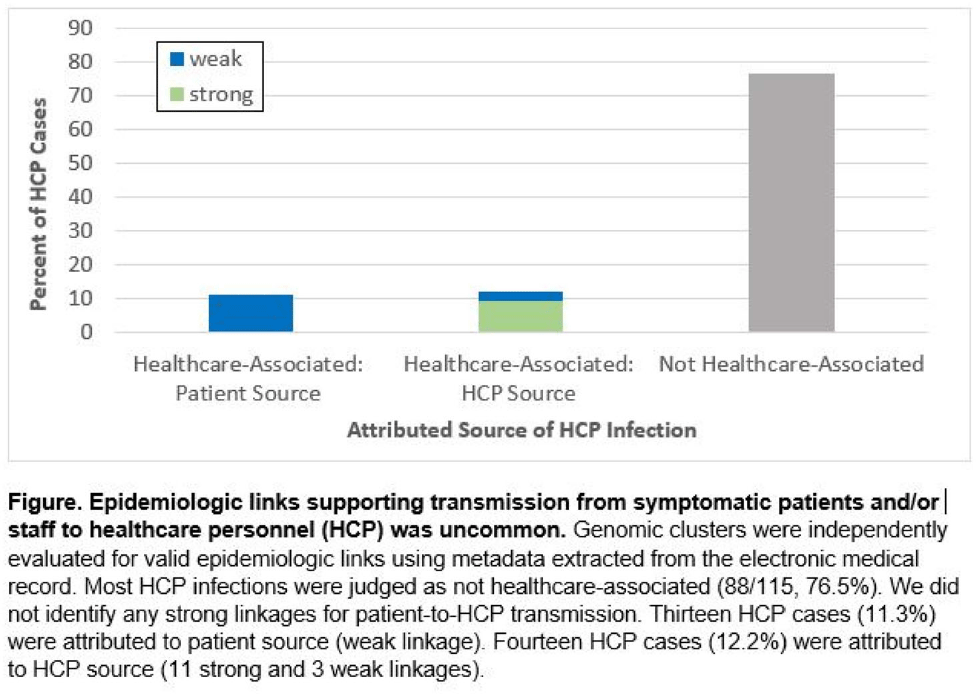
Genomic investigation to identify the source of SARS-CoV-2 infection among healthcare personnel
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 2 / Issue S1 / July 2022
- Published online by Cambridge University Press:
- 16 May 2022, pp. s74-s75
-
- Article
-
- You have access
- Open access
- Export citation
‘He Saw Heaven Opened’: Heavenly Temple and Universal Mission in Luke-Acts
-
- Journal:
- New Testament Studies / Volume 68 / Issue 1 / January 2022
- Published online by Cambridge University Press:
- 09 December 2021, pp. 38-51
- Print publication:
- January 2022
-
- Article
-
- You have access
- Open access
- HTML
- Export citation
Whole-genome sequencing for neonatal intensive care unit outbreak investigations: Insights and lessons learned – ADDENDUM
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 1 / Issue 1 / 2021
- Published online by Cambridge University Press:
- 03 August 2021, e18
-
- Article
-
- You have access
- Open access
- HTML
- Export citation
Next Generation File Formats and Platforms
-
- Journal:
- Microscopy and Microanalysis / Volume 27 / Issue S1 / August 2021
- Published online by Cambridge University Press:
- 30 July 2021, p. 2842
- Print publication:
- August 2021
-
- Article
-
- You have access
- Export citation
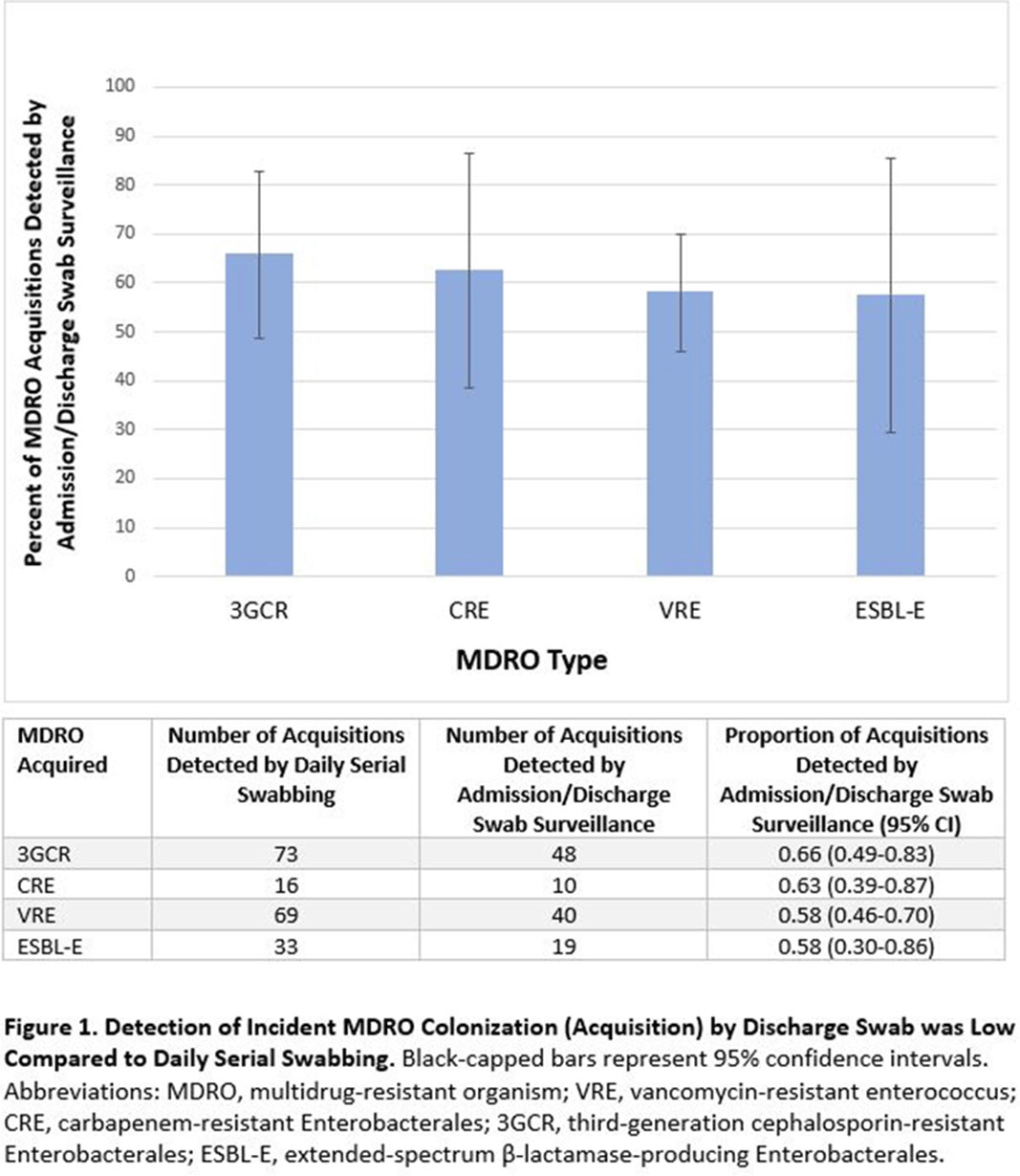
Admission and Discharge Sampling Underestimates Multidrug-Resistant Organism (MDRO) Acquisition in an Intensive Care Unit
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 1 / Issue S1 / July 2021
- Published online by Cambridge University Press:
- 29 July 2021, p. s28
-
- Article
-
- You have access
- Open access
- Export citation
Whole-genome sequencing for neonatal intensive care unit outbreak investigations: Insights and lessons learned
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 1 / Issue 1 / 2021
- Published online by Cambridge University Press:
- 24 June 2021, e2
-
- Article
-
- You have access
- Open access
- HTML
- Export citation
A history of high-power laser research and development in the United Kingdom
- Part of
-
- Journal:
- High Power Laser Science and Engineering / Volume 9 / 2021
- Published online by Cambridge University Press:
- 27 April 2021, e18
-
- Article
-
- You have access
- Open access
- HTML
- Export citation
KPC-Producing Enterobacter cloacae Transfer Through Pipework Between Hospital Sink Waste Traps in a Laboratory Model System
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s308-s309
- Print publication:
- October 2020
-
- Article
-
- You have access
- Export citation
Baseline results from the European non-interventional Antipsychotic Long acTing injection in schizOphrenia (ALTO) study
-
- Journal:
- European Psychiatry / Volume 52 / August 2018
- Published online by Cambridge University Press:
- 01 January 2020, pp. 85-94
-
- Article
-
- You have access
- Open access
- HTML
- Export citation
An analysis of the impacts of Cretaceous oceanic anoxic events on global molluscan diversity dynamics
-
- Journal:
- Paleobiology / Volume 45 / Issue 2 / May 2019
- Published online by Cambridge University Press:
- 10 April 2019, pp. 280-295
-
- Article
- Export citation
The Hennepin Ketamine Study Investigators’ Reply
-
- Journal:
- Prehospital and Disaster Medicine / Volume 34 / Issue 2 / April 2019
- Published online by Cambridge University Press:
- 03 May 2019, pp. 111-113
- Print publication:
- April 2019
-
- Article
-
- You have access
- HTML
- Export citation
2507 A novel multi-photon microscopy method for neuronavigation in deep brain stimulation surgery
-
- Journal:
- Journal of Clinical and Translational Science / Volume 2 / Issue S1 / June 2018
- Published online by Cambridge University Press:
- 21 November 2018, pp. 2-3
-
- Article
-
- You have access
- Open access
- Export citation
9 - Balancing Privacy and Public Safety in the Post-Snowden Era
- from Part II - Surveillance Applications
-
-
- Book:
- The Cambridge Handbook of Surveillance Law
- Published online:
- 20 October 2017
- Print publication:
- 12 October 2017, pp 227-247
-
- Chapter
- Export citation
Modifiable Risk Factors for the Spread of Klebsiella pneumoniae Carbapenemase-Producing Enterobacteriaceae Among Long-Term Acute-Care Hospital Patients
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 38 / Issue 6 / June 2017
- Published online by Cambridge University Press:
- 11 April 2017, pp. 670-677
- Print publication:
- June 2017
-
- Article
- Export citation
The Last Interglacial Ocean
-
- Journal:
- Quaternary Research / Volume 21 / Issue 2 / February 1984
- Published online by Cambridge University Press:
- 20 January 2017, pp. 123-224
-
- Article
- Export citation
Age Dating and the Orbital Theory of the Ice Ages: Development of a High-Resolution 0 to 300,000-Year Chronostratigraphy1
-
- Journal:
- Quaternary Research / Volume 27 / Issue 1 / January 1987
- Published online by Cambridge University Press:
- 20 January 2017, pp. 1-29
-
- Article
- Export citation