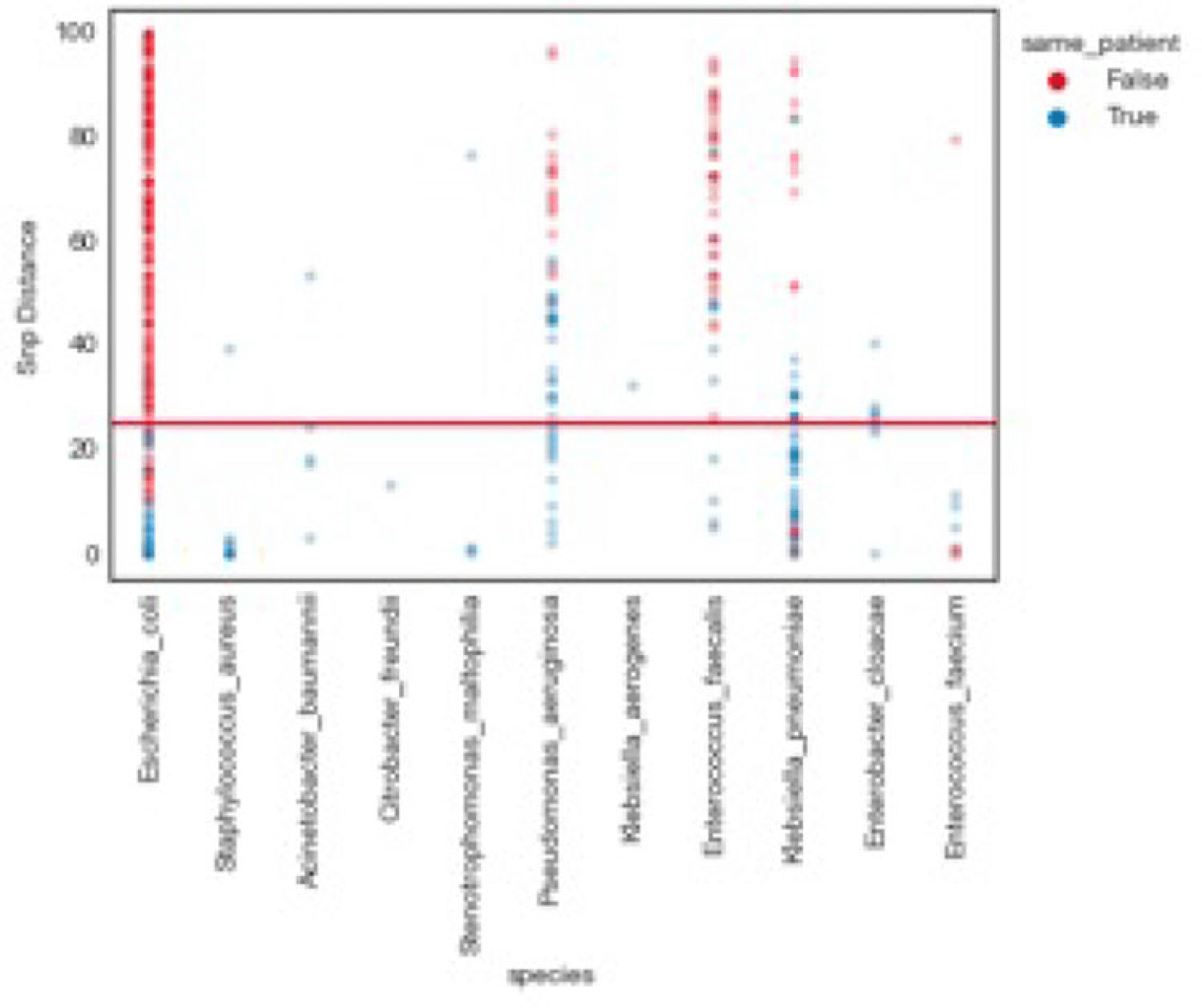
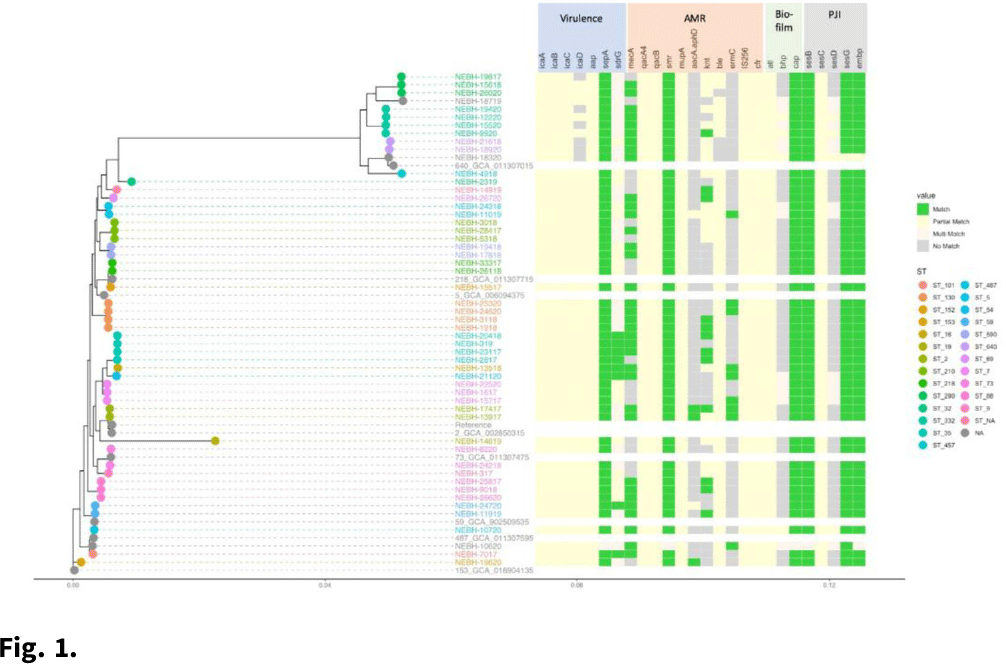
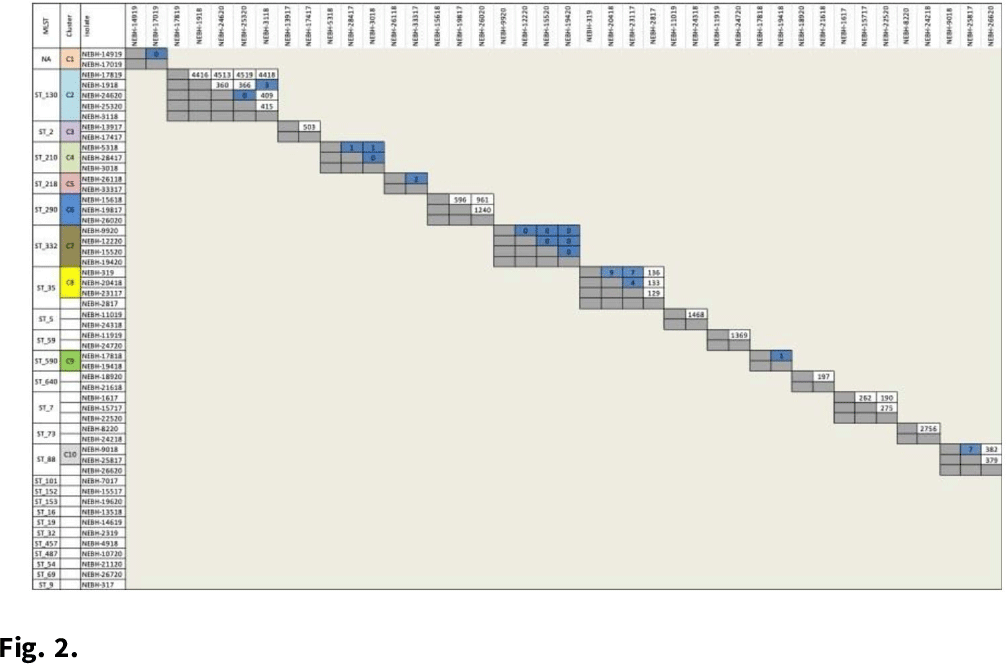
Background: Whole-genome sequencing (WGS) is well established as a high-resolution method for measuring bacterial relatedness to better understand infection transmission in cases of healthcare-associated infections (HAIs). However, sequencing is still rarely used in HAI investigations due to a lack of access to computational analysis platforms with actionable turnaround times. Single-nucleotide polymorphism (SNP) analysis is typically used to determine bacterial relatedness. However, SNP-based methods often require a suite of bioinformatics tools that can be difficult to use and interpret without the expertise of a trained computational biologist. These obstacles become more significant in the case of prospective, real-time surveillance of HAIs, which can require the analysis of a large number of isolates. To enable the use of WGS for proactive determination of infection outbreaks, a rapid, automated method that can scale to large data sets is needed. Methods: Here, we demonstrate the capabilities of ksim, a novel automated algorithm to determine the clonality of bacterial samples using WGS. ksim measures the number of shared kmers (genomic subsequences of length k) between bacterial samples to determine their relatedness. ksim also filters out accessory genomic regions, such as plasmids, that can confound genetic relatedness estimates. We benchmarked the accuracy and speed of ksim relative to an SNP-based pipeline on simulated data sets (with sequencing reads generated in silico) and on 9 clinical-cluster data sets (6 publicly available and 3 real-time data sets from Massachusetts General Hospital [MGH]). We also used ksim to determine the relatedness of >5,000 historical clinical bacterial isolates from MGH, collected between 2015 and 2019. Results: ksim first preprocesses raw sequencing data to generate a common data structure, after which it computes the genomic distance between bacterial samples in ∼0.2 seconds in simple cases and in ∼4 seconds in complex cases when accessory genome filtering is required. In simulations across 5 species, ksim determined clonality (defined as <40 SNPs) with high accuracy (sensitivity, 99.7% and specificity, 99.6%). ksim performance on 9 clinical HAI data sets demonstrated its sensitivity (99.4%) and specificity (90.8%) compared to an SNP-based pipeline. ksim efficiently analyzed >5,000 clinical samples from MGH and found previously unidentified transmission clusters. Conclusions:ksim shows promise for rapid clonality determination in HAI outbreaks and has the potential to scale to tens of thousands of samples. This method could enable infection control teams to use WGS for prospective outbreak detection via an automated computational pipeline without the need for specialized computational biology training.
Funding: Day Zero Diagnostics and the NIH provided Funding: for this study.
Disclosures: Mohamad Sater reports salary from Day Zero Diagnostics.