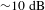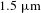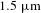7 results
Brightness enhancement on a narrow linewidth FBG-based MOPA fiber laser
-
- Journal:
- High Power Laser Science and Engineering / Accepted manuscript
- Published online by Cambridge University Press:
- 09 May 2024, pp. 1-24
-
- Article
-
- You have access
- Open access
- Export citation
A 110 W fiber gas Raman laser at 1153 nm
-
- Journal:
- High Power Laser Science and Engineering / Volume 11 / 2023
- Published online by Cambridge University Press:
- 28 February 2023, e10
-
- Article
-
- You have access
- Open access
- HTML
- Export citation
Automated Analysis of Grain Growth Under in-situ Irradiation Using Convolutional Neural Network
-
- Journal:
- Microscopy and Microanalysis / Volume 28 / Issue S1 / August 2022
- Published online by Cambridge University Press:
- 22 July 2022, pp. 2036-2037
- Print publication:
- August 2022
-
- Article
-
- You have access
- Export citation
Mitigation of stimulated Raman scattering in kilowatt-level diode-pumped fiber amplifiers with chirped and tilted fiber Bragg gratings
-
- Journal:
- High Power Laser Science and Engineering / Volume 7 / 2019
- Published online by Cambridge University Press:
- 18 March 2019, e18
-
- Article
-
- You have access
- Open access
- HTML
- Export citation
10 watt-level tunable narrow linewidth
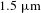 $1.5~\unicode[STIX]{x03BC}\text{m}$ all-fiber amplifier
$1.5~\unicode[STIX]{x03BC}\text{m}$ all-fiber amplifier
- Part of
-
- Journal:
- High Power Laser Science and Engineering / Volume 6 / 2018
- Published online by Cambridge University Press:
- 06 June 2018, e33
-
- Article
-
- You have access
- Open access
- HTML
- Export citation
18 - Inducible RNAi as a forward genetic tool in Trypanosoma brucei
-
-
- Book:
- RNA Interference Technology
- Published online:
- 31 July 2009
- Print publication:
- 17 January 2005, pp 247-256
-
- Chapter
- Export citation