The Colorado Twin Registry (CTR) database includes the extensive information about twins and their families recruited from birth records and school records that is housed at the University of Colorado Boulder’s Institute for Behavioral Genetics (IBG). This population-based registry holds records from twins born in Colorado, or who were living in Colorado at the time of ascertainment through school records, beginning with 1968 births. Two previous reports have been published in this series of Twin Research and Human Genetics special issues: the first (Rhea et al., Reference Rhea, Gross, Haberstick and Corley2006) gave a history of the CTR and described samples drawn from the CTR participating in particular studies; the second (Rhea et al., Reference Rhea, Gross, Haberstick and Corley2013) provided updates on additional studies using the samples described in 2006.
The purposes of this update are to provide information about additional studies using the ongoing longitudinal samples described in 2013, to describe new samples formed from the CTR, to describe the genotyping that has been done on some samples and to describe how phenotypic and genotypic data from CTR samples have contributed to various multi-sample collaborations and consortia. A companion piece in this issue will describe how one twin sample from the CTR, the Longitudinal Twin Sample, has been combined with a longitudinal adoption study, the Colorado Adoption Project, to create a genetically informative archive of infancy-to-mid-adulthood data for the study of aging (CATSLife; Wadsworth et al., Reference Wadsworth, Corley, DeFries, Plomin and Reynolds2019, this issue).
Ascertainment, Recruitment, Assessment, Samples and Studies
The methods used for CTR ascertainment are described in more detail in Rhea et al. (Reference Rhea, Gross, Haberstick and Corley2006). To summarize, recruitment of families began in 1984 with mailings from the Colorado Department of Health (CDH) to the parents of twins born in 1982. For twins born between 1982 and 1998, families that expressed interest in twin research were enrolled in the CTR. A similar procedure was used to contact and enroll twins born between 1968 and 1981 as young adults. From 1999 to the present a negative consent protocol has been followed, in which the CDH contacted eligible (no record of twin death) twin families on IBG’s behalf, but only forwarded information about families who did not refuse to participate and whose mail was not returned as undeliverable. Since the negative consent procedure began, approximately 85% of twin births have been forwarded each year to IBG for further contact. For years in which the CDH kept records of why approximately 15% of twin families were not made available (2006–2009), absence of forwarding addresses was twice as common as outright refusals (only 5% of total eligible twin families). Addresses were generally forwarded to IBG in the second half of the year following the twins’ birth year. Additional twin families identified through mailings from participating school districts were asked if they also wished to be enrolled in the CTR. A small number of older twin pairs were identified through surname/birthdate pairings in Department of Motor Vehicles records available until 2000. The CDH’s publications and generous cooperation has allowed IBG researchers to compare demographic trends over time (see Rhea et al., Reference Rhea, Corley, Heath, Iacono, Neale and Hewitt2017 as an example) and compare participating twin families with nonparticipants (Rhea et al., Reference Rhea, Gross, Haberstick and Corley2006).
From 1999 to 2017 (the last year of available twin births), the CTR has grown almost exclusively from the information supplied by the CDH through its negative consent procedure. In that period, a total of 17,000 twin families have been identified for potential contact, including 5445 female–female pairs, 5383 male–male pairs and 6172 opposite-sex (OS) twin pairs. The number of twin families forwarded by the CDH to the registry each year has been relatively constant, ranging from a low of 772 families in 1999 to a high of 1007 families in 2007. Because no zygosity information is available directly from the CDH, it is only after contact and recruitment into a study performing zygosity ascertainment that it is possible to determine what percentage of the same-sex samples are monozygotic (MZ) or dizygotic (DZ). Using Weinberg’s rule (MM + FF − MF/total = MZ rate) as employed by Rhea et al. (Reference Rhea, Corley, Heath, Iacono, Neale and Hewitt2017) to examine birth cohort trends in MZ twin percentages, we would conclude that the MZ twin rate among the same-sex individuals is approximately 43% (assuming no bias on the basis of zygosity for refusal to participate or moving without leaving a forwarding address).
Once contacted by mail, and beginning in 1995, family respondents were asked to send back information about their family structure (number and ages of siblings) and information about education and job titles for the twin parents, along with additional contact information such as phone numbers and email addresses. In years prior to 1995, information requested from families upon initial contact was more limited. Families identified as willing to participate in twin research represent the pool of subjects from which twin samples were drawn.
As described in the two earlier papers (Rhea et al., Reference Rhea, Gross, Haberstick and Corley2006, Reference Rhea, Gross, Haberstick and Corley2013), the CTR distinguishes between twin samples and twin studies, as several of the longitudinal samples have been used for multiple studies, with different funding mechanisms and research goals. After initial sample recruitment, primarily from among those families who had returned a baseline information form, and after initial assessments for a particular study were completed, families were sometimes asked to participate in additional studies. Rhea et al. (Reference Rhea, Gross, Haberstick and Corley2013) describe four active samples. The longest running sample of twins (recruited between 1976 and the present, with birth years as early as 1965) is that of the Colorado Learning Disabilities Research Center (CLDRC; identified as the CLS in Rhea et al., Reference Rhea, Gross, Haberstick and Corley2006), focusing on twins aged 8–16 at recruitment. The youngest-in-age twin sample, recruited during infancy and early toddlerhood (5–14 months), is Colorado’s Longitudinal Twin Sample (LTS) based on twins born between 1984 and 1990. A sample of adolescent twins recruited between 1997 and 2002 and with birth years between 1979 and 1990 forms the Community Twin Sample (CTS). A fourth sample of preschool twins born between 1995 and 2000 formed the Colorado subsample of the International Longitudinal Twin Study of Early Reading Development sample (ERDS; Olson et al., Reference Olson, Keenan, Byrne, Samuelsson, Coventry, Corley and Hulslander2007). Characteristics of a sample are partially dependent on the criteria imposed by the initial study for which the sample was recruited. Thus, for most of the samples, families were located within an approximately 2-h driving distance of Boulder, Colorado, where IBG is located. Additional details about sample characteristics and the studies using those samples are found in Rhea et al. (Reference Rhea, Gross, Haberstick and Corley2006).
Sample Overlap
The LTS, CTS and ERDS longitudinal samples are nonoverlapping. However, some of the CLDRC sample partially overlaps with LTS and CTS. For the Haworth et al. (Reference Haworth, Wright, Luciano, Martin, de Geus, van Beijsterveldt and Plomin2010) mega-analysis, described in the Collaborations and Consortia section below, overlaps were detected and duplicate families deleted using surname/birthdate/forename matching (a slight underestimate due to name/birthdate changes), yielding an 18.5% estimate of overlap in the CLDRC sample with LTS and CTS. As CLDRC recruitment has continued, overlap has dropped below 15%.
Samples versus Studies
As described in Rhea et al. (Reference Rhea, Gross, Haberstick and Corley2013), multiple studies can use a given sample, and a study can use multiple samples. To avoid excessive subject burden, studies using the same samples have synchronized their testing schedules. To briefly summarize the relationships between samples and studies prior to this current update, we describe studies in terms of the sample, with a hyphenated suffix to indicate the particular study. The CLDRC sample does not require a suffix, although a follow-up study using the sample will be described in the next section. The ERDS sample, with multiple waves of assessment, also represents just a single study. The Longitudinal Twin Sample has a long history as the Longitudinal Twin Study (LTS), tracing back to infancy and currently extended by a new study (LTS-CATSLife), but families were also asked to participate in the Center for Antisocial Drug Dependence (LTS-CADD) starting at age 12, and subsequently to participate in an Executive Function study (LTS-EF) starting at age 17. The CTS sample has branched into several substudies: the CTS-CADD, for which it was recruited and an Executive Function extension of the CADD for a subset of the sample (CTS-EF). Changes to the studies associated with these samples will be described in more detail in the next section. An overview of the connections between samples and studies is shown in Figure 1, with the acronyms for the sample-study combinations shown in parentheses.

Fig. 1. Colorado Twin Registry samples and subjects active since 2012. Note: Adapted and updated from Table 1, Rhea et al. (Reference Rhea, Gross, Haberstick and Corley2013).
Update on Studies Using Previously Described Samples
In this section, we briefly describe work done with CTR samples described in the Rhea et al. (Reference Rhea, Gross, Haberstick and Corley2013) article, including studies which were in progress in 2013 and additional studies that are currently being conducted in 2019. Because the CLDRC has become more closely integrated with the CTR, and because it was less thoroughly described in the last CTR update (Rhea et al., Reference Rhea, Gross, Haberstick and Corley2013), we describe it at some length.
Colorado Learning Disabilities Research Center
The CLDRC (in various permutations and names; DeFries et al., Reference DeFries, Filipek, Fulker, Olson, Pennington, Smith and Wise1997) is the longest running of the twin studies based at the IBG, University of Colorado Boulder, and has amassed the largest number of twin pairs in any single sample. Through December 2018, the CLDRC has tested a cumulative total of 2497 twin pairs and 561 non-twin siblings. The central focus of the CLDRC has been the study of genetic and environmental influences on reading performance and reading difficulties and comorbid disorders such as attention-deficit hyperactivity disorder (ADHD), and difficulties with mathematical calculation and comprehension have also been assessed. Initial psychometric assessments of these measures have been complemented by laboratory assessments of executive functions, reading processing in twin brains and genotyping of buccal- and blood-derived DNA (see the genotyping section below for more details about genetic assessments). The most recent round of testing was completed in 2016, with a newly funded grant starting in 2018. This center renewal features functional magnetic resonance imaging (fMRI), expanding on earlier work on structural MRI assessment (Pennington et al., Reference Pennington, Filipek, Lefly, Chhabildas, Kennedy, Simon and DeFries2000). Zygosity determination in this sample has been based primarily on tester ratings of twin similarity on physical characteristics with questionable zygosities resolved using Short Tandem Repeat (STR) markers in DNA collected from buccal swabs. Between 2000 and 2017, 10.8% of twin-pair zygosities have been resolved through DNA analysis. Most twin pair ascertainment occurs through participating school districts; only 11% have been recruited directly from Colorado birth records in the CTR. However, the majority of twins (62.4%) with birth years in 1995 and later who have been tested by the CLDRC have also been identified in the birth records Those CDLRC twins not found in the birth records represent a combination of twin families with twins born outside of Colorado and families who declined to enroll in the CTR or who could not be reached by mailings from the CDH
The CLDRC sample has, in part, been longitudinally assessed (Wadsworth et al., Reference Wadsworth, DeFries, Willcutt, Pennington and Olson2015), first through a separate grant (DC01590), and since 2012 as part of the main CLDRC center grant. (HD027082). For subjects initially tested from 1996 onward, eligible twin pairs were recontacted by mail and invited to participate in a follow-up round of assessments approximately 5 years after their initial assessments. Eligibility required that the twins be willing to come as a pair to the testing facility in Boulder, Colorado, and that they have not aged out of the adolescent age window (currently limited to 13–19 years of age). Eligibility for follow-up did not depend on reading difficulty status of either pair member at initial assessment. During a full-day test session, assessments of reading and language processes, executive functions, cognitive performance and achievement, and behavioral issues that mirrored those conducted during initial assessments were completed. In the last concluded grant cycle for 2012–2017, a total of 302 twins were retested (49.3% female) at an average age of 15.7 years (SD = 1.8 years). Another cycle of follow-up testing began in 2018. A list of CLDRC publications can be found in Table S1 in the Supplemental material.
LTS-CADD
Completion of a third wave of testing using the CADD protocol occurred in 2014. This wave of testing, funded by NIDA center grant DA011015, focused on substance use and comorbid psychopathology into early adulthood (N = 762, mean age = 22.8, SD = 1.3), and HIV-relevant risk behaviors.
LTS-EF
The second wave of the LTS-EF study was completed in 2014 (N = 751, mean age = 22.8, SD = 1.3; Gustavson et al., Reference Gustavson, Stallings, Corley, Miyake, Hewitt and Friedman2017). Testing between the LTS-CADD third wave and LTS-EF second wave was synchronized, with both the CADD protocol and the EF protocol typically administered within the same week. An additional wave of data collection from these subjects (fMRI Twin Study of Cognitive Control), beginning in 2014 with targeted completion in 2019, focuses on fMRI, with both resting state and executive function processing segments.
LTS-CATSLife
A third study using the LTS sample is the CATSLife study of cognitive aging as described in more detail in the companion piece in this issue (Wadsworth et al., Reference Wadsworth, Corley, DeFries, Plomin and Reynolds2019). Testing began in 2015 with scheduled completion in 2020 and is synchronized when possible with the third wave of LTS-EF.
LTS-COMN
A fourth study using the LTS sample, in conjunction with the CTS sample, is the collaborative study between Colorado and Minnesota studying the impact of marijuana legalization on the substance use behaviors of twins in states with permissive attitudes toward recreational marijuana use (Colorado) and more restrictive attitudes (including toward medical marijuana, Minnesota). This will represent the fifth wave of substance use assessment for these subjects. The first assessments occurred in 2018 and are scheduled to complete in 2021.
Although the LTS sample was originally recruited from a relatively small geographic area in Colorado, because of the limitation on driving distance to Boulder, Colorado, as twins have aged and been assessed by different studies, more than 20% of the sample is reporting current addresses (obtained via LTS-CATSLife) outside of Colorado. Figure 2 shows the contrast between locations at birth, and locations as of 2019, for those living in the US’ 48 contiguous states.
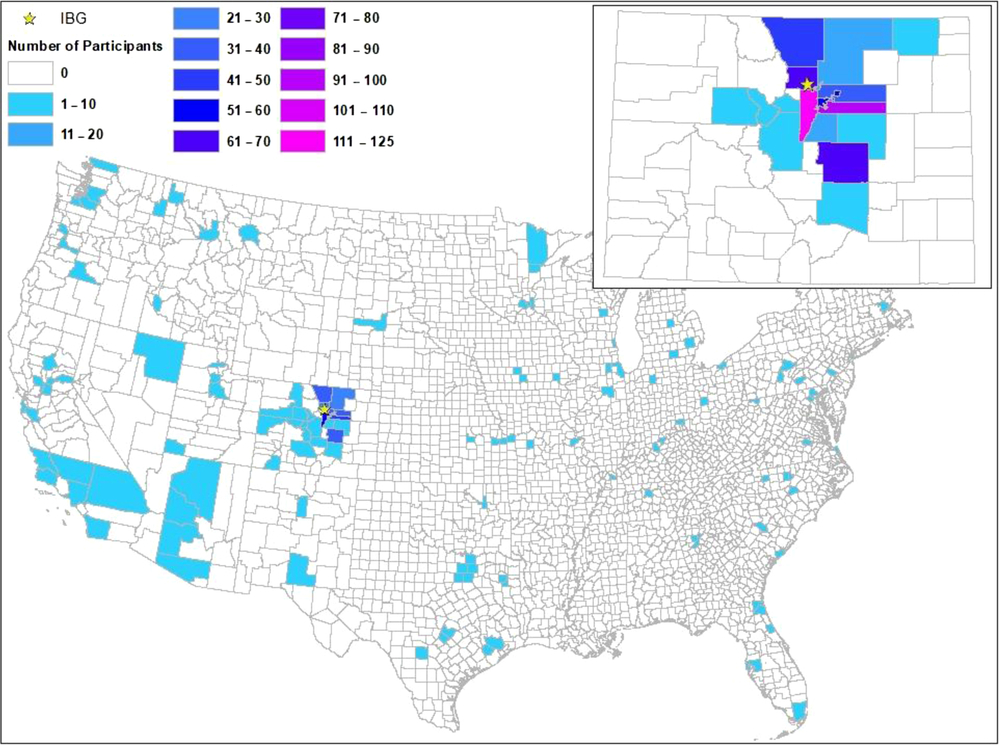
Fig. 2. Counties of origin (in Colorado), inset above, and counties of current residence in 48 US states (below).
CTS-CADD
In parallel with the LTS-CADD testing, a third wave of data completion, focusing on substance use, comorbid psychopathology and risky behaviors, was finished in 2014 (N = 1740, mean age = 26.70, SD = 2.29).
CTS-EF
Selected twins tested at wave 2 of the CADD using the CTS-EF protocol were asked to participate in an fMRI study (A Neurogenetic Basis for Risky Decisions) during wave 3 of the CADD. The study was designed to assess brain function under a risk-taking experiment (the Colorado Balloon Game). Two aspects in particular were of interest: making a decision to engage in a risky decision and responding to the reward/loss feedback from the decision. A total of 80 CTS twins provided usable fMRI data.
CTS-COMN
In parallel with the LTS-COMN testing, assessment of the CTS sample began in 2018, with scheduled completion in 2021. This will represent the fourth wave of substance use assessments for this sample, following subjects into their fourth decade. In addition to assessment of substance use and associated psychopathology in the presence or absence of recreational marijuana legalization, the assessment batteries include questions about current living situations, personality, relationships, parenting, finances, problem-solving, and beginning in 2019, sleep behaviors.
CTS-PAIN
In this study, ‘Brain and genetic predictors of individual differences in pain and analgesia,’ a selected sample of CTS twins is participating in an fMRI scan while experiencing different types of safe, non-damaging painful stimulation; for example, heat or pressure on fingers and thumbs. Nonintrusive sensing of physiological measurements, including heart rate and respiration rate, is measured while in the magnet. Initial testing began in 2018, with scheduled completion in 2022.
ERDS
Final testing of this sample, focused on reading comprehension and academic achievement, was completed in 2015 after subjects had completed ninth grade (N = 931, mean age = 15.46, SD = 0.32). These longitudinal subjects have thus been assessed up to six times: before kindergarten, after kindergarten, after first grade, after second grade, after fourth grade and after ninth grade.
New Samples Derived from the CTR
CoTwins
The Colorado Online Twin Study (CoTwins) was initiated in 2015. The study was designed to combine traditional survey methods (in-person interviews, online questionnaires) with smartphone technology to assess short-term changes in adolescent substance use and related behaviors using phone-pushed questionnaires. Additional data available through smartphone technology, such as location data and internet search histories, were also collected. Sample 1 consisted of 335 twin pairs (141 female–female, 106 male–male and 88 opposite-sex), recruited in 2015 and 2016 at ages 14–17 years (mean age = 16.06, SD = 1.13) followed for up to 2 years. Zygosity determination was based primarily on tester ratings of twin similarity based on six physical characteristics, including eye color and hair characteristics; same-sex pairs were rated 55.5% DZ and 45.5% MZ (see Brazel et al., Reference Brazel, Corley, Phelan, Friesr, Subramonyam, Rhea and Vrieze2017, e.g., analyses illustrating MZ, DZ and OS twin proximity similarity). Although most of the sample represented new recruits from the registry, 14.0% overlapped with the previously ascertained CLDRC sample and 24.2% overlapped with the ILTS/ERDS sample.
Recruitment for a second twin sample began in 2019. In this iteration, recruitment focused on high school juniors and seniors for a 2-year assessment span intended to focus on the transition from high school to the post-high school year. Subjects from sample 1 are also eligible for continuing participation. Although subject protocols were largely comparable for the two samples, participants in the second were asked to be more actively engaged in data collection through auditory sampling and requests for photographs from their immediate environs.
ABCD — Twin Hub
The Adolescent Brain Cognitive Development (ABCD) Study (https://www.nimh.nih.gov/research/research-funded-by-nimh/research-initiatives/adolescent-brain-cognitive-development-abcd-study.shtml; https://www.addictionresearch.nih.gov/abcd-study) is one of the National Institute of Health’s most ambitious collaborative efforts. Subject recruitment of 9- and 10 year olds at 22 different sites began in 2016, and when completed in October 2018 had enrolled 11,875 individuals. In Colorado, a total of 129 non-twin adolescents and 215 same-sex twin pairs were recruited. Colorado is one of the four sites (the others based in Minnesota, Missouri and Virginia) participating in a twin hub with a total recruitment goal of 800+ twin pairs, with a particular emphasis in Colorado on twins with at least one Hispanic parent. This sample of 9- and 10 year olds was drawn from the birth registry and does not overlap with other existing IBG-based twin samples
All ABCD adolescent subjects undergo an initial baseline standardized assessment, including an approximately 1.5-h structural and fMRI session, and assessments of cognitive development, substance exposure, environmental perceptions and screens for psychopathology. The goal is to see each subject on a yearly basis, with follow-up fMRIs scheduled for every other year. In Colorado, the initial baseline wave of assessments began in September 2016 and was completed in August 2018; the first annual follow-up began in September 2017 and the second follow-up (with the additional fMRI session) began in September 2018. From 2017 onward, two waves of assessments are being conducted in the same year, with half the sample undergoing a follow-up year without fMRI testing, and the other half of the sample undergoing a more extensive test protocol, including fMRI testing.
Domains
Table 1 provides an overview of the domains covered for the different twin samples. Although different samples may have coverage in the same domains, test instruments and domain foci may differ between samples, reflecting the priorities of different funding agencies. For example, in the Home Environment domain, the CLDRC and the ERDS measures include measures of print exposure and familial reading habits, while the other studies include measures indicating degree of parental monitoring or familial conflict.
Table 1. Phenotypic domains assessed by sample

Note: X indicates that measures in the domain were collected on a partial or selected subsample, XX indicates that measures within the domain were collected systematically.
Genotyping and Genetic Analyses in the CTR Samples
Different genotyping methods and analyses have been used for the various CTR samples, depending on the period in which the corresponding studies were conducted, the genotyping resources available at that time and the analytic goals of the studies. Our previous report on the CTR (Rhea et al., Reference Rhea, Gross, Haberstick and Corley2013) listed some of the genotyping that had been done for two samples, the LTS and CTS. This section is intended to develop the topic further, to cite some representative papers using genetic variation from the samples and to provide an overview of the extent of different genotyping efforts within the CTR samples. Genotyping in the twin samples has often been done on a subsample of individuals or pairs due to the needs of a particular study. Some examples include selecting only one twin per pair to obtain a sample of unrelated individuals, or selecting twins as matched controls for clinical or affected subjects, or selecting only one twin from a known MZ pair. Table 2 at the end of this section indicates which methods have been used for which twin samples and distinguishes between partial or selected sampling, and more systematic genotyping efforts. For all the twin samples for which DNA was collected, donation of samples was not a prerequisite for participation in a particular study, although participation rates were high (>95%) when asked to give samples.
Table 2. Samples by genotyping description

Note: X indicates that genotyping was done on a partial or selected subsample, XX indicates that systematic genotyping was done within the sample.
Zygosity Imputation
The use of genetic markers to impute zygosity has been the most commonly used type of genotyping in the CTR samples. Samples for which at least some zygosity genotyping has been done include CLDRC, LTS, CTS, ERDS, CoTwins and ABCD. The most common method of genotyping has been the use of a small number of short tandem repeat (STR) microsatellite markers with high degrees of heterozygosity similar to the panel of STR markers used in the US’ Combined DNA Index System (CODIS), but for privacy concerns, with nonoverlapping markers. However, in the ABCD study, zygosity is imputed from within twin-pair variation across multiple single-nucleotide polymorphisms (SNPs), currently from the more than 500,000 SNPs available through BioRealm’s Smokescreen genotyping array developed for addiction research in collaboration with the National Institute on Drug Abuse (NIDA). For most samples, the primary source of DNA has been saliva buccal cells, but blood samples have been collected in both the CTR (through CATSLife) and in ABCD from willing participants.
Markers for Linkage Analysis
Early analyses in the CTR samples searching for genetic sources of variation for reading disability included twins from the CLDRC sample (Cardon et al., Reference Cardon, Smith, Fulker, Kimberling, Pennington and DeFries1994, Reference Cardon, Smth, Fulker, Kimberling, Pennington and DeFries1995). This study used five STR markers to infer the location of a genetic region of interest on chromosome 6 within the human leukocyte antigen complex. The evolution and refinement of additional genetic analyses for reading disability can be traced in the Supplementary table of publications S1 for the CLDRC. In particular, Deffenbacher et al. (Reference Deffenbacher, Kenyon, Hoover, Olson, Pennington, DeFries and Smith2004) used a combination of linkage analysis and direct SNP association tests in the 6p21.3 region of chromosome 6 with both STR and SNP markers to narrow in on a small set of genes in this chromosomal region.
Variable Number Tandem Repeats
A number of variable number tandem repeat (VNTR) genetic polymorphisms have been of recurring interest to researchers in psychiatric genetics. Some of the most prominent include those in genes for neurotransmitter receptors and transporters and include variants in the genes DRD4, MAOA, SLCA3 and SLC6A4. Selected VNTRs have been genotyped in the LTS and CTS samples (Heiman et al., Reference Heiman, Larsson, Stallings, Young, Smolen and Hewitt2003). A recent report included VNTR data from twins in a validation sample indicating high imputation accuracy for VNTRs from genome-wide SNP data (Border et al., Reference Border, Smolen, Corley, Stallings, Brown, Conger and Evans2019).
Targeted SNP Genotyping
Selected SNP genotyping has been carried out in CLDRC, CTS and LTS and has varied in magnitude from single SNPs to custom SNP arrays designed to efficiently assay multiple SNPS within multiple genes of interest. As an example, Corley et al. (Reference Corley, Zeiger, Crowley, Ehringer, Hewitt, Hopfer and Krauter2008) included CTS and LTS twins as controls in a case–control study using a targeted gene assay (custom SNP chip) yielding more than a 1000 SNPs spread across 49 genes selected as candidates influencing substance use disorders. Results from this type of selected SNP analysis were used in some of the consortia efforts described in the next section. In the CATSLife study, direct genotyping of two SNPs in APOE in the LTS twins was conducted via TaqMan assays. These TaqMan data have recently been used in a study of the effects of APOE variants on longitudinal cognitive performance (Reynolds et al., Reference Reynolds, Smolen, Corley, Munoz, Friedman, Rhee and Wadsworth2019). Additional targeted SNP genotyping papers using twin samples can be found in the Supplemental table of publications S2.
Genome-Wide Genotyping Arrays
Selected subjects from the CLDRC have been genotyped using the Illumina HumanOmniExpress chip, with 749 subjects included in a collaborative analysis of genetic variants influencing reading and language traits (Gialluisi et al., Reference Gialluisi, Newbury, Wilcutt, Olson, DeFries, Brandler and Fisher2014). LTS and CTS twins were included in the subjects genotyped using the Affymetrix 6.0 chip (the AFFY6 set) and analyzed by Derringer et al. (Reference Derringer, Corley, Haberstick, Young, Demmitt, Howrigan and McQueen2015) for associations with behavioral disinhibition and by Benca et al. (Reference Benca, Derringer, Corley, Young, Keller, Hewitt and Friedman2017) for associations with EF performance variation using derived risk scores. Approximately 40% of both the LTS and CTS families contributed twins to this genotyped sample, but typically only one twin per family was included. A small number of twins have also been genotyped as part of NIDA’s opioid consortium using the Smokescreen array (the SMOKESCR set). Systematic genotyping of LTS twins, done as part of the CATSLife study, is ongoing, using an Affymetrix Axiom array (the AXIOM set). To date, over 90% of families with available DNA samples have had at least one twin genotyped. Both members of DZ twin pairs are targeted, while one twin in 90% and both twins in 10% of MZ pairs are targeted for genotyping. The Affymetrix 6.0 chip and the Affymetrix Axiom Array calls have been imputed to both the 1000 Genomes Version 3 and HRC reference panels. A small degree of subject overlap exists between these three genome-wide genotyped sets.
Targeted Sequencing
Kamens et al. (Reference Kamens, Corley, Richmond, Darlington, Dowell, Hopfer and Ehringer2016) included subsets of LTS and CTS twins in estimating the contribution of low-frequency variants in nicotinic acetylcholine receptor (CHRN) genes to a measure of antisocial drug dependence. All 16 CHRN genes, including base pairs up to 5000 pairs upstream and downstream of the coding regions, were sequenced using bar-coded DNA to allow for individual read identification. Burden method tests estimated the cumulative effects of low-frequency variants on the outcome measure.
Microbiome Analysis
DNA from buccal cell saliva samples has been analyzed for bacterial oral microbiome constituents from some LTS and CTS samples. Stahringer et al. (Reference Stahringer, Clemente, Corley, Hewitt, Knights, Walters and Krauter2012) took advantage of repeated DNA collections at approximately 5-year intervals to assess stability across time and differential twin similarity between MZ and DZ twin pairs. One conclusion is that twin pairs become less similar in their microbiomes as they get older and no longer cohabitate. Demmitt et al. (Reference Demmitt, Corley, Huibregtse, Keller, Hewitt, McQueen and Krauter2017) used saliva samples from 752 LTS and CTS twin pairs to show that MZ twins show significantly less microbiome diversity within-pair than do DZ twins, supporting the idea that the human genome is influencing oral microbiomes.
Collaborations and Consortia
Data from the various twin studies have been used in a number of cross-site collaborations and consortia focused on particular phenotypes and genotypes. Generally, these multisite study extensions can be organized by whether they involve phenotype data only or both phenotypes and genotypes, and whether analyses are done at each individual site with results later combined, or whether analyses are done once using combined data sets. A brief description of some of the collaborations and consortia to which CTR twin data have contributed follows. This section is not intended to be exhaustive, but should show the breadth of the collaborative investigations to which CTR studies have contributed.
Phenotypic Only
Longitudinal data on height, weight and body mass index from the LTS and CTS samples represent part of the COllaborative project of Development of Anthropometrical measures in Twins (CODATwins) project described in this issue by Silventoinen et al. (Reference Silventoinen, Jelenkovic, Yokoyama, Sund, Sugawara, Tanaka and Kaprio2019). Much of the height and weight data were self-report or parental report, but in the LTS, at key ages including birth and years 7, 12 and 16, these variables were directly measured. Analyses were conducted from a unified, harmonized database managed in Helsinki.
Cognitive data from CLDRC, CTS and LTS twins became part of the 11,000+ twin pair sample in the Genetics of High Cognitive Abilities consortium (Haworth et al., Reference Haworth, Wright, Luciano, Martin, de Geus, van Beijsterveldt and Plomin2010). A measure of general cognitive ability created from IQ subtests was normed by age and sex in Colorado with duplicate data due to subject overlap across samples eliminated. This paper detected a significant increase in heritability for IQ from childhood to young adulthood and suggested active G–E correlation as a plausible explanation.
Longitudinal data from LTS and CTS twins on tobacco use from the CADD were included in a large mega-analysis of smoking initiation based on nearly 20,000 pairs (Developmental Genetic Epidemiology of Smoking and Health; Maes et al., Reference Maes, Prom-Wormley, Eaves, Rhee, Hewitt, Young and Neale2017). Although prevalence differed by site, the authors found a nearly linear increase in smoking initiation across sites during adolescence, and an increasing role for genetic influences later in adolescence.
Both Phenotypic and Genotypic
Reading, language and ADHD data from the CLDRC have been frequently used in multisite analyses, as seen in the Deffenbacher et al. (Reference Deffenbacher, Kenyon, Hoover, Olson, Pennington, DeFries and Smith2004) and Gialluisi et al. (Reference Gialluisi, Newbury, Wilcutt, Olson, DeFries, Brandler and Fisher2014) papers cited above, and in the references cited in Supplemental Table S1. As exemplified by Gialluisi et al. (Reference Gialluisi, Andlauer, Mirza-Schreiber, Moll, Becker, Hoffmann and Schulte-Körne2019), these collaborations have been expanded recently by participation in the GenLang consortium.
NIDA Genetic Consortium’s (NGC) Collaborative Meta-analysis of OPRM1 (Schwantes-An et al., Reference Schwantes-An, Zhang, Chen, Hartz, Culverhouse, Chen and Saccone2016) associated variation on a single genetic polymorphism (rs1799971) in OPRM1 with six measures of substance dependence, using data from the LTS and CTS samples in addition to other subjects with data collected as part of the CADD. The International Cannabis Consortium (Stringer et al., Reference Stringer, Minică, Verweij, Mbarek, Bernard, Derringer and Vink2016) focused on cannabis specifically, using genome-wide analyses of SNP data from the Affymetrix 6.0 subjects, including LTS and CTS twins, with the primary phenotype whether subjects had ever used cannabis in any form. In this study, each collaborating site conducted their own analysis following a standardized protocol and contributed their results for a combined meta-analysis. This model, in which each contributing site follows a standardized protocol for analysis, has been the basis for a number of additional collaborations involving substance use and substance use disorders. The NGC Cooperative Analysis of Opioid Dependence effort, led by Eric Johnson, is using this strategy to compile site-specific results from existing studies for which data on opioid exposure, use and dependence, including from the LTS and CTS twins genotyped on the Affymetrix 6.0 chip, have already been collected. In the GWAS (Genome-Wide Association Study) & Sequencing Consortium of Alcohol and Nicotine use (GSCAN), phenotypic and genotyping data from the Affymetrix 6.0 twins, along with other chipped subjects, were used to create sample-specific association results from an imputed SNP set, then compiled across a large number of studies to create risk scores for smoking phenotypes (Liu et al., Reference Liu, Jiang, Wedow, Li, Brazel, Chen and Vrieze2019). More than 550 CADD subjects with either lifetime cocaine or opioid dependence or both, including some twins, have been newly genotyped as a part of the NGC’s Opioid Dependence Smokescreen genotyping initiative, with genotypic and dependence data destined for the database of Genotypes and Phenotypes (dbGaP). The Affymetrix 6.0 genotypic data and substance use data on tobacco, alcohol, cannabis and other drugs are already a part of the Psychiatric Genetic Consortium’s (PGC) database on Substance Disorders (Walters et al., Reference Walters, Polimanti, Johnson, McClintick, Adams, Adkins and Agrawal2018).
In the COgnitive GENomics consorTium (COGENT), features of the genome have been used to predict general cognitive ability. Twin data taken from the AFFY6 set are part of the COGENT database (Davies et al., Reference Davies, Lam, Harris, Trampush, Luciano, Hill and Deary2018). In an example analysis, Howrigan et al. (Reference Howrigan, Simonson, Davies, Harris, Tenesa, Starr and Keller2016) used the extent of runs of homozygosity in the SNP data as a marker of potential inbreeding depression in an ethnically homogenous set of subjects of European ancestry and found evidence for a deleterious effect largely attributable to long, rare, presumably autozygous tracts.
Recently collected structural imaging data from the LTS-fMRI study are currently being processed for inclusion in Enhancing Neuro Imaging Genetics Through Meta Analysis (ENIGMA; (http://enigma.ini.usc.edu; Thompson et al., Reference Thompson, Stein, Medland, Hibar, Vasquez and Renteria2014), specifically for the working group on GWAS Meta-Analysis of Cortical Thickness and Surface Area. Site-localized analyses use genotyping results and imputed SNPs for both the Affymetrix 6.0 and Affymetrix Axiom arrays.
Several consortia efforts involving CTR twins involve a combination of genetic data and developmentally relevant measures of problem behavior assessed by parents and teachers. For EAGLE (The EArly Genetics and Lifecourse Epidemiology Consortium), imputed genotypes from the AFFY6 set and AXIOM set for subjects with Aggression and Attention subscale data from Achenbach’s CBCL (Achenbach, Reference Achenbach1991a) and TRF (Achenbach, Reference Achenbach1991) checklists were analyzed locally and summary results sent to the consortia analysts for meta-analysis. For the Total Child Psychiatric Problems GWAS consortium (Alexander Neumann, lead investigator), the maternally rated total problems score from the CBCL was the outcome measure, and genotypes were limited to the AFFY6 set. One additional developmentally oriented consortium involves toddler cognition as measured by the Bayley’s Mental Developmental Index. A small number of LTS subjects with AFFY6 genotypic data were included in this Bayley’s Toddler Score GWAS Meta-analysis project (Tarun Ahluwalia & Lærke Sass, Investigators). And finally, the DEVelopmental CONsortium (DEVCON; Scott Vrieze & Michael Stallings, Lead investigators) is in its initial stages of organizing and harmonizing developmental data from multiple longitudinal studies with both genetic and phenotypic data, including both LTS and CTS.
Discussion
The CTR has proven its worth many times over during its nearly 40 years of operation. Samples and studies using the CTR have covered a wide range of research topics, and the ability to maintain continuing subject participation in longitudinal studies is a tribute to the dedicated teams of investigators, coordinators, research staff and support staff that have made subjects feel welcomed and appreciated. And to the participants in these studies, who so generously made room in their busy lives to continue their involvement, the IBG and its collaborators owe heartfelt thanks. The CDH has maintained its ability to provide the CTR with contact information for a high percentage of the twin families giving birth in the state of Colorado, and family participation remains strong when contacted. The large number of collaborations and consortia to which CTR twin data have contributed speaks to the relevance of the phenotypic and genotypic data collected from the CTR studies, and the importance of phenotypic data that cover multiple developmental periods, from the first years of life to mid-life. There are some issues with twin research based in Colorado that are recognized and make collaborations with other investigators in other locations even more important. Examples include that Colorado was one of the first states to legalize recreational marijuana, has a high concentration of subjects along its urbanized Front Range, has a relatively low proportion of ethnic minorities, has higher than national average educational and activity levels and has experienced a high degree of both in-migration and out-migration in its recent past. But, as shown in Figure 2, even when participants have relocated outside Colorado, they may still be willing to participate in studies in which they feel invested. We hope the proven track record of the CTR, and the diverse backgrounds, expertise and willingness to collaborate of the investigators associated with the CTR will continue to make it a valuable resource for future research and potential expanded twin-based designs.
Acknowledgments
The authors gratefully acknowledge support from the National Institutes of Health, NIH AG046938 [MPIs, Chandra A. Reynolds, Sally J. Wadsworth] for the preparation of this manuscript. The content of this manuscript is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. Our thanks to Kyle Gebelin for the preparation of Figure 2 and to John DeFries for his assistance with Supplementary Table S1. We would also like to acknowledge the continuing support for the maintenance and updating of the CTR from the CDH, Division of Vital Statistics. Support for the CTR across the years has come from a CRCW grant from the University of Colorado, NIH grants HD19802, HD010333, HD18426, MH43899, AA023974, DA015131, DA011015, DA041120 and HD027802; the John D. and Catherine T. MacArthur Foundation; and Veterans Administration grant 1296.07.1629B. The founders and directors were R. Plomin, J. C. DeFries, R. N. Emde, D. W. Fulker, J. K. Hewitt and R. K. Olson. LTS and CTS participation in the Center for Antisocial Drug Dependence was funded by DA011015 (Michael Stallings, Component PI). LTS participation in the Executive Functions/fMRI study is funded by MH063207 (Naomi Friedman, PI). LTS participation in CATSLife is funded by AG046938 (Chandra Reynolds & Sally Wadsworth, MPIs). LTS and CTS participation in COMN is funded by DA042755 (John Hewitt, Colorado PI), CTS participation in Executive Function/fMRI was funded by DA011015 (Thomas Crowley, Component PI). CTS participation in the fMRI study of pain is funded by DA046064 (Tor Wager, PI). ERDS was funded by HD038526 (Erik Willcutt, PI). CLDRC is funded by HD027802 (Erik Willcutt, PI). CoTwins is funded by AA023974 and U01DA046413 (Scott Vrieze, PI). The ABCD twin hub is funded by DA041120 (Marie Banich & John Hewitt, Colorado PIs). Most importantly, we would like to acknowledge the many Colorado twin families who have so graciously given their time to the studies associated with the CTR. We thank for your continuing interest and participation!
Supplementary material
To view supplementary material for this article, please visit https://doi.org/10.1017/thg.2019.50.








