The use of a horse as an athlete requires the provision of a high-energy ration to the individual. Commonly, this is achieved by complementing forages with concentrates rich in starch and non-fibre carbohydrates. However, horses have moderate capacity to digest starch in the small intestine due to low concentrations of pancreatic amylase( Reference Kienzle, Radicke and Landes1 ). In this respect, high amounts of starch in the ration are not entirely degraded in the pre-caecal gut and flow into the hindgut, constituted by the caecum and the colon, where it can trigger microbiota dysbiosis. Compared with high-fibre diets, high-starch diets lead indeed to an increase in starch-utilising micro-organisms, at the expense of fibre-utilising micro-organisms that are essential for degrading plant cell walls. This results in a shift in fermentation patterns with higher lactate and propionate concentrations leading to increasing acidity. This has recently been reviewed by Julliand & Grimm( Reference Julliand and Grimm2 ). Moreover, an overload of starch was reported to induce histological damage of the hindgut epithelial mucosa in horses( Reference Krueger, Kinden and Garner3 ). It was also shown to increase hindgut mucosa permeability both in vitro ( Reference Weiss, Evanson and Green4 ) and in vivo ( Reference Weiss, Evanson and MacLeay5 ) in equine studies. It was hypothesised that the low pH and/or the lactate produced by starch fermentation could be responsible for epithelium disruption. In livestock, an increase in intestinal permeability can result in leakage of bacteria, or bacterial products, such as lipopolysaccharides (LPS), through the intestinal epithelium to the bloodstream, with subsequent activation of an acute phase response( Reference Gómez-Laguna, Salguero, Pallarés and Veas6 , Reference Mani, Weber and Baumgard7 ).
Some nutritional solutions have been developed with the objective of supporting the equine hindgut microbiota during dietary challenges. A new product combining the yeast Saccharomyces cerevisiae strain CBS 493·94 and the microalgae Aurantiochytrium limacinum has been proposed. S. cerevisiae strain CBS 493·94 has long been used in equine nutrition for its beneficial effect on the hindgut microbial ecosystem in horses fed either high-fibre or high-starch diets. Globally, supplementation may have a positive effect on microbiota by enhancing its fibre-digesting capacity and by limiting dysbiosis with high-risk feeding practices, such as ingestion of large quantities of starch( Reference Jouany, Medina and Bertin8 – Reference Moore, Newman and Spring10 ). Conversely, the effect of A. limacinum on the hindgut microbial ecosystem in horses had yet to be studied. Initial investigations in ruminants suggested that, thanks to its high content in n-3 fatty acids (mainly in DHA), it could modify ruminal microbiota composition and activity( Reference Boeckaert, Vlaeminck and Dijkstra11 , Reference Castro-Carrera, Toral and Frutos12 ). However, it is not known whether those modifications are beneficial for the animal. n-3 Fatty acids have also been reported to have immunomodulatory and anti-inflammatory effects in various mammals( Reference Calder13 ). Therefore, supplementing A. limacinum in horses fed high-starch diets could modulate the hindgut microbiota structure and activity and help limit inflammation where integrity of the intestinal mucosa is compromised leading to LPS translocation.
In equine studies, in vivo evaluation of nutritional solution effects on the hindgut microbiota was performed either by studying directly caecum or colon content using fistulated horses( Reference Jouany, Medina and Bertin8 , Reference Medina, Girard and Jacotot9 , Reference Respondek, Goachet and Julliand14 , Reference Grimm, Julliand and Philippeau15 ), or, more frequently, using faeces, which is non-invasive and less labour-intensive( Reference Furtado, Barboza and Brandi16 – Reference Faubladier, Chaucheyras-Durand and da Veiga22 ). However, to our knowledge, no study has ever verified that changes observed in the faeces under supplementation were consistent with those in the hindgut microbial ecosystem. In horses, the caecum and the right ventral (RV) colon (first part of the large colon) are anatomically far from the rectal ampulla, being separated by the left ventral colon, the left dorsal colon and the right dorsal colon plus the small colon. The retention times of the digesta in the whole hindgut is longer than some 20 h( Reference Van Weyenberg, Sales and Janssens23 ). Along the different compartments, there is an active absorption (SCFA, amino acids, water) and an evolution of the bacterial structure( Reference Julliand and Grimm24 ). It is thus assumed that the response to a supplementation could be different between the proximal hindgut and the faeces.
The present study aimed to assess the impact of supplementing the combination of S. cerevisiae and A. limacinum on both the caecal, colonic and faecal microbial ecosystem and blood inflammatory markers of horses fed a high-fibre or high-starch diet.
Methods
All experimental procedures were evaluated and approved by the local ethical committee (Comité d’Ethique de l’Expérimentation Animale Grand Campus Dijon).
Animals and management
Six adult crossbred geldings (495 (SD 18) kg body weight (BW)) fitted with cannulas in the caecum (5/6 horses) and the RV colon (6/6 horses) were individually housed in 14 m² free stalls bedded with wood shavings. Each day all horses had access to a dry paddock for 2 h. They were exercised six times a week in an automatic walker for 1 h/d at 4–6 km/h. They were up to date with their vaccinations against influenza and tetanus (ProteqFlu, Merial SAS) and were dewormed (Hippomectin, Audevard).
Diets and supplementation
Throughout the experimental trial, horses were successively fed two different iso-energetic diets providing 105 % energy requirements of a horse in ‘very light exercise’ according to French recommendations( 25 ). The first diet (H) was composed of 100 % meadow hay (2·1 % BW DM hay/d, Table 1). The second diet (B) contained 56 % meadow hay and 44 % rolled barley (1·4 % BW DM/d of which 0·8 % BW DM hay/d, Table 1). The hay/barley diet was formulated for providing starch at 0·20 % BW per meal which is the maximum recommendation of starch per meal( Reference Julliand, De Fombelle and Varloud26 ). It also provided 0·8 % BW DM/d which is less than the minimal recommendation of 1 % BW DM of hay( 27 ). Both diets were equally distributed across two meals offered at 08.00 and 16.30 hours. The test supplement provided 100 g of microalgae A. limacinum (All-G-Rich™, Alltech®) and 10 g (1 × 109 CFU/g) of yeast S. cerevisiae (Yea-sacc™, Alltech®) daily. The microalgae powder was pelleted with molasses and anise infusion to improve its palatability. The supplementation was equally split across the two meals. Water and a block of mineralised salt containing sodium chloride (Na 39 %/SOLSEL®; European Salt Company) were offered free choice.
Table 1. Nutritional and biochemical composition of the diets

BW, body weight; CP, crude potein; NDF, neutral-detergent fibre; ADF, acid-detergent fibre; ADL, acid-detergent lignin.
* Cellulose = ADF – ADL and hemicelluloses = NDF – ADF.
Experimental design
Horses were randomly assigned to two treatment groups over two experimental periods in a 2 × 2 Latin square design. Each experimental period was a total of 8 weeks, separated by a wash-out period of 7 weeks (Fig. 1). Group 1 received the yeast/microalgae product in experimental period 1 (P1) only, while Group 2 received the product during experimental period 2 (P2). In each experimental period, both groups were fed the H diet over the first 4 weeks and then, over 5 d, gradually moved onto the B diet for the remainder of that experimental period. During the wash-out period, all horses received hay as the only feedstuff, without supplementation.
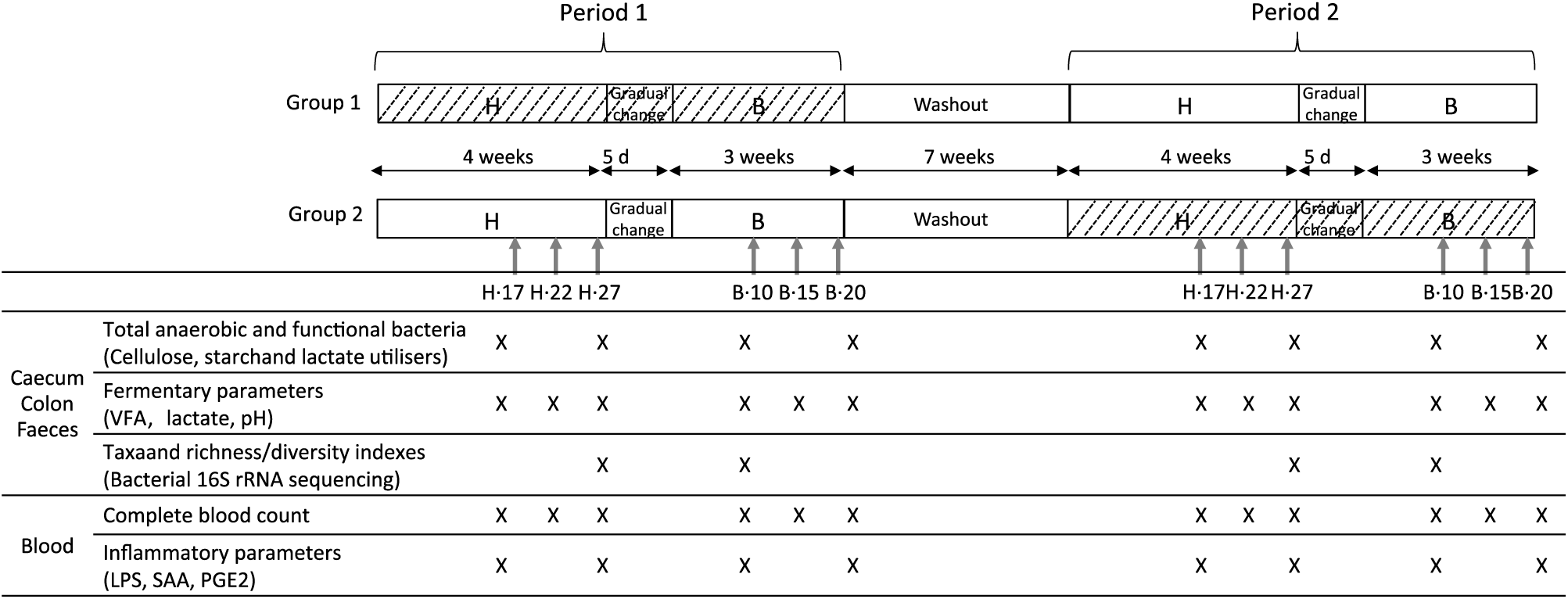
Fig. 1. Experimental design of the study and analyses performed during the H diet (100 % hay) on the 17th, 22nd and 27th days (H.17, H.22 and H.27) and during the B diet (56 % hay + 44 % barley) on the 10th, 15th and 20th days (B.10, B.15 and B.20) in the caecum, right ventral colon, faeces and blood. VFA, volatile fatty acids; LPS, lipopolysaccharides; SAA, serum amyloid A. ![]() , Yeast/microalgae supplementation.
, Yeast/microalgae supplementation.
Sample collection
During each experimental period, caecal, colonic, faecal and blood samples were collected 4 h after the morning meal on days 17, 22 and 27 during the H diet (H.17, H.22 and H.27) and days 10, 15 and 20 during the B diet (B.10, B.15 and B.20) (Fig. 1).
The caecum and colon contents were obtained by gravity via the cannulas. Fresh faeces were collected by rectal grab. An aliquot of unfiltered content was sampled in a container filled to capacity (to avoid the presence of oxygen) for microbiological analyses performed immediately after the sample collection. Another aliquot of the unfiltered contents was sampled for molecular biology analysis (1 ml, storage at –80°C until analysis). The remaining part of the contents was filtered (100 μm filter), and 1 ml was immediately stored at –20°C until analysis after the addition of a preservative (0·1 ml of a solution of 5 % H3PO4 + 1 % HgCl2) for further volatile fatty acid (VFA) analysis. Another filtered 1 ml was immediately stored at –20°C until analysis for further lactate analysis.
Blood samples were taken from the jugular vein into 4 ml vacutainer tubes (Venosafe, Terumo) containing either K3-EDTA as an anticoagulant (for LPS, PG-E2 and complete blood count analyses) or not (for serum amyloid A analysis). Tubes for complete blood count analysis were directly sent to an extern laboratory, and all other tubes were centrifuged (10 min at 1000 g ). Plasma or serum was removed and frozen at –20°C (for LPS and PG-E2 analysis) or directly assayed (for serum amyloid A analysis).
Analyses were carried out according to the sampling day (Fig. 1).
Functional bacterial groups enumeration
Total anaerobic bacteria and cellulose-utilising, starch-utilising and lactate-utilising bacteria were assayed using conventional anaerobic culture techniques. The content of caecum, colon and faeces (1 g) was serially diluted to decimal in a mineral solution( Reference Bryant and Burkey28 ) and then inoculated in roll tubes on specific media under continuous flow of CO2 ( Reference Hungate29 ).
Concentrations of total anaerobic bacteria (dilutions representing 10−5, 10−6 or 10−7 ml of intestinal contents) were determined on a non-selective modified complete agar medium( Reference Leedle and Hespell30 , Reference Julliand, de Vaux and Millet31 ) after 48 h of incubation at 38°C. Concentrations of lactate-utilising bacteria (dilutions representing 10−3, 10−4 or 10−5 ml of intestinal contents) were determined on a selective medium( Reference Mackie and Heath32 ) after 48 h of incubation at 38°C. Concentrations of starch-utilising bacteria (dilutions representing 10−4, 10−5 and 10−6 ml of intestinal contents) were determined on a modified selective medium containing 1 % (w/v) soluble starch as the main energy source( Reference Lowe, Theodorou and Trinci33 ), after 72 h of incubation at 38°C. Most probable number( Reference Clarke and Owens34 ) of cellulose-utilising bacteria (dilutions representing 10−5, 10−6 and 10−7 ml of intestinal contents) were determined using a modified broth medium( Reference Julliand, de Vaux and Millet31 , Reference Halliwell and Bryant35 , Reference Baruc, Dawson and Baker36 ), after 15 d of incubation at 38°C.
Bacterial 16S rDNA extraction, sequencing and bioinformatical analysis
Total DNA was extracted from 0·25 g caecal, RV colonic or faecal content as described by Yu & Morrison( Reference Yu and Morrison37 ). The quantity of obtained DNA was assessed on a spectrophotometer (Eppendorf), and its purity was evaluated by calculating A260:280 ratios to verify contamination by proteins (ratio values between 1·6 and 2 were acceptable for nucleic acid extractions from digestive samples). DNA samples were then frozen at –80°C. The V3–V4 hypervariable region (459 bp in E. coli) of the bacterial 16S ribosomal RNA gene (16S rRNA, 1542 bp in E. coli) was amplified by PCR using the forward and reverse primers F343 (CTTTCCCTACACGACGCTCTTCCGATCTACGGRAGGCAGCAG)( Reference Liu, Lozupone and Hamady38 ) and R784 (GGAGTTCAGACGTGTGCTCTTCCGATCTTACCAGGGTATCTAATCCT)( Reference Andersson, Lindberg and Jakobsson39 ). The PCR mix was constituted of 10 ng of DNA, 5 µl of buffer (MP Biomedicals), 1 µl of dNTP mix, 0·5 µl of Taq polymerase (5 U/µl MTPTM Taq DNA Polymerase, Sigma), 1·25 µl (20 µm) of forward and reverse primers and was completed to 50 µl with sterile water. The conditions of PCR were: one cycle at 94°C during 1 min, followed by 30× (94°C during 1 min, 65°C during 1 min and 72°C during 1 min). The amplification was followed by a melting program (72°C during 10 min). To verify the correct amplification of the V3–V4 region, the DNA samples were electrophoresed on a 2 % agarose gel. The PCR products (4 µl) were applied per well and electrophoresed for 75 min at 45°C using a fixed voltage of 70 V on a Mupid One system. After purification with magnetic beads, amplicons were submitted to a second PCR aiming at ligate Illumina adapters and an index allowing the identification of the sample. The this second PCR mix was the same than for the first PCR with a forward primer (AATGATACGGCGACCACCGAGATCTACACTCTTTCCCTACACGAC) and a reverse primer (CAAGCAGAAGACGGCATACGAGAT-Index-GTGACTGGAGTTCAGACGTGT). The conditions of this second PCR were similar to the previous ones but twelve cycles of elongation were performed instead of thirty. PCR products were again purified using magnetic beads. The resulting PCR products were sequenced using an Illumina MiSeq run of 250 base paired-ends according to the manufacturer instructions (Illumina Inc.). The quality of the run was checked using control libraries generated from the PhiX virus (Illumina PhiX control; Illumina Inc.).
Bioinformatic analyses were performed using the FROGS pipeline (Find Rapidly operational taxonomic unit (OTU) with Galaxy Solution), which associates several tools to work on the file of sequences and obtain both the abundance table of OTU and their taxonomic affiliation( Reference Escudié, Auer and Bernard40 ). OTU which are not present in at least two samples, or whose abundance was less than 5 × 10−5, were removed. Remaining OTU were then aligned to the silva123 16S database( Reference Pruesse, Quast and Knittel41 ) using BLAST and were affiliated to the highest taxonomic rank possible. The abundance table and the associated multi-hit list were created. Richness (number of species) and diversity (Shannon and inverse Simpson) indexes were calculated from the abundance table.
Biochemical analyses
The pH of the caecum, colon and faeces was measured directly after the samples collection on filtered contents using the HI 1053 electrode (Hanna instrument) connected to an electronic pH-meter (CyberScan pH 510, Eutech Instrument). VFA (acetate (C2), propionate (C3), butyrate (C4), valerate (C5)) concentrations were assayed by a GLC (Clarus 500, PerkinElmer)( Reference Jouany42 ). Each VFA was also expressed as a percentage of the total VFA concentration, and the ratio ((C2 + C4)/C3) was calculated according to Sauvant et al.( Reference Sauvant, Chapoutot and Archimède43 ). d-Lactate and l-lactate concentrations were measured spectrophotometrically at 340 nm (MRX revelation, Dynatech Laboratories) in microplate using an enzymatic colorimetric method (Megazyme, d-/l-lactic acid (d-/l-lactate) (Rapid) Assay Kit, Megazyme International Ireland Ltd) as published in Grimm et al.( Reference Grimm, Philippeau and Julliand44 ).
Blood analyses
Blood LPS concentration was determined by direct quantitation of 3-hydroxymyristic acid using HPLC coupled with tandem MS (HPLC-MS/MS, Agilent QQQ 6460 device) as presented by Pais de Barros et al.( Reference Pais de Barros, Gautier and Sali45 ). Plasma PG-E2 concentration was evaluated using an immunoassay kit (DetectX Prostaglandin E2, Arbor Assays), following the regular format protocol. Serum amyloid A concentrations were assayed in an extern laboratory (LABEO Frank DUN-COMBE) using an ELISA Phase kit from Eurobio. Complete blood counts were performed in an extern laboratory (LABEO Frank DUNCOMBE) using a haematology system (ADVIA 120, Siemens). The blood cell count included count of erythrocytes and leucocytes with their distribution into neutrophils, basophils, eosinophils, monocytes, small and large lymphocytes. The Hb concentration and haematocrit percentage were also measured, as well as blood platelets.
Statistical analyses
As horses fitted with cannulas both in caecum and right ventral colon represent a scarce resource due to ethical concerns, six animals were recruited in the present study. A Latin square design was chosen so that each animal acted as its own control, thus reducing the individual variability and allowing to reach a significance level of 0·05 with a power of 80 % for the variables tested.
Richness and diversity indexes, as well as normalised taxa (phylum, class, family and genus level), were statistically analysed by the MIXED procedure of SAS (version 9.3, SAS Institute Inc.) for each digestive compartment separately, testing the period (P1 and P2), the diet (H and B), the supplementation (supplemented or not) and their interactions. The model included horse and horse × period in the random statement. For all other data, the model was similar, but tested the sampling days instead of the diet and included the sampling day as repeated measures with horse × period as subject. In case of a significant sampling day effect, orthogonal contrast was performed to compare the two diets (H v. B). For both models, least-square means were calculated for all variables using Tukey multiple comparison procedure. The significance threshold was set at P < 0·05, and trends were considered at P < 0·1. Principal coordinates analysis based on Bray–Curtis distance of the 1931 OTU was performed using the Vegan package in R.
Results
Two horses were removed before the end of the trial due to health troubles. The first horse developed colic during P1 the day after the B.10 samples collection and was thus immediately removed. It was included in P2. The second horse developed a disease unrelated to digestion and had to be euthanised after the first collection (H.17) in P2.
Sequencing metrics
The initial 2 059 711 sequences of the 16S rDNA V3–V4 regions obtained from the sixty-two caecal, colonic and faecal samples were reduced to 1 032 444 after quality filtering during bioinformatics processes, with a mean of 16 652 (SD 4377) sequences per sample. These sequences were clustered into 1931 OTU (1622 in the caecal samples, 1856 in the colonic samples and 1793 in the faecal samples) that grouped into eleven phyla, twenty-five classes, thirty-eight orders, seventy-one families and 189 genera. The captured diversity was large, as the coverage estimates were comprised between 98·0 and 99·7 % in the sixty-two samples (caecum: 99·16 (SD 0·07); RV colon: 99·16 (SD 0·06); faeces: 98·91 (SD 0·06)).
Out of the 1931 OTU, 1697 were considered as unknown species (87·9 %) and 186 were multi-affiliated (9·6 %). Forty-three OTU in the caecum (2·5 % of the total abundance), forty-six in the colon (2·8 % of the total abundance) and thirty-seven in the faeces (1·1 % of the total abundance) were identified at species level.
Firmicutes and Bacteroidetes were identified as the two major bacterial phyla in the caecum (40·1 and 48·1 %, respectively), colon (57·3 and 30·2 %, respectively) and faeces (51·6 and 34·2 %, respectively) (online Supplementary Fig. S1).
Richness and diversity indexes
In all compartments, no significant interaction period × diet × supplementation was observed on the different richness and diversity indexes. In the caecum, the species number (P = 0·039), the Shannon (P = 0·009) and the inverse Simpson (P = 0·0002) indexes were significantly lower during the B diet than during the H diet, irrespective of the supplementation (Table 2). In the colon, the inverse Simpson index (P = 0·006) was significantly lower and the Shannon index tended to be significantly lower (P = 0·099) during the B diet compared with the H diet. In the faeces, the species number (P = 0·026) and the Shannon index (P = 0·020) displayed significant interaction between the diet and the supplementation, with their values decreasing only in non-supplemented horses fed the B diet. The faecal inverse Simpson index significantly increased with the yeast/microalgae supplementation irrespective of the diet (P = 0·043).
Table 2. Richness and diversity indexes in the caecum, right ventral (RV) colon and faeces of horses fed the diets H (100 % hay) and B (56 % hay + 44 % barley) and receiving (S) or not (NoS) a yeast/microalgae supplementation
(Mean values and pooled standard errors)
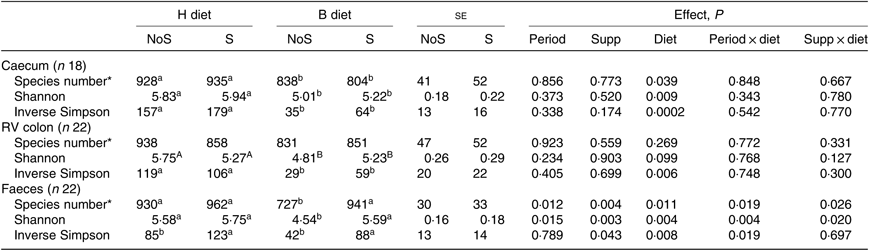
a,b Mean values within a row with unlike superscript letters are significantly different (P < 0·05).
A,B Mean values within a row with unlike superscript letters tend to be significantly different (P < 0·1).
* Number of operational taxonomic units.
Bacterial taxa
In the caecum, colon and faeces, significant effect of the diet was observed at all levels of bacterial taxonomy on a large number of taxa (online Supplementary Table S2). Using principal coordinates analysis, caecal, colonic and faecal samples formed clear distinct groups according to the diet (H or B) fed (Fig. 2).
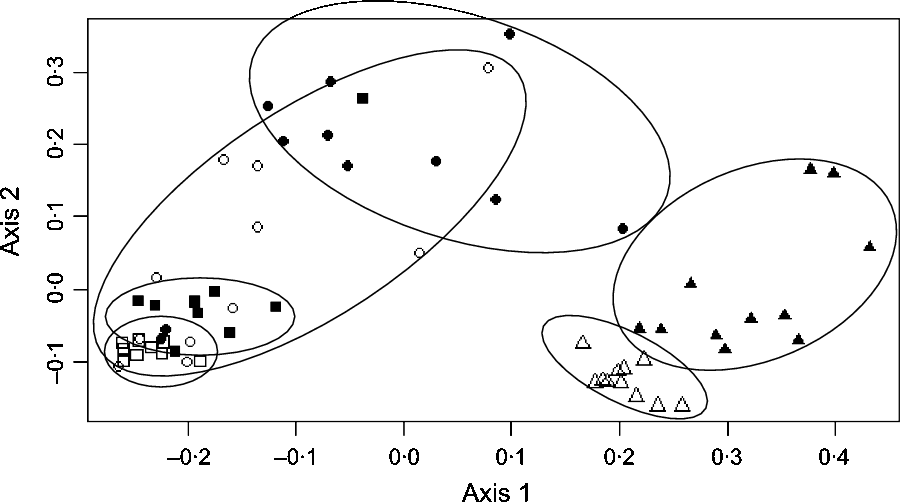
Fig. 2. Principal coordinates analysis at the operational taxonomic units level showing the bacterial structure in the caecum, right ventral (RV) colon and faeces of horses fed either 100 % DM hay (H) or 56 % DM hay + 44 % DM barley (B). ![]() , Caecum H;
, Caecum H; ![]() , caecum B;
, caecum B; ![]() , RV colon H;
, RV colon H; ![]() , RV colon B;
, RV colon B; ![]() , faeces H;
, faeces H; ![]() , faeces B.
, faeces B.
In the caecum (Table 3), a significant diet × supplementation interaction was found on the relative abundance of the genus Ruminiclostridium (P = 0·033) that decreased when the diet changed from H to B diet only in non-supplemented horses. Significant effects of the yeast/microalgae supplementation were observed on the relative abundance of Clostridiales family XIII (P = 0·025), and on the relative abundance of the genus Pseudobutyrivibrio (P = 0·033), which significantly increased in supplemented horses, irrespective of the diet.
Table 3. Relative abundance of bacterial taxa showing significant (or trend) supplementation × diet interaction or supplementation effect in the caecum, right ventral (RV) colon and faeces of horses fed the diets H (100 % hay) and B (56 % hay + 44 % barley) and receiving (S) or not (NoS) a yeast/microalgae supplementation
(Mean values with their pooled standard errors)
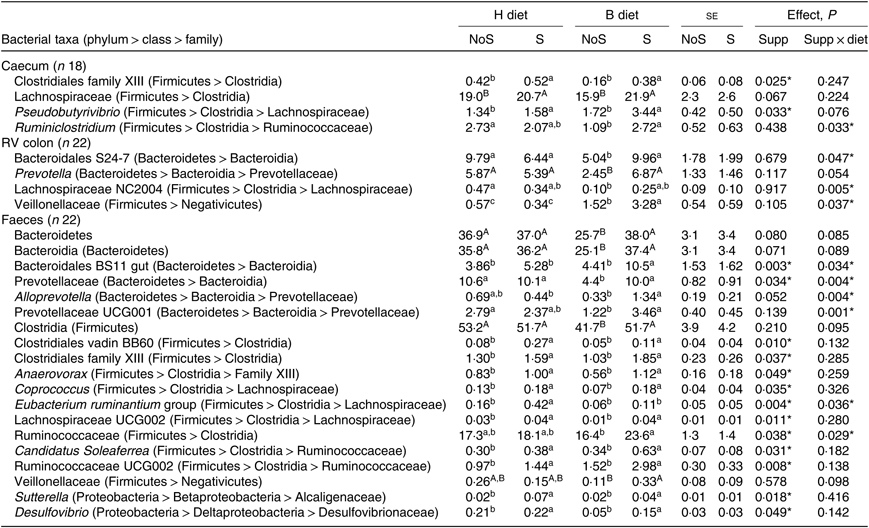
a,b,c Mean values within a row with unlike superscript letters are significantly different (P < 0·05).
A,B Mean values within a row with unlike superscript letters tend to be significantly different (P < 0·1).
In the colon (Table 3), when horses changed from the H to the B diet, a decrease of family Bacteroidales S24-7 (P = 0·047) and genus group Lachnospiraceae NC2004 (P = 0·005) relative abundances was observed in non-supplemented horses only. The Veillonellaceae relative abundance increased significantly in horses fed the B diet compared with the H diet, with a larger increase in horses receiving the supplementation (P = 0·037).
In the faeces (Table 3), the relative abundances of faecal Bacteroidales BS11 gut (P = 0·034) and Alloprevotella (P = 0·004) were significantly higher in supplemented horses fed the B diet. When horses changed from the H to the B diet, the decrease of Prevotellaceae (P = 0·004) and Prevotellaceae UCG001 (P = 0·001) relative abundances was observed in non-supplemented horses only. During the H diet, the relative abundance of the Eubacterium ruminantium group increased with supplementation (P = 0·036). During the B diet, the Ruminococcaceae relative abundance was higher in supplemented horses (P = 0·029). Significant effects of the yeast/microalgae supplementation were observed at the family level on the relative abundances of Clostridiales vadin BB60 (P = 0·010), family XIII of Clostridiales (P = 0·037), Alcaligenaceae (P = 0·018) and Desulfovibrionaceae (P = 0·049) and at the genus level on the relative abundances of Anaerovorax (P = 0·049), Candidatus soleaferrea (P = 0·031), Coprococcus (P = 0·035), Desulfovibrio (P = 0·049), Lachnospiraceae UCG002 (P = 0·011), Ruminococcaceae UCG002 (P = 0·008) and Sutterella (P = 0·018). All these bacterial taxa relative abundances were higher when horses received the yeast/microalgae supplementation, irrespective of diet.
Bacterial functional groups
Except for the cellulose-utilising bacterial concentration in the caecum (P = 0·037), no significant interaction period × diet × supplementation was observed on the functional bacterial concentration. No significant interaction supplementation × sampling day was found on the bacterial groups (Table 4). No significant effect of the supplementation was observed. A significant effect of the sampling day associated with a significant diet effect was seen on the caecal lactate utilisers (P = 0·007) and on all the RV colonic and faecal bacterial groups (P ≤ 0·008). The total anaerobes, starch utilisers, lactate utilisers increased and the cellulose utilisers decreased when the diet changed from H to B.
Table 4. Bacterial functional group concentrations in the caecum, right ventral (RV) colon and faeces of horses depending on sampling day within the diets H (100 % hay) and B (56 % hay + 44 % barley) irrespective of supplementation
(Mean values with their pooled standard errors)
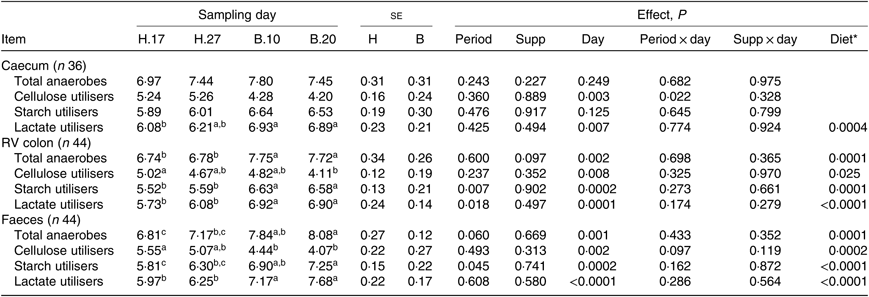
a,b,c Mean values within a row with unlike superscript letters are significantly different (P < 0·05).
* Orthogonal contrasts testing the diet (H v. B) were performed only if a sampling day effect was significant.
Biochemical parameters
No significant period × sampling day × supplementation interaction, supplementation × sampling day interaction and supplementation effect was observed on biochemical parameters, irrespective of sample type. A significant effect of sampling day was found on several parameters, as well as a significant effect of the diet: the change of diet from H to B diet caused a decline in total VFA (P = 0·028) and acetate (P = 0·002) concentration and an increase in propionate (P = 0·007) concentration in the caecum. In the RV colon, there was an increase in valerate concentration (P = 0·0005) and, in the faeces, an increase in all VFA concentration (Table 5). The VFA ratio significantly decreased in the caecum (P < 0·0001) and RV colon (P = 0·017), but not in the faeces. No significant dietary variations were observed on lactate concentrations.
Table 5. Biochemical parameters in the caecum, right ventral (RV) colon and faeces of horses depending on sampling day within diets H (100 % hay) and B (56 % hay + 44 % barley) irrespective of supplementation
(Mean values with their pooled standard errors)
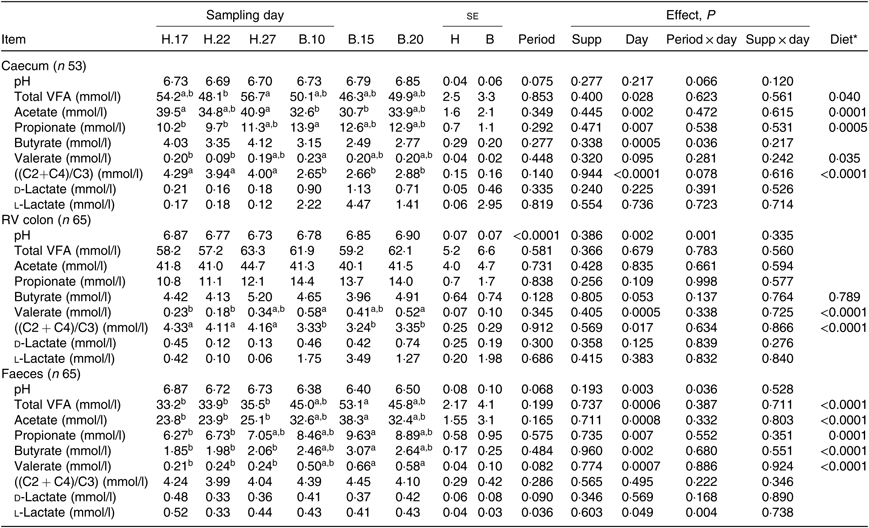
VFA, volatile fatty acids; C2, acetate; C4, butyrate; C3, propionate.
a,b Mean values within a row with unlike superscript letters are significantly different (P < 0·05).
* Orthogonal contrasts testing the diet (H v. B) were performed only if a sampling day effect was significant.
No significant interaction supplementation × sampling day and supplementation effect was observed on VFA proportions. A significant effect of day of sampling concomitant with a significant effect of diet was observed on acetate and propionate proportions both in the caecum (P < 0·0001) and the RV colon (P = 0·011 and P = 0·023 for acetate and propionate proportions, respectively), but not in faeces (Fig. 3). In the hindgut compartments, the proportions of acetate were significantly lower and the proportions of propionate significantly higher when horses were fed the B diet compared with the H diet.
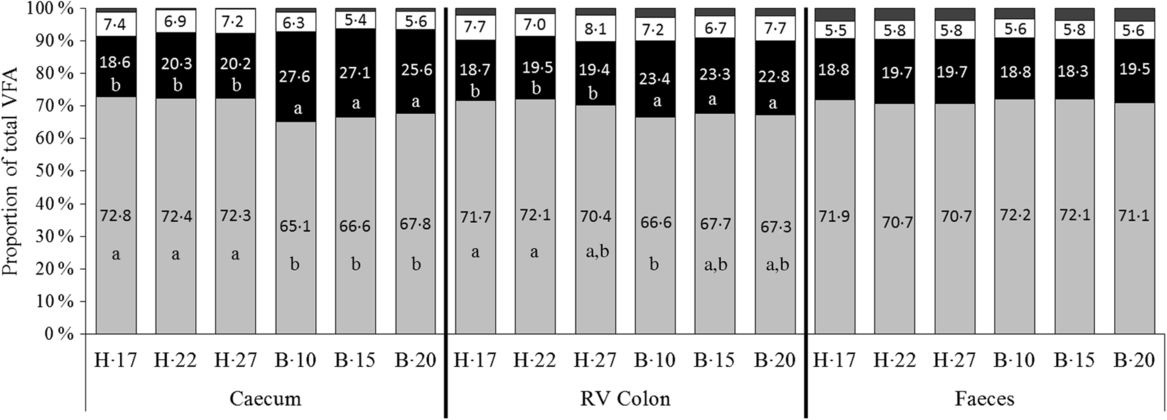
Fig. 3. Proportions of volatile fatty acids (VFA) in the caecum, right ventral (RV) colon and faeces of horses depending on the sampling day within diets H (100 % hay) and B (56 % hay + 44 % barley) irrespective of supplementation. ![]() , Others;
, Others; ![]() , butyrate (C4);
, butyrate (C4); ![]() , propionate (C3);
, propionate (C3); ![]() , acetate (C2).
, acetate (C2).
Blood parameters
No significant interaction period × diet × supplementation was observed on the blood parameters except on the serum amyloid A concentration (P = 0·028). No significant interaction between the diet and the supplementation nor simple significant effects of the diet or supplementation were observed on the different blood parameters (online Supplementary Table S3).
Discussion
The present study provides a better insight into the phenomenon of dysbiosis in the equine hindgut thanks to the holistic approach combining the use of culture and molecular techniques simultaneously with microbial activity measurements. Furthermore, the present study investigated for the first time a possible way for alleviating digestive dysbiosis using a combination of yeast and microalgae supplementation both in the hindgut and the faeces of horses.
When horses changed from a 100 % hay diet to a 56/44 hay/barley diet, their hindgut microbial ecosystems were clearly challenged in terms of diversity and function. The hay/barley diet was associated with a lower bacterial diversity both in the caecum and RV colon in agreement with a previous study( Reference Hansen, Avershina and Mydland46 ) and with a lower richness in the caecum only. A lack of sufficient diversity in the bacterial community has been reported to diminish its stability and ability to withstand perturbation( Reference Bäckhed, Fraser and Ringel47 ). This would imply that the microbial ecosystem of the horse hindgut under the hay/barley diet would be less resistant to stress than under the hay diet, thus leading potentially to a dysbiosis responsible for colic. Interestingly, the individual that developed colic on the 11th day of the high-starch diet (non-supplemented) showed the lowest diversity in the caecum (Shannon index: 4·12) and the faeces (Shannon index: 2·41) compared with other horses.
Similarly to previous data( Reference Medina, Girard and Jacotot9 , Reference Julliand, de Fombelle and Drogoul48 ), the fibrolytic function was disturbed under the hay/barley diet compared with hay, as demonstrated by the lower cellulose-utilising bacteria and ((C2 + C4)/C3) ratio. This was in accordance with the decrease of the relative abundance of several bacterial taxa, such as Fibrobacteres, Prevotellaceae, Rikenellaceae, Bacteroidales S24-7 and several genera of Lachnospiraceae and Ruminococcaceae that could be implicated in plant cell wall degradation( Reference Flint, Scott and Duncan49 – Reference Ormerod, Wood and Lachner51 ). Concomitantly, the amylolytic function was higher with the hay/barley diet compared with the hay diet, as shown by the increase of the starch-utilising bacteria and propionate and valerate concentrations. Lactobacilli and streptococci have been the most studied starch-utilising bacteria using culture technics( Reference Julliand and Grimm24 ). Our sequencing data confirmed that in the proximal hindgut, Streptococcaceae family relative abundances were higher with the hay/barley diet (from 0·06 to 0·13 % in the caecum and from 0·02 to 0·59 % in the RV colon in non-supplemented horses). This increase even if it remained limited was in accordance with previous studies using sequencing technics( Reference Hansen, Avershina and Mydland46 ) or classical culture technics( Reference Medina, Girard and Jacotot9 , Reference Julliand, de Fombelle and Drogoul48 ). On the contrary, we did not observe changes in lactobacilli by sequencing 16S bacterial rDNA, whereas a large increase was reported in studies using classical culture technics. In agreement with previous data( Reference Hansen, Avershina and Mydland46 ), the relative abundance of the Succinivibrionaceae family (member of the Proteobacteria phylum) that possesses several species able to degrade starch( Reference Hippe, Hagelstein and Kramer52 ) largely increased in both equine caecum and RV colon when the high-starch diet was fed. We demonstrated that the lactate-utilising function was higher with the hay/barley diet than that with the hay diet: the number of lactate-utilising bacteria increased in the caecum and RV colon not only using culture technics but also using sequencing technics, as we observed an increase of the Veillonellaceae family, confirming previous findings( Reference Hansen, Avershina and Mydland46 , Reference Daly, Proudman and Duncan53 ).
Despite the fact that it induced disturbances in the hindgut microbial ecosystem, the high-starch diet did not lead to alteration of plasma LPS concentration as expected, nor alteration of other measured inflammatory markers. It may be assumed that the hay/barley diet used in the present study was not sufficiently disturbing to induce an inflammatory reaction, precluding demonstration of any anti-inflammatory potential of the supplementation. Indeed, we did not record a decrease of pH with the hay/barley diet, thus preventing from a high acidity in the hindgut and subsequent mucosa disruption, LPS translocation into the plasma and later development of an acute-phase response( Reference Weiss, Evanson and Green4 , Reference Weiss, Evanson and MacLeay5 ).
Regarding supplementation, we expected to observe the combined effects of yeast (fibrolytic function improvement and dysbiosis limitation as reported in horses) and microalgae (microbial ecosystem alteration in ruminants and inflammatory modulation in mammals) when the product was supplemented. Factually, we found an increase of the relative abundance of Pseudobutyrivibrio in the caecum that could be due to the microalgae supplementation rich in long-chain unsaturated fatty acids, mostly DHA. Indeed, it has been reported that this genus has a role in lipid biohydrogenation in the rumen( Reference Paillard, McKain and Chaudhary54 , Reference Boeckaert, Vlaeminck and Fievez55 ). Besides in the rumen, bacteria belonging to Pseudobutyrivibrio produce multiple xylanases responsible for high xylanolytic activity and are mainly butyrate producers( Reference Čepeljnik, Križaj and Marinšek-Logar56 , Reference Zorec, Vodovnik and Marinšek-Logar57 ). Yeast/microalgae supplementation could thus contribute to increase the fibrolytic function in the caecum.
Another significant impact of yeast/microalgae supplementation was the reduction of the variations in the bacterial taxa relative abundance due to the high-starch diet. Firstly, the supplementation avoided the decrease in relative abundance of some potential fibre-utilising bacteria in the caecum (Ruminiclostridium) and in the RV colon (Bacteroidales S24-7, Prevotella and Lachnospiraceae NC2004) that was observed in non-supplemented horses when the hay/barley diet was fed. In addition, the increase in relative abundance of the lactate-utiliser Veillonellaceae reported during the hay/barley diet was amplified in the RV colon of supplemented horses. This suggested a higher capacity to lower the lactate concentration, thus limiting acidity and associated issues. The increase in lactate-utilising bacteria with the high-starch diet was in accordance with previous equine( Reference Jouany, Medina and Bertin8 , Reference Moore, Newman and Spring10 ) and ruminant studies( Reference Pinloche, McEwan and Marden58 ) that investigated S. cerevisiae supplementation effects. However, this increase did not modify the lactate-utilising function (culture technics) or lactate concentration.
The changes in bacterial taxonomy with the yeast/microalgae supplementation, either in the sense of the increase of fibre-utilising representatives or in the sense of dysbiosis limitation, had limited consequences on the cultivable cellulose-utilising bacteria and VFA production from the microbial ecosystem. It is possible that some other bacterial functions not investigated in the present study, such as the hemicellulose-utilising or pectin-utilising, were altered with supplementation. Assessing different enzymes would allow better characterisation of the impact of supplementation.
The absence of a yeast supplementation effect on the cellulolytic function could be explained by the difference in terms of quantity of S. cerevisiae strain CBS.493.94 used between studies mentioned in the literature and ours( Reference Jouany, Medina and Bertin8–Reference Moore, Newman and Spring10 ). We fed our horses top dressed 10 g/d Yea-Sacc, which represented 1 × 1010 CFU/d per horse as recommended. This was slightly lower than the dose of 4·5 × 1010 CFU/d distributed in two earlier studies( Reference Jouany, Medina and Bertin8 , Reference Medina, Girard and Jacotot9 ). Another study did not specify the dose of Yea-Sacc that was used( Reference Moore, Newman and Spring10 ).
In ruminants, many studies highlighted a diminution of the total VFA concentration in the rumen, accompanied by a higher pH when a DHA-rich product (microalgae A. limacinum or fish oil) was supplemented. They also reported a lower molar proportion of acetate in parallel to a higher proportion of propionate in the rumen of supplemented ruminants( Reference Boeckaert, Vlaeminck and Dijkstra11 , Reference Shingfield, Kairenius and Ärölä59 , Reference Zhu, Fievez and Mao60 ). Contrary to ruminants, the microalgae A. limacinum supplemented to horses passed firstly through the stomach and the small intestine before reaching the fermentative reservoir in the hindgut. The resistance of the microalgae to stomach acidity and pre-caecal digestion was unknown. Moreover, the quantity supplemented (100 g/d) was maybe not sufficient to have an effect on hindgut microbial functions.
In many mammals, faeces are very often used to study the impact of a nutritional supplement on the digestive ecosystem. However as explained before, the response to a supplementation could be different between the proximal hindgut and the faeces of horses. In our study, we thus compared changes observed in the faeces under yeast/microalgae supplementation with those measured in the proximal hindgut microbial ecosystem.
In a coherent way with what was observed in the caecum and RV colon, the bacterial functional groups and the biochemical parameters did not vary in the faeces of horses supplemented with S. cerevisiae and A. limacinum. Moreover, a very limited number of taxa varied in the same direction in the hindgut and faeces: the relative abundance of the Family XIII of Clostridiales increased both in the caecum and the faeces when horses were supplemented irrespective of diet, and the relative abundance of Veillonellaceae and the belonging genus Anaerovibrio was greater with supplementation in colon and faeces during the hay/barley diet.
However, while we expected more noticeable effects of the yeast/microalgae in the hindgut than in the faeces, our results demonstrated greater changes in the faecal bacterial diversity and structure. Indeed, whereas the bacterial richness and diversity were unchanged in the hindgut with supplementation, the faecal diversity of supplemented horses increased whatever the diet (higher inverse of Simpson index). The loss of richness and diversity that occurred in the faeces during the high-starch diet was avoided with supplementation (higher number of species and Shannon index). Moreover, compared with the hindgut, the supplementation had a greater impact on the bacterial structure in faeces, as more bacterial taxa had a modified relative abundance with supplementation. As in the hindgut, the variations in faeces suggested that supplementation could increase fibrolytic function whatever the diet and limit dysbiosis when the horses’ diet changed from high fibre to high starch.
Conclusion
We highlighted that some parameters of the caecal and RV colonic microbial ecosystems were positively affected by the supplementation combining S. cerevisiae and A. limacinum. Nevertheless, more studies using microalgae alone might be beneficial, as well as studies combining microalgae and yeast, to characterise the effects of the algae more definitively in horse. It has recently been reported that microalgae could possess prebiotic properties( Reference de Jesus Raposo, de Morais and de Morais61 ) and this could be explored in horses.
In view of our data, the faecal microbial ecosystem appeared to be affected by the supplementation even more than the proximal hindgut microbial ecosystem. This suggests that faeces can be a good indicator for evaluating modifications in the hindgut microbial ecosystem, reflecting not only variations in the proximal but probably also those of the distal hindgut. More investigation into how representative faeces are of the hindgut environment is required to make accurate evaluation of nutritional products on intestinal function using faeces.
Acknowledgements
We would like to thank Alltech for the financial support of the present study, and for the revisions of this manuscript. We thank the entire technical staff who contributed actively to the care of horses, samplings and laboratory analyses at Agrosup Dijon. We are grateful to the genotoul bioinformatics platform Toulouse Midi-Pyrenees and Sigenae group for providing help and storage resources, thanks to Galaxy instance (http://sigenae-workbench.toulouse.inra.fr).
This work was also supported by the Regional Council of Bourgogne – Franche Comté, the Fonds Européen de DEveloppement Régional (FEDER).
P. G. and V. J. conceived and oversaw the entire project. They elaborated and conducted the experimental study and managed the laboratory analysis of the samples, statistics and interpretation of results. S. C., G. P. and L. C. supervised the sequencing, bioinformatical and biostatistical analyses. All authors participated in article drafting or revising.
There are no conflicts of interest.
Supplementary material
For supplementary materials referred to in this article, please visit https://doi.org/10.1017/S0007114519002824













