Introduction
Salmonella spp. are Gram-negative bacilli from the family Enterobacteriaceae that are capable of colonizing the intestinal tract of most vertebrates. Non-typhoidal Salmonella are important food-borne pathogens that cause gastroenteritis, bacteraemia and focal infections in human and animals [Reference Hohmann1]. Transmission of salmonellae to human typically occurs by ingesting meat, dairy products and other food contaminated by animal faeces, or by cross-contamination from food contaminated with salmonellae. Zoonotic transmission of non-typhoidal Salmonella can also occur by direct exposure to the faeces of reptiles, pets and other animals [2–Reference Holmberg, Wells and Cohen6].
Several outbreaks of salmonellosis in human related to exposure to contaminated dog food and dog treat products have been reported. In 1999, laboratory and epidemiological investigations identified pig ear-based dog treats as a source of Salmonella enterica subsp. enterica serovar Infantis infection in human in Canada [Reference Clark7]. As a consequence of the Canadian outbreak, the Food and Drug Administration Center for Veterinary Medicine (FDA CVM) in the USA performed a retail sampling study investigating the prevalence of Salmonella in pet treats available in the US pet stores. Therein, 158 pet treats were collected, of which 41% were contaminated with Salmonella [Reference White8]. Twenty-four serotypes were identified, including S. Anatum, S. Typhimurium and S. Infantis. Of these, 36% were resistant to at least one antimicrobial, whereas 13% were resistant to four or more antimicrobials [Reference White8]. In 2002, 2004, 2005 and 2013, some human infections of Salmonella were attributed to pet treats in Canada and the USA [Reference Pitout9–Reference Cavallo11]. A human infection in Canada in 2002 was caused by CMY-2 AmpC β-lactamase-producing S. Newport strains [Reference Pitout9]. In many countries, the incidence of human infections caused by extended-spectrum cephalosporin-resistant Salmonella has increased dramatically [Reference Su12–Reference Harrois18]. In Japan, extended-spectrum cephalosporin-resistant Salmonella harbouring AmpC or extended-spectrum β-lactamase (ESBL) genes, such as bla CTX−M−14 and bla CTX−M−15, have been isolated from human [Reference Izumiya19, Reference Morita20].
As no routine surveillance of dog treats for Salmonella contamination is performed in Japan, the main objective of the current study was to determine the current prevalence of Salmonella contamination in such dog treats. The second objective was to investigate the prevalence of β-lactam resistance among Salmonella from dog treats in Japan using a molecular approach to detect ESBL and AmpC β-lactamase genes.
Methods
Salmonella isolation and identification
A total of 303 product samples were collected, consisting of domestic products (n = 255) and imported products (n = 48). It was estimated that 300 samples would provide a 95% probability that at least one sample would be positive for Salmonella, assuming a minimum prevalence of 1%. All samples were collected in the Okayama and Osaka Prefectures in Japan from April 2016 to December 2016. The main pet supply chain from which we obtained our samples carried a variety of brands of imported and domestically packaged products. Other miscellaneous brands sold in supermarkets and in a chain store selling a large variety of products were purchased off the shelves. Prior to each sampling day, three stores were selected randomly in the scheduled city. We purchased 10 samples randomly at one store. We purchased three samples at another store, as only three were available. Other stores sold more than 30 kinds of dog treats. The samples were transported to the laboratory and kept at ambient temperature until analysis. Salmonella were isolated following the procedure of the US FDA Bacteriological Analytical Manual [Reference Wallace21]. The isolates were identified using API 20E identification kits (bioMerieux, l'Etoile, France) and were serotyped by using slide and tube agglutination tests with commercially available antisera (Denka Seiken Co., Ltd., Tokyo, Japan). In addition, polymerase chain reaction (PCR) was used to serotype the Salmonella isolates [Reference Alvarez22].
Antimicrobial susceptibility testing
Escherichia coli ATCC 25922 was used as the quality-control strain in the experiments. The minimum inhibitory concentrations (MICs) of the following drugs were determined using the microbroth dilution method on Eiken dry plates (Eiken Kagaku Co., Ltd., Tokyo, Japan): ampicillin (ABPC), cefazolin (CEZ), cefotaxime (CTX), chloramphenicol (CP), tetracycline (TC), gentamycin (GM), kanamycin (KM), nalidixic acid (NA), ciprofloxacin (CPFX) and trimethoprim (TMP). MIC breakpoints were interpreted according to the Clinical and Laboratory Standards Institute guidelines [23]. Susceptibility to streptomycin (SM) was determined by using the standard disk diffusion method [23] with Sensi-Discs (Japan Becton Dickinson Company, Tokyo, Japan). Isolates resistant to CTX were also tested for ESBL production, using a phenotypic confirmatory test [23].
Detection of antimicrobial resistance genes
All DNA templates for analysis were prepared by the boiling method, as described elsewhere [Reference Matayoshi24]. Briefly, bacterial cells were suspended in 200 µl of distilled water and boiled for 10 min. The cells were then pelleted by centrifugation for 1 min. The supernatants (5 µl) were used for PCR to detect the presence of the class 1 and class 2 integron genes and antimicrobial resistance genes. PCR was performed in a final volume of 25 µl using GoTaq® Green master mix 2× (Promega, Madison, WI, USA), according to the manufacturer's instructions; primers are listed in Table 1.
Table 1. Primer sequences and expected PCR product sizes
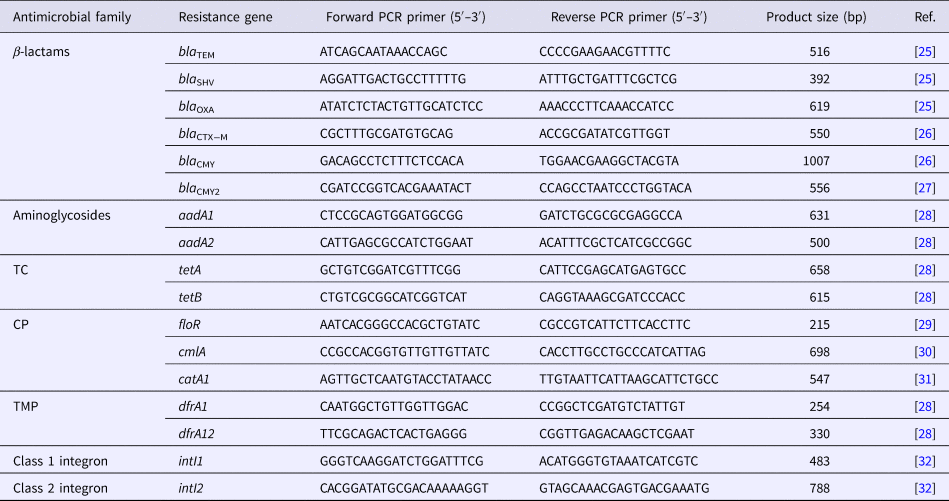
TC, tetracycline; CP, chloramphenicol; TMP, trimethoprim
The resistant isolates were screened for the presence of 15 resistance genes corresponding to their resistance phenotypes. β-lactam antibiotic-resistant isolates (n = 3) were screened for the presence of the six β-lactamase genes (bla TEM, bla SHV, bla OXA, bla CTX−M, bla CMY and bla CMY2) [Reference Colom25–Reference Dahshan27]. Aminoglycoside-resistant isolates (n = 3) were screened for the presence of the aadA1 and aadA2 genes [Reference Chuanchuen and Padungtod28]. TC-resistant isolates (n = 3) were screened for the presence of the tetA and tetB genes [Reference Chuanchuen and Padungtod28]. CP-resistant isolates (n = 2) were screened for the presence of the floR, cmlA and catA1 genes [Reference Bolton29–Reference Maynard31]. TMP-resistant isolates (n = 4) were screened for the presence of the dfrA1 and dfrA12 genes [Reference Chuanchuen and Padungtod28]. All resistant isolates (n = 7) were screened for the presence of the intI1 and intI2 genes [Reference Sáenz32].
Statistical methods
For all prevalence estimates, we calculated 95% confidence intervals using the Wilson score interval method [Reference Wilson33]. Fisher's exact test was used to calculate the statistical differences between the prevalence of Salmonella in the imported and domestic dog treat samples.
Results
Isolation of Salmonella and identification of isolate serotypes
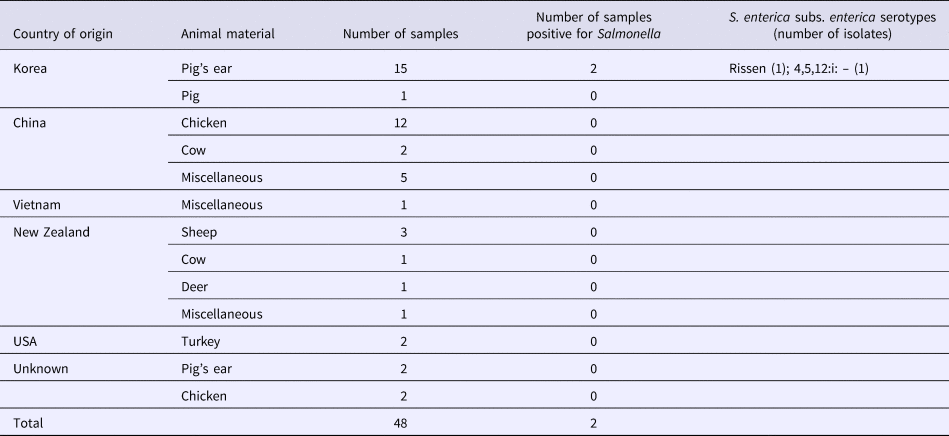
S. enterica subspecies enterica was isolated from seven (2.3%; 95% CI 1.1–4.7) of 303 dog treat samples, including five (2.0%; 95% CI 0.9–4.6) domestic and two (4.2%; 95% CI 1.2–14.0) imported products. There was no significant difference in the contamination levels between domestic and imported products. Three of these isolates represented serovar 4,5,12:i:–, two were serovar Rissen, and two were serovar Thompson. Five isolates were found in the domestic dog treats made from chicken, pig's ear and cow (Table 2). Two isolates were found in dog treats imported from Korea, made from pig's ear (Table 3).
Table 2. Salmonella isolated from domestic dog treats
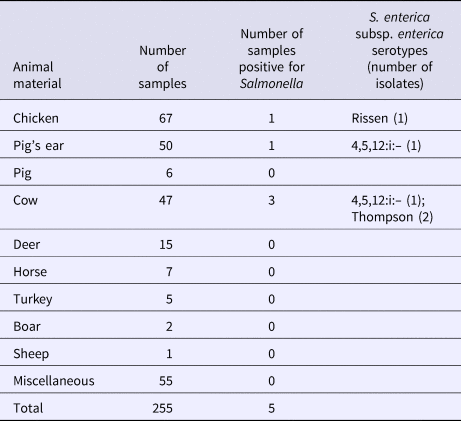
Table 3. Salmonella isolated from imported dog treats
Antimicrobial susceptibility profiles of the isolates
All Salmonella isolates were susceptible to CTX, CPFX and NA. Two isolates (serovar Rissen) (29% of all isolates) was fully susceptible to all tested antimicrobial agents (Tables 4 and 5). Five isolates were resistant to one or more drugs, including Salmonella serovars 4,5,12:i:– (n = 3) and Thompson (n = 2). Three isolates representing serovar 4,5,12:i:– were resistant to four and more antimicrobials. Because all Salmonella isolates were susceptible to CTX, ESBL production was not pursued.
Table 4. MICs for Salmonella isolates
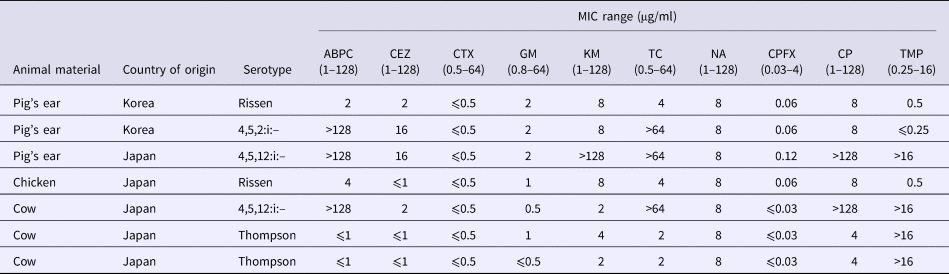
ABPC, ampicillin; CEZ, cefazolin; CTX, cefotaxime; GM, gentamycin; KM, kanamycin; TC, tetracycline; NA, nalidixic acid; CPFX, ciprofloxacin; CP, chloramphenicol; TMP, trimethoprim.
Table 5. Summary of the Salmonella isolate resistance profiles
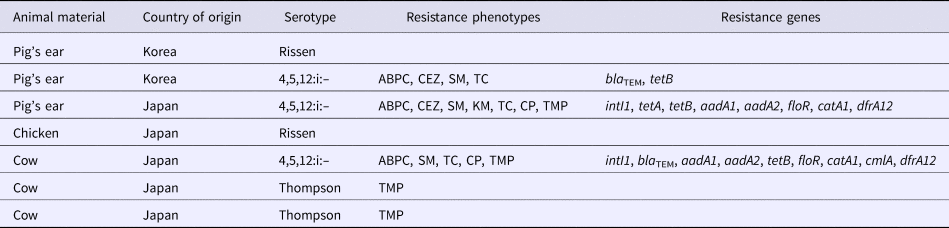
ABPC, ampicillin; CEZ, cefazolin; SM, streptomycin; KM, kanamycin; TC, tetracycline; CP, chloramphenicol; TMP, trimethoprim
Detection of antimicrobial resistance genes harboured by the isolates
PCR screening of the Salmonella isolates for the presence of integron genes revealed that two isolates were positive for the class 1 integron gene and none were positive for the class 2 integron gene (Table 5). The distribution of the various resistance genes in the isolates is shown in Table 5. Two of the three ABPC-resistant isolates contained the bla TEM gene; none of the ABPC-resistant isolates contained any other β-lactamase genes. The tetB gene was detected in all (three) TC-resistant isolates. The floR and catA1 genes were detected in all (two) CP-resistant isolates. The aadA1 and aadA2 genes were detected in two out of the three SM-resistant isolates. The dfrA12 gene was detected in two out of the four TMP-resistant isolates.
Discussion
In the current study, we showed that dog treats in Japan might harbour Salmonella. The determined prevalence of Salmonella in dog treats from Japan was 2%. Although the difference was not significant, the prevalence of Salmonella was slightly higher for imported treats than for domestic treats. A survey performed in the UK by Willis identified Salmonella in 7.8% imported dog chews [Reference Willis34]. In this aforementioned study, samples imported from Asia or South Africa were the main target of the survey. A total of 2369 dog chew samples imported from Thailand, China, India, Sri Lanka, Argentina, Brazil, Colombia and the USA were examined at public health laboratories in Ashford, Chelmsford, Ipswich, London and Southampton. Overall, Salmonella species were detected in 184 samples (7.8%). Wong et al. found Salmonella in 6.7% dog chews in New Zealand [Reference Wong35]. The authors collected 600 samples, consisting of New Zealand-produced (domestic) and imported samples. Most samples were purchased from two stores of a major pet supply chain in Christchurch and by direct mail order from a major supplier in Wellington, New Zealand; there was no significant difference in the contamination levels between imported and domestic samples. The prevalence of Salmonella identified in the current study was lower than that of previous reports. For instance, Clark et al. reported that 51% of retail Canadian pig ear treats collected from Alberta, Saskatchewan, Ontario, Québec, Newfoundland and Nova Scotia were positive for Salmonella [Reference Clark7]. White et al. found 41% of retail dog treats collected in the USA by 16 district offices and seven regional laboratories of the FDA to be positive for Salmonella in the USA [Reference White8]. Li et al. reported that the prevalence of Salmonella in pet food or pet treats collected under the FDA CVM Feed Contaminants Program in the years 2007–2009 was significantly (P < 0.05) lower than that reported for the years 2002–2006 [Reference Li36]. It is thought that this reduction of Salmonella prevalence is associated with the various countermeasures undertaken by each country. The American Feed Industry Association [37] and the European Pet Food Industry Federation [38] have developed guidelines for the manufacturing of pet products. In addition, to monitor the trend of Salmonella contamination in animal feed, since 2002, the FDA CVM has established a Salmonella surveillance programme, which includes dog treats, in the USA. Subsequently, the FDA CVM has established a second surveillance programme, including dog treats, in the USA. In Japan, the Law for Ensuring the Safety of Pet Food came into force in 2009, and the Ministry of Agriculture, Forestry, and Fisheries published a proper manufacturing manual for pet food (including dog treats) in 2014. Nevertheless, the Japanese government have not carried out any Salmonella surveillance programmes that include dog treats. Since Salmonella contamination was indeed detected in dog treats in the current study, the government of Japan should take stronger measures to counteract the possible associated health threat. It is not known how many dog treats are sold in Japan. Neither is it known how many companies are manufacturing dog treats in Japan, as the government of Japan have not published a report with these data. However, the government of Canada have published a report about dog treats in Japan, which estimated that retail sales of dog treats and mixers would reach US$553.9 million in 2016 [39]. However, the figures described in this report were not examined by the government itself but were estimated by a market research company. Therefore, we could not verify the accuracy of the content. Thus, the government of Japan should make efforts to determine the distribution volume and sales of dog treats in Japan, which would help determine sample size in future studies of this nature.
In the current study, the Salmonella-positive dog treats originated from Japan and Korea. However, we also found dog treats of unknown country of origin. In Japan, the Law for Ensuring the Safety of Pet Food states that sellers of pet food, including dog treats, must label the content, country of origin, expiration date, materials, location, company name and company location on the pet food products.
An increasing incidence of multidrug-resistant Salmonella has been widely reported in the past and is presumably attributed to the extensive use of antimicrobial agents in human and veterinary medicine [Reference Threlfall, Rowe and Ward40]. In a US study, White et al. found that Salmonella isolated from dog treats harboured class 1 integrons [Reference White8]. In Japan, Futagawa-Saito et al. reported that the rates of antimicrobial resistance among faecal isolates from healthy pigs obtained in the years 2004–2005 were significantly higher than those of isolates from the years 1998–1999 [Reference Futagawa-Saito41]. In the current study, we detected the bla TEM gene but not the bla CTX−M, bla CMY or bla CMY2 genes in Salmonella isolates from dog treats in Japan. More extensive monitoring of dog treats must be undertaken as part of surveillance of multidrug-resistant Salmonella.
In conclusion, a small percentage of dog treats in Japan are contaminated with Salmonella, including antimicrobial-resistant isolates. In the USA, an outbreak of Salmonella Typhimurium occurred in humans that had been exposed to dog treats [Reference Cavallo11]. Therefore, care should be taken when handling dog treats. It is recommended that people wash their hands after feeding dogs and after any contact with dog treats [Reference Kukanich42]. Veterinarians have the responsibility to disseminate accurate information about the potential contamination risks, so that appropriate precautions can be implemented. In Japan, there are currently no reports of human salmonellosis caused by dog treats. Although the risk of salmonellosis from contaminated dog treats may be low, adhering to safety recommendations will help to minimise the risk of infections with Salmonella in dogs and family members.
Author ORCIDs
S. Yukawa, 0000-0002-4453-5029.
Acknowledgements
We would like to thank Editage (http://www.editage.com) for editing and reviewing this manuscript for English language.
Financial support
This research received no specific grant from any funding agency, commercial or not-for-profit sectors.
Conflict of interest
None.









