Health literacy is ‘the degree to which individuals have the capacity to obtain, process, and understand basic health information and services needed to make appropriate health decisions’ (Institute of Medicine, 2004). This capacity is thought to be important for navigating all aspects of health care, including the ability to seek out and act on appropriate health information and self-manage health conditions (Baker, Reference Baker2006; Institute of Medicine, 2004). Tests of functional health literacy have been used to investigate the association between health literacy and health. Individuals with lower health literacy have been found to be less likely to take part in health-promoting behaviors (von Wagner et al., Reference von Wagner, Knight, Steptoe and Wardle2007). Lower health literacy is associated with poorer overall health status (Berkman et al., Reference Berkman, Sheridan, Donahue, Halpern and Crotty2011), lower self-reported physical and mental health (von Wagner et al., Reference von Wagner, Knight, Steptoe and Wardle2007; Wolf et al., Reference Wolf, Gazmararian and Baker2005) and greater self-reported depressive symptoms (Gazmararian et al., Reference Gazmararian, Baker, Parker and Blazer2000). One study (Wolf et al., Reference Wolf, Gazmararian and Baker2005) found that individuals with inadequate health literacy were 48% more likely to report a diagnosis of diabetes and 69% more likely to report having heart failure, compared to those with adequate health literacy, after adjusting for sociodemographic variables and health behaviors. Using prospective studies, lower health literacy predicted incident dementia (Kaup et al., Reference Kaup, Simonsick, Harris, Satterfield, Metti, Ayonayon and Yaffe2014; Yu et al., Reference Yu, Wilson, Schneider, Bennett and Boyle2017) and risk of dying (Baker et al., Reference Baker, Wolf, Feinglass, Thompson, Gazmararian and Huang2007; Berkman et al., Reference Berkman, Sheridan, Donahue, Halpern and Crotty2011; Bostock & Steptoe, Reference Bostock and Steptoe2012).
Compared with those of health literacy, similar associations with health have been found for cognitive function. Individuals with higher cognitive function tend to participant more in health-promoting behaviors (Mons et al., Reference Mons, Schottker, Muller, Kliegel and Brenner2013; Richards et al., Reference Richards, Jarvis, Thompson and Wadsworth2003; Taylor et al., Reference Taylor, Hart, Smith, Starr, Hole, Whalley and Deary2003; Wraw et al., Reference Wraw, Der, Gale and Deary2018). Vascular risk factors, including diabetes and hypertension, have been associated with poorer cognitive function and greater cognitive decline (Knopman et al., Reference Knopman, Mosley, Catellier and Coker2009; Mõttus et al., Reference Mõttus, Luciano, Starr and Deary2013; Pavlik et al., Reference Pavlik, Hyman and Doody2005; Singh-Manoux & Marmot, Reference Singh-Manoux and Marmot2005). Cognitive function, measured early in life, has been found to predict later life physical functioning and health status (Wraw et al., Reference Wraw, Deary, Gale and Der2015), psychological distress (Gale et al., Reference Gale, Hatch, Batty and Deary2009), psychiatric illness (Batty et al., Reference Batty, Mortensen and Osler2005; Dickson et al., Reference Dickson, Laurens, Cullen and Hodgins2012; Gale et al., Reference Gale, Deary, Boyle, Barefoot, Mortensen and Batty2008; Scult et al., Reference Scult, Paulli, Mazure, Moffitt, Hariri and Strauman2017; Zammit et al., Reference Zammit, Allebeck, David, Dalman, Hemmingsson, Lundberg and Lewis2004), dementia (McGurn et al., Reference McGurn, Deary and Starr2008), and death (Batty et al., Reference Batty, Deary and Gottfredson2007; Calvin et al., Reference Calvin, Deary, Fenton, Roberts, Der, Leckenby and Batty2011, Reference Calvin, Batty, Der, Brett, Taylor, Pattie and Deary2017; Christensen et al., Reference Christensen, Mortensen, Christensen and Osier2016).
Performance on tests of health literacy and cognitive function are moderately to highly correlated (Boyle et al., Reference Boyle, Yu, Wilson, Segawa, Buchman and Bennett2013; Mõttus et al., Reference Mõttus, Johnson, Murray, Wolf, Starr and Deary2014; Murray et al., Reference Murray, Johnson, Wolf and Deary2011; Reeve & Basalik, Reference Reeve and Basalik2014). Murray et al. (Reference Murray, Johnson, Wolf and Deary2011) found that the correlations between general cognitive ability and three tests of health literacy, tested in older adulthood, ranged from .35 to .53 (p < .001). Given these correlations, researchers have sought to determine whether the relationship between health literacy and health remained when also accounting for cognitive function. Cognitive function has been consistently found to attenuate the size of the association between health literacy and health; however, whereas some studies have found that health literacy no longer predicted better health when controlling for cognitive function (Fawns-Ritchie et al., Reference Fawns-Ritchie, Starr and Deary2018b; Mõttus et al., Reference Mõttus, Johnson, Murray, Wolf, Starr and Deary2014; O’Conor et al., Reference O’Conor, Wolf, Smith, Martynenko, Vicencio, Sano and Federman2015; Serper et al., Reference Serper, Patzer, Curtis, Smith, O’Conor, Baker and Wolf2014), others have found that a small but significant association remained between higher health literacy scores and better health when also controlling for cognitive function (Baker et al., Reference Baker, Wolf, Feinglass and Thompson2008; Bostock & Steptoe, Reference Bostock and Steptoe2012; Fawns-Ritchie et al., Reference Fawns-Ritchie, Starr and Deary2018a; Mõttus et al., Reference Mõttus, Johnson, Murray, Wolf, Starr and Deary2014; Yu et al., Reference Yu, Wilson, Schneider, Bennett and Boyle2017).
Whereas there is a wealth of evidence reporting a relationship between health literacy, cognitive function and health, it is less well understood why these associations are found. One possibility is that they share genetic influences. Cognitive function is substantially heritable (Deary et al., Reference Deary, Harris and Hill2019; Haworth et al., Reference Haworth, Wright, Luciano, Martin, de Geus, van Beijsterveldt and Plomin2010; Plomin & Deary, Reference Plomin and Deary2015). With increasing samples sizes, the specific genetic variants associated with cognitive function are being identified (Davies et al., Reference Davies, Lam, Harris, Trampush, Luciano, Hill and Deary2018; Savage et al., Reference Savage, Jansen, Stringer, Watanabe, Bryois, de Leeuw and Posthuma2018). One study (Hagenaars et al., Reference Hagenaars, Harris, Davies, Hill, Liewald, Ritchie and Deary2016) sought to explore the shared genetic architecture between cognitive function and health, using two complementary genetic techniques: linkage disequilibrium (LD) score regression (Bulik-Sullivan et al., Reference Bulik-Sullivan, Loh, Finucane, Ripke, Yang and Patterson2015) and polygenic profile scoring (Purcell et al., Reference Purcell, Wray, Stone, Visscher, O’Donovan, Sullivan and Sklar2009). The first technique involves calculating the genetic correlations between two traits of interest using summary results from previous genome-wide association studies (GWAS). The second technique uses summary GWAS data for a specific trait (e.g., type 2 diabetes) and tests whether the genetic variants found to be associated with this trait are also associated with the same (e.g., type 2 diabetes) or a different (e.g., cognitive function) phenotype in an independent sample. Using these techniques, Hagenaars et al. (Reference Hagenaars, Harris, Davies, Hill, Liewald, Ritchie and Deary2016) found substantial shared genetic influences between cognitive function and physical and mental health. Negative genetic correlations were found between a test of verbal-numerical reasoning and Alzheimer’s disease (r g = −0.39, p = .002), and schizophrenia (r g = −0.30, p = 3.5 × 10-11), among others. Polygenic profiles for various mental and physical health-related variables were associated with performance on tests of cognitive function, including coronary artery disease, Alzheimer’s disease and schizophrenia. The shared genetic architecture between cognitive function and health has been subsequently replicated using larger samples (Davies et al., Reference Davies, Lam, Harris, Trampush, Luciano, Hill and Deary2018).
Summaries are available regarding the advances made in understanding the genetic architecture of cognitive function and its overlap with physical and mental health (Deary et al., Reference Deary, Harris and Hill2019; Hill, Harris et al., Reference Hill, Harris and Deary2019). However, to the best of our knowledge, no one has investigated the genetic contributions to people’s differences in health literacy. The aim of the present study was to explore the genetic contributions to health literacy and its overlap with cognitive function and health. Using data from the English Longitudinal Study of Ageing (ELSA), a sample of English adults aged 50 years and older, the present study conducted a GWAS of health literacy, estimated its single nucleotide polymorphism (SNP)-based heritability, and used polygenic profile scoring to examine the genetic overlap between health literacy and cognitive function, and health literacy and various health-related traits.
Materials and Methods
Participants
This study used data from ELSA (https://www.elsa-project.ac.uk/), a cohort study designed to be representative of English adults aged 50 years and older (Steptoe et al., Reference Steptoe, Breeze, Banks and Nazroo2013). The original sample (wave 1) was recruited in 2002–2003 and consisted of 11,391 participants. Participants have been followed up every 2 years and the sample has been refreshed at subsequent waves to ensure the sample’s representativeness. Interviews took place via computer-assisted personal interviews and self-completion questionnaires in the participants’ own homes. The topics assessed included health, financial and social circumstances. A nurse visit was carried out every second wave to measure biomarkers. Blood samples collected during the nurse visit have been used to genotype ELSA participants. More information on the ELSA sampling procedures are reported elsewhere (Steptoe et al., Reference Steptoe, Breeze, Banks and Nazroo2013). For the present study, a subsample of participants was used who completed the health literacy test at wave 2 (2004–2005) and who had genome-wide genotyping data (n = 5783).
Procedure
Health literacy
Health literacy was measured using a four-item reading and comprehension test. This test was designed to mimic written materials, such as drug labels, that would be encountered in a health-care setting. A piece of paper containing instructions for an over-the-counter packet of medicine was given to participants. Participants were asked four questions about the information on the medicine packet (e.g., ‘What is the maximum number of days you may take this medication?’). One point was awarded for each correct answer. As has been done in other studies (Gale et al., Reference Gale, Deary, Wardle, Zaninotto and Batty2015; Kobayashi et al., Reference Kobayashi, Wardle and von Wagner2014), participants were categorized as having adequate (4/4 questions correct) or limited (<4 correct) health literacy.
Genotyping and quality control
A total of 7597 ELSA participants who had provided blood samples were genotyped in two batches (batch 1, n = 5652; batch 2, n = 1945) by UCL Genomics using the Illumina Omni 2.5-8 chip. Quality control procedures were performed by UCL Genomics and by the present authors. This included removal of SNPs based on call rate, minor allele frequency (MAF) and deviation from Hardy–Weinberg equilibrium. Individuals were removed based on call rate, relatedness, gender mismatch and non-Caucasian ancestry. A sample of 7358 participants remained following quality control procedures.
Imputation
Prephasing and imputation to the 1000 Genome Phase 3 reference panel (Altshuler et al., Reference Altshuler, Durbin, Abecasis, Bentley, Chakravarti and Clark2015) was performed using the Sanger Imputation Service (McCarthy et al., Reference McCarthy, Das, Kretzschmar, Delaneau, Wood and Teumer2016), EAGLE2 (v2.0.5) (Loh et al., Reference Loh, Danecek, Palamara, Fuchsberger, Reshef, Finucane and Price2016) and PBWT (Durbin, Reference Durbin2014) pipeline.
Curation of summary results from GWAS of cognitive and health-related traits
Summary results from 21 GWAS of cognitive function, general health status variables, chronic diseases, health behaviors, neuropsychiatric disorders, years of schooling, social deprivation and the personality traits of conscientiousness and neuroticism were collected. For each trait, we checked the samples used in the GWAS to ensure ELSA was not included. Sources of summary statistics and key references are given in the Supplementary materials and Supplementary Table S1.
Statistical Analyses
Genome-wide association analyses
SNP-based association analyses were performed using the BGENIE v1.2 analysis package (https://jmarchini.org/bgenie/). A linear SNP association model was tested which accounted for genotype uncertainty. Prior to these analyses, the health literacy phenotype was adjusted for the following covariates: age, sex and 15 genetic principal components.
Genomic risk loci characterization using FUMA
Genomic risk loci were defined from the SNP-based association results, using FUnctional Mapping and Annotation of genetic associations (FUMA; Watanabe et al., Reference Watanabe, Taskesen, van Bochoven and Posthuma2017). First, independent significant SNPs were identified using the SNP2GENE function and defined as SNPs with a p value of < 1 × 10-5 and independent of other genome-wide suggestive SNPs at r 2 < .6. Using these independent significant SNPs, tagged SNPs to be used in subsequent annotations were identified as all SNPs that had an MAF ≥ 0.0005 and were in LD of r 2 ≥ .6 with at least one of the independent significant SNPs. These tagged SNPs included those from the 1000 Genomes Phase 3 reference panel and need not have been included in the GWAS performed in the current study. Genomic risk loci that were 250 kb or closer were merged into a single locus. Lead SNPs were also identified using the independent significant SNPs and were defined as those that were independent from each other at r 2 < .1.
Comparison to previous findings
A look-up of the independent significant and tagged SNPs for health literacy in the current study was performed in previous GWAS of general cognitive ability (Davies et al., Reference Davies, Lam, Harris, Trampush, Luciano, Hill and Deary2018) and years of schooling (Okbay et al., Reference Okbay, Beauchamp, Fontana, Lee, Pers, Rietveld and Benjamin2016). We identified whether significant SNPs and tagged SNPs reported here reached either genome-wide (p < 5 × 10-8) or suggestive (p < 1 × 10-5) significance in these previous GWAS.
Gene-based analysis implemented in FUMA
Gene-based association analyses were conducted using MAGMA (Multi-marker Analysis of GenoMic Annotation; de Leeuw et al., Reference de Leeuw, Mooij, Heskes and Posthuma2015). The test carried out using MAGMA, as implemented in FUMA, was the default SNP-wise test using the mean χ2 statistic derived on a per gene basis. SNPs were mapped to genes based on genomic location. All SNPs that were located within the gene body were used to derive a p-value describing the association found with health literacy. The SNP-wise model from MAGMA was used and the NCBI build 37 was used to determine the location and boundaries of 18,199 autosomal genes. LD within and between each gene was gauged using the 1000 genomes Phase 3 release. A Bonferroni correction was applied to control for multiple testing across 18,199 genes; the genome-wide significance threshold was p < 2.75 × 10−6.
Functional annotation implemented in FUMA
The independent significant SNPs and those in LD with the independent significant SNPs were annotated for functional consequences on gene functions using ANNOVAR (Wang et al., Reference Wang, Li and Hakonarson2010) and the Ensembl genes build 85. Functionally annotated SNPs were then mapped to genes based on physical position on the genome and chromatin interaction mapping (all tissues). Intergenic SNPs were mapped to the two closest up- and downstream genes, which can result in their being assigned to multiple genes.
Gene-set analysis implemented in FUMA
In order to test whether the polygenic signal measured in the GWAS clustered in specific biological pathways, a competitive gene-set analysis was performed. Gene-set analysis was conducted in MAGMA (de Leeuw et al., Reference de Leeuw, Mooij, Heskes and Posthuma2015) using competitive testing, which examines whether genes within the gene-set are more strongly associated with health literacy than other genes. A total of 10,675 gene-sets, sourced from Gene Ontology (Ashburner et al., Reference Ashburner, Ball, Blake, Botstein, Butler and Cherry2000), Reactome (Fabregat et al., Reference Fabregat, Sidiropoulos, Garapati, Gillespie, Hausmann, Haw and D’Eustachio2016) and SigDB (Subramanian et al., Reference Subramanian, Tamayo, Mootha, Mukherjee, Ebert, Gillette and Mesirov2005) were examined for enrichment of health literacy. A Bonferroni correction (p < .05/10,675 = 4.68 × 10-6) was applied to control for the multiple tests performed.
Gene-property analysis implemented in FUMA
A gene-property analysis was conducted using MAGMA in order to indicate the role of particular tissue types that influence differences in health literacy. The goal of this analysis was to test if, in 30 broad tissue types and 53 specific tissues, tissue-specific differential expression levels were predictive of the association of a gene with health literacy. Tissue types were taken from the GTEx v6 RNA-seq database (Ardlie et al., Reference Ardlie, DeLuca, Segre, Sullivan, Young and Gelfand2015) with expression values being log2 transformed with a pseudocount of 1 after winsorising at 50, with the average expression value being taken from each tissue. Multiple testing was controlled for using a Bonferroni correction (p < .05/53 = 9.43 × 10-4).
Estimation of SNP-based heritability
The proportion of variance explained by all common SNPs was estimated using univariate genome-wide complex trait analysis (GCTA-GREML; Yang et al., Reference Yang, Benyamin, McEvoy, Gordon, Henders, Nyholt and Visscher2010). The sample size for the GCTA-GREML is slightly smaller than that used in the association analysis (n = 5661), because one individual was excluded from any pair of individuals who had an estimated coefficient of relatedness of >.025 to ensure that effects due to shared environment were not included. The same covariates were included in the GCTA-GREML as for the SNP-based association analysis.
Polygenic profile analyses
Polygenic profile scores were created using PRSice version 2 (Euesden et al., Reference Euesden, Lewis and O’Reilly2015; https://github.com/choishingwan/PRSice). First, we used the GWAS results for health literacy to create health literacy polygenic profile scores in an independent sample and used these scores to predict health literacy, cognitive function and educational attainment phenotypes. Polygenic profile scores for health literacy were created in 1005 genotyped participants from the Lothian Birth Cohort 1936 (LBC1936) study (Deary et al., Reference Deary, Gow, Taylor, Corley, Brett, Wilson and Starr2007) by calculating the sum of alleles associated with health literacy across many genetic loci, weighted by the effect size for each loci. Before the polygenic scores were created, SNPs with a MAF of < 0.01 were removed and clumping was used to obtain SNPs in LD (r 2 < .25 within a 250 kb window). Five scores were then created that included SNPs according to the significance of the association with health literacy, based on the following p-value thresholds: p < .01, p < .05, p < 0.1, p < .5 and all SNPs. Linear regression was used to investigate whether polygenic profiles for health literacy were associated with performance on the Newest Vital Sign (Weiss et al., Reference Weiss, Mays, Martz, Castro, DeWalt, Pignone and Hale2005), a test of health literacy similar in content to the ELSA health literacy test, a measure of general cognitive ability and years of schooling (see Supplementary Methods for more detail on these phenotypes). Models were adjusted for age, sex and four genetic principal components, and standardized betas were calculated.
Next, we used summary GWAS results from 21 GWAS of cognitive and health-related phenotypes to create polygenic profile scores for cognitive and health-related traits in ELSA participants. As the creation of polygenic scores requires summary GWAS results from an independent sample, the GWAS of general cognitive ability (Davies et al., Reference Davies, Lam, Harris, Trampush, Luciano, Hill and Deary2018) was rerun removing ELSA participants. SNPs with a MAF of < .01 were removed and clumping was used to obtain SNPs in LD (r 2 < .25 within a 250 kb window) prior to the creation of the polygenic scores. Five scores were created for each phenotype based on the p-value thresholds detailed above. For Alzheimer’s disease, we created a second set of scores with a 500 kb region around the APOE locus removed [hereafter called ‘Alzheimer’s disease (500 kb)’] to create a polygenic risk score of Alzheimer’s disease with and without the APOE locus.
These polygenic scores were converted to z scores. Logistic regression was used to investigate whether polygenic profiles for cognitive and health-related traits were associated with having adequate, compared to limited, health literacy in ELSA participants. All models were adjusted for age at wave 2, sex and the 15 genetic principal components to control for population stratification. For each phenotype, five logistic regression models were run using the five polygenic scores created based on the p-value thresholds; thus, a total of (5 × 21) 105 models were run. To control for multiple testing, the reported p-values are false discovery rate-corrected. This method controls for the number of false positive results in those that reach significance (Benjamini & Hochberg, Reference Benjamini and Hochberg1995). A multivariate logistic regression model was run including all of the significant polygenic scores, controlling for age, sex and 15 genetic principal components to test whether these polygenic scores independently contributed to health literacy.
Results
Of the 7358 participants who remained following genotyping quality control procedures, 5783 (3160 female; 54.6%) had completed the health literacy test at wave 2 and form the analytic sample (mean age = 65.49, SD = 9.55). A total of 4012 (69.4%) participants had adequate health literacy, whereas 1771 (30.6%) participants had limited health literacy. Participants with limited health literacy were older (mean age = 67.76, SD = 10.00) than participants with adequate health literacy (mean age = 64.72, SD = 9.19; t(3140.90) = 10.91, p < .001).
Genome-wide association study
A genome-wide association analysis of health literacy found no genome-wide significant (p < 5 × 10-8) SNP associations. There were 131 suggestive SNP associations (p < 1 × 10−5). The SNP-based Manhattan plot is shown in Figure 1 (the SNP-based QQ plot is shown in Supplementary Figure S1; suggestive SNPs are reported in Supplementary Data S1). Genomic risk loci characterization performed using FUMA with the genome-wide suggestive significance threshold (p < 1 × 10−5) identified 39 ‘independent’ significant SNPs distributed within 36 loci; see Methods section for the description of independent SNP selection criteria. For consistency, we use the term ‘independent suggestively significant SNP’ here according to the definition that is used in the relevant analysis package and the significance threshold described above. Details of functional annotation of these independent suggestively significant SNPs and tagged SNPs within the 36 loci can be found in Supplementary Data S2.
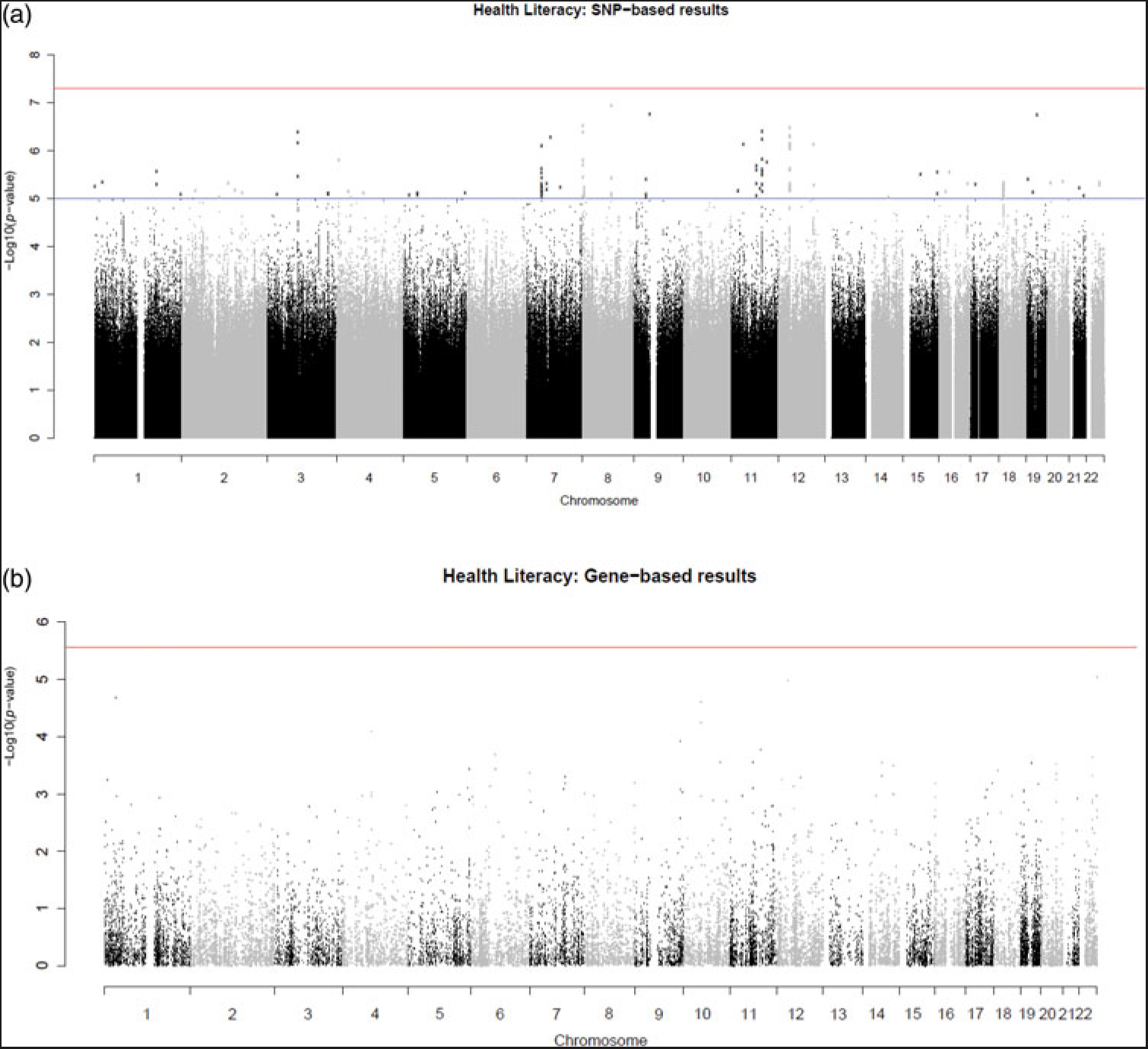
Fig. 1. (Color online) SNP-based (a) and gene-based (b) association results for health literacy. The red line indicates the threshold for genome-wide significance: p < 5 × 10−8 for (a), p < 2.75 × 10−6 for (b); the blue line in (a) indicates the threshold for suggestive significance: p < 1 × 10−5.
Comparison with previous findings
Of the 39 independent suggestively significant and 253 tagged SNPs (those in LD with the independent suggestively significant SNPs), none had been reported as reaching genome-wide (p < 5 × 10-8) or suggestive (p < 1 × 10-5) significance in previous GWAS of general cognitive ability or years of education.
Gene-based analyses
No genome-wide significant findings were found from the gene-based association analysis; the gene-based association results are shown in Supplementary Data S3 (the gene-based Manhattan plot is shown in Figure 1; the QQ plot is shown in Supplementary Figure S1). The gene-set and gene-property analyses also did not identify any significant results (Supplementary Data S4 and S5).
SNP-based heritability
We estimated the proportion of variance explained by all common SNPs to be 0.085 (SE = 0.072). We note that, with the large standard error, this does not rule out zero SNP-based heritability.
We did not calculate genetic correlations between health literacy and those phenotypes included in the polygenic profile analyses as we did not have adequate power in this sample to utilize either the LD score regression method or, for those phenotypes also available in ELSA, bivariate GCTA-GREML. The mean chi-squared value for the health literacy phenotype was 1.009, which is below the LD score regression recommended threshold of 1.02 (Bulik-Sullivan et al., Reference Bulik-Sullivan, Loh, Finucane, Ripke, Yang and Patterson2015). This indicates that there is too small a polygenic signal for these methods to work with.
Health literacy polygenic profile scores predicting health literacy, cognitive function and educational attainment in LBC1936
Polygenic profile score for health literacy did not significantly predict performance on the Newest Vital Sign, cognitive ability or years of schooling in LBC1936 (Supplementary Table S2).
Cognitive and health-related polygenic scores predicting health literacy in ELSA
Table 1 shows the results of the association between cognitive and health-related polygenic scores and health literacy in ELSA participants, using the most predictive threshold. Supplementary Table S3 reports the full results for all thresholds.
Table 1. Association between polygenic profiles of cognitive, socioeconomic, health and personality traits with having adequate health literacy, controlling for age, sex and 15 genetic principal components

Note: FEV1 = forced expiratory volume in 1 s; BMI = body mass index. The associations between the polygenic profile with the largest effect size (threshold) and the health literacy phenotype are reported.
* Nagelkerke Pseudo R 2. R 2 is calculated by subtracting the value of a model containing only the covariates (age, sex and 15 genetic principal components) from the model including the polygenic profile score and covariates.
† p-values reported have been FDR-adjusted. FDR-adjusted significant p-values are shown in bold.
Increased odds of having adequate, compared to limited, health literacy were associated with a one standard deviation higher polygenic profile score for general cognitive ability [OR = 1.34, 95% CI (1.26, 1.42)], verbal-numerical reasoning [OR = 1.30, 95% CI (1.23 -1.39)] and years of schooling [OR = 1.29, 95% CI (1.21, 1.36)]. Reaction time and childhood IQ polygenic scores did not predict health literacy. Decreased odds of having adequate health literacy were associated with a 1 standard deviation higher polygenic profile score for poorer self-rated health [OR = 0.92, 95% CI (0.87, 0.98)] and schizophrenia [OR = 0.91, 95% CI (0.85, 0.96)]. No other polygenic scores predicted health literacy.
To examine whether each polygenic profile score improved the prediction of health literacy, the Nagelkerke pseudo R 2 value for a model with only the covariates (age, sex and 15 genetic principal components) was subtracted from the Nagelkerke pseudo R 2 for the model with both covariates and the polygenic score (Table 1). Polygenic profile scores for general cognitive ability, verbal-numerical reasoning and years of schooling accounted for 2.2%, 1.8% and 1.7%, respectively, of the variance in health literacy. The variance in health literacy accounted for by the self-reported health and schizophrenia polygenic scores was small, at 0.2% and 0.3%, respectively.
Table 2 shows the results of the multivariate logistic regression in which polygenic scores for general cognitive ability, verbal-numerical reasoning, years of schooling, self-rated health and schizophrenia were all entered simultaneously. The odds ratios for all polygenic scores were attenuated in this model. Increased odds of having adequate, compared to limited, health literacy were significantly associated with the following: higher polygenic scores for general cognitive ability [OR = 1.18, 95% CI (1.06, 1.32)] and years of schooling [OR = 1.19, 95% CI (1.11, 1.27)]; and lower polygenic risk for schizophrenia [OR = 0.93, 95% (CI 0.88, 0.99)]. Together, these polygenic profile scores accounted for 3.0% of the variance in health literacy. In this multivariate model, the association between the verbal-numerical reasoning polygenic profile score and health literacy was attenuated and nonsignificant. This is not surprising as the general cognitive ability polygenic score is derived from a metaanalysis, which includes the verbal-numerical reasoning test (Davies et al., Reference Davies, Lam, Harris, Trampush, Luciano, Hill and Deary2018). The self-rated health polygenic score was also attenuated and nonsignificant in this model.
Table 2. Multivariate models of the association between polygenic profiles of cognitive and health traits with having adequate health literacy, controlling for age, sex and 15 genetic principal components
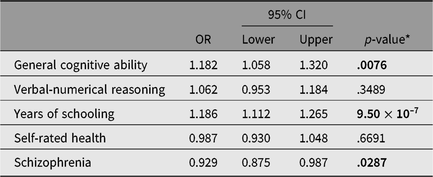
Note: ORs and 95% CIs are from a model in which all five polygenic scores are entered simultaneously, controlling for age, sex and 15 genetic principal components.
* p values reported have been FDR-adjusted (after a false discovery rate correction across five tests). FDR-adjusted significant p-values are shown in bold.
Discussion
Using a sample of 5783 middle-aged and older adults living in England, no SNPs were found to be significantly associated with health literacy; however, we report 131 suggestive SNP associations within 36 independent genomic loci. Using polygenic profile scoring, this study found that genetic variants previously associated with higher general cognitive ability, verbal-numerical reasoning and more years of schooling were associated with having adequate health literacy, whereas genetic variants previously found to be associated with poorer self-rated health and a diagnosis of schizophrenia were associated with having limited health literacy. These results suggest that the phenotypic associations frequently reported between health literacy and cognitive function, and health literacy and health might be partly due to shared genetic etiology. In a multivariate model in which all the significant polygenic scores were entered simultaneously, higher polygenic scores for general cognitive ability, years of schooling and schizophrenia remained significant, suggesting these polygenic scores independently contribute to performance on a health literacy test.
A number of studies have reported phenotypic associations between performance on tests of health literacy and cognitive function (Boyle et al., Reference Boyle, Yu, Wilson, Segawa, Buchman and Bennett2013; Mõttus et al., Reference Mõttus, Johnson, Murray, Wolf, Starr and Deary2014; Murray et al., Reference Murray, Johnson, Wolf and Deary2011; Reeve & Basalik, Reference Reeve and Basalik2014). Due to the strength of these reported associations, some researchers (Mõttus et al., Reference Mõttus, Johnson, Murray, Wolf, Starr and Deary2014; Reeve & Basalik, Reference Reeve and Basalik2014) have proposed that health literacy and cognitive function are not separate constructs and are instead assessing to a substantial extent the same underlying ability. To investigate this overlap, Reeve and Basalik (Reference Reeve and Basalik2014) entered three health literacy tests and six cognitive tests into an exploratory factor analysis. No unique health literacy factor emerged, and in fact, the three health literacy tests each loaded on different factors (Reeve & Basalik, Reference Reeve and Basalik2014). The authors concluded that there is very little evidence that health literacy is unique from cognitive function (Reeve & Basalik, Reference Reeve and Basalik2014). The current study found that the genetic variants associated with cognitive function make significant contributions to performance on tests of health literacy, providing additional evidence that health literacy and cognitive function are intrinsically related and that they might, in part, be associated with the same underlying construct.
Some researchers have suggested that educational attainment can be used as a proxy for cognitive ability in genetic studies (Hill, Marioni et al., Reference Hill, Marioni, Maghzian, Ritchie, Hagenaars, McIntosh and Deary2019; Okbay et al., Reference Okbay, Beauchamp, Fontana, Lee, Pers, Rietveld and Benjamin2016) because: (a) there are large phenotypic and genetic correlations between cognitive function and educational attainment (Hagenaars et al., Reference Hagenaars, Harris, Davies, Hill, Liewald, Ritchie and Deary2016) and (b) it is much easier to collect information on educational attainment than it is to administer cognitive assessments in large studies. In the current study, when all significant polygenic scores were entered simultaneously, the general cognitive ability polygenic score and the years of schooling polygenic score both had independent associations with health literacy. Thus, at least when measuring health literacy, it might not be appropriate to consider cognitive function and educational attainment polygenic scores as proxies for the same underlying ability. On the other hand, it is possible that educational attainment was indexing some aspects of cognitive function not tapped by the phenotypes that went into the cognitive GWAS, which tended to be more fluid in characterization.
The results of the current study provide some evidence that the frequently reported associations between health literacy and health (Berkman et al., Reference Berkman, Sheridan, Donahue, Halpern and Crotty2011) might be partly due to shared genetic influences. We found that genetic variants associated with poorer self-reported health and having a diagnosis of schizophrenia were associated with having poorer health literacy. Many studies have reported phenotypic associations between health literacy and self-reported health status (Berkman et al., Reference Berkman, Sheridan, Donahue, Halpern and Crotty2011; von Wagner et al., Reference von Wagner, Knight, Steptoe and Wardle2007; Wolf et al., Reference Wolf, Gazmararian and Baker2005). There has been relatively little research investigating health literacy and schizophrenia; however, health literacy has been found to be negatively associated with other mental health outcomes including mental health status (Wolf et al., Reference Wolf, Gazmararian and Baker2005) and depressive symptoms (Gazmararian et al., Reference Gazmararian, Baker, Parker and Blazer2000). The ELSA sample used here consisted of relatively healthy community-dwelling adults. In this sample of participants without schizophrenia, having a higher polygenic risk of schizophrenia was associated with poorer health literacy. This mimics the results seen for schizophrenia and cognitive function. Individuals with higher polygenic risk of schizophrenia tend to perform more poorly on tests of cognitive function (Hagenaars et al., Reference Hagenaars, Harris, Davies, Hill, Liewald, Ritchie and Deary2016; McIntosh et al., Reference McIntosh, Gow, Luciano, Davies, Liewald, Harris and Deary2013). In this study, whereas the association between polygenic risk of schizophrenia and poorer health literacy was attenuated when also controlling for cognitive polygenic scores, polygenic risk of schizophrenia remained a significant predictor of health literacy, suggesting the associations reported here are not simply because of any overlap between cognitive function and schizophrenia.
One strength of the current study is that we used GWAS summary results from a large number of cognitive and health-related traits, which enabled a comprehensive investigation of the shared genetic influences between health literacy, cognitive function and health. Whereas phenotypic associations between health literacy and health-related traits such as type 2 diabetes (Wolf et al., Reference Wolf, Gazmararian and Baker2005) and Alzheimer’s disease (Kaup et al., Reference Kaup, Simonsick, Harris, Satterfield, Metti, Ayonayon and Yaffe2014; Yu et al., Reference Yu, Wilson, Schneider, Bennett and Boyle2017) have been identified, we did not find that genetic variants previously associated with these health-related traits were associated with health literacy in this study. One limitation of the current study is that the quality of the polygenic profile scores created depends on the quality of the original GWAS. Many of the GWAS are metaanalyses, which introduces heterogeneity in both the genetic methods used and in measuring the phenotype. Some of the GWAS have relatively small sample sizes. It is possible that we did not find an association between some of the health and cognitive polygenic scores with health literacy because the original GWAS was underpowered to identify genetic associations with the phenotype.
Unlike recent GWAS of cognitive function (Davies et al., Reference Davies, Lam, Harris, Trampush, Luciano, Hill and Deary2018; Savage et al., Reference Savage, Jansen, Stringer, Watanabe, Bryois, de Leeuw and Posthuma2018), which found many genetic variants associated with cognitive function, we found no SNPs significantly associated with health literacy. It is now well known that for polygenic traits, the effect of individual genetic variants on a trait is likely to be very small and therefore larger sample sizes than the one used here are required to identify such associations (Deary et al., Reference Deary, Harris and Hill2019). Identification of many genetic variants associated with cognitive function is only now possible because of the ever-increasing sample sizes. For cognitive function, early studies of approximately 3500 individuals found no significant SNPs (Davies et al., Reference Davies, Tenesa, Payton, Yang, Harris, Liewald and Deary2011). However, a more recent GWAS of cognitive function used data from over 300,000 individuals and found over 1000 significant SNPs (Davies et al., Reference Davies, Lam, Harris, Trampush, Luciano, Hill and Deary2018). The GWAS reported here is therefore underpowered. Given the cognitive literature, much larger samples sizes — at least 10 times larger — than the sample size used here are probably needed to begin to understand the specific genetic variants involved in health literacy. There are, however, few large studies that measure health literacy. The present study is the first investigation of the molecular genetic contributions to health literacy. We encourage other groups with both health literacy and genetic data to explore the genetic associations of health literacy. In an effort to increase power, future studies should look to conduct a metaanalysis of GWAS of health literacy.
None of the suggestively significant SNPs identified in this study have previously been reported as genome-wide significantly associated with cognitive function (Davies et al., Reference Davies, Lam, Harris, Trampush, Luciano, Hill and Deary2018) or years of schooling (Okbay et al., Reference Okbay, Beauchamp, Fontana, Lee, Pers, Rietveld and Benjamin2016). If health literacy, cognitive ability and education do have shared genetic influences, and the suggestive health literacy findings reported here are found to be true associations, then they appear to be associated with an aspect of health literacy that does not have shared genetic etiology with cognitive function or years of education. However, given the small sample size, some of the observed suggestive signals reported here may be due to chance.
The present study found that the SNP-based heritability for health literacy was 8.5% (SE = 7.2%), which is lower than has been reported in studies testing the SNP-based heritability of cognitive phenotypes (Davies et al., Reference Davies, Tenesa, Payton, Yang, Harris, Liewald and Deary2011, Reference Davies, Lam, Harris, Trampush, Luciano, Hill and Deary2018). The heritability estimates for cognitive function were calculated using much larger samples than were used here. For example, the SNP-based heritability for general cognitive function was found to be 25% (SE = 0.6%) using a sample of 86,010 UK Biobank participants (Davies et al., Reference Davies, Lam, Harris, Trampush, Luciano, Hill and Deary2018). Measurement characteristics of health literacy in the present study probably lowered the estimate of SNP-based heritability. In common with other studies using this test (Gale et al., Reference Gale, Deary, Wardle, Zaninotto and Batty2015; Kobayashi et al., Reference Kobayashi, Wardle and von Wagner2014), we dichotomized test scores into adequate (4/4 correct) and limited (<4 correct) health literacy. Given the brief nature of this test, and given that we have defined health literacy as a dichotomy, it is possible that the SNP-based heritability estimate reported here was attenuated. Longer tests, which can capture the continuum in health literacy variance, would be preferable and could test this possibility. On the other hand, we note that dichotomous variables in this general area can result in larger SNP-based heritability; for example, having or not having a college or university degree in the UK Biobank sample had a SNP-based heritability of 21% (SE = 0.6%; Davies et al., Reference Davies, Marioni, Liewald, Hill, Hagenaars, Harris and Deary2016).
One strength of this study is that health literacy was measured consistently in all participants. One limitation is that the health literacy measure used in ELSA is a brief, four-item test that has a ceiling effect. That is, 70% of participants scored full marks (4/4) on this test. Despite the brief nature of the ELSA health literacy test, and despite the ceiling effect, this measure has been found to be associated with various health outcomes, including mortality (Bostock & Steptoe, Reference Bostock and Steptoe2012). In the current study, this health literacy test was sensitive enough to identify associations with polygenic scores for cognitive and health-related traits. Future research examining the genetic contributions to health literacy should use more detailed and continuous measures of health literacy.
Phenotypic associations have consistently been found between health literacy — the skills and ability required to manage ones health — and cognitive function and health. This study is the first to investigate whether health literacy and cognitive function, and health literacy and health, share genetic architecture. In this study we investigated the genetic associations of health literacy and tested whether genetic contributions to cognitive function and health are associated with health literacy. No SNPs had genome-wide significant associations with health literacy. Polygenic scores for cognitive function, years of schooling, self-reported health and schizophrenia were significantly associated with performance on a brief test of health literacy. These results indicate that the phenotypic associations between health literacy and cognitive function, and health literacy and health may be partly due to shared genetic etiology between these traits. Future studies should build on the heritability estimate and polygenic profile results reported here and explore the genetic overlap and distinctiveness of health literacy, cognitive ability, education and health. As the number of participants increases, we will be able to determine the SNPs, genes, and gene-sets that are shared and distinct between health literacy, cognitive function and health.
Supplementary material
To view supplementary material for this article, please visit https://doi.org/10.1017/thg.2019.28.
Acknowledgments
The English Longitudinal Study of Ageing is jointly run by University College London, Institute for Fiscal Studies, University of Manchester and National Centre for Social Research. Genetic analyses have been carried out by UCL Genomics and funded by the Economic and Social Research Council and the National Institute on Aging. All GWAS data have been deposited in the European Genome-Phenome Archive. Data governance was provided by the METADAC data access committee, funded by ESRC, Wellcome and MRC. (2015-2018: Grant Number MR/N01104X/1 2018-2020: Grant Number ES/S008349/1).
The present study was supported by the University of Edinburgh Centre for Cognitive Ageing and Cognitive Epidemiology, part of the cross council Lifelong Health and Wellbeing Initiative, funded by the Biotechnology and Biological Sciences Research Council (BBSRC), and Medical Research Council (MRC) (grant number MR/K026992/1). The Lothian Birth Cohort 1936 is funded by Age UK (Disconnected Mind grant). SPH is funded by the Medical Research Council (MR/S0151132). This study presents independent research part supported by the National Institute for Health Research (NIHR) Biomedical Research Centre at South London and Maudsley NHS Foundation Trust and King’s College London. The views expressed are those of the author(s) and not necessarily those of the NHS, NIHR, Department of Health or King’s College London.
Conflict of interest
None.
Ethical standards
The authors assert that all procedures contributing to this work comply with the ethical standards of the relevant national and institutional committees on human experimentation and with the Helsinki Declaration of 1975, as revised in 2008.