Iodine, a reactant in the synthesis of thyroid hormones, is an essential nutrient absorbed mainly through drinking water and food to meet the needs of the human. Autoimmune thyroiditis (AIT) is a common thyroid disease, and the characteristic of AIT is the infiltration of lymphocytes and positivity for thyroid antibodies in the thyroid. Studies have confirmed that iodine as a nutritional factor intake could change the occurrence of AIT(Reference Dayan and Daniels1).
The association of genetic susceptibility factors and environmental factors could cause the development of AIT. Among environmental factors, iodine nutrition is an identified risk factor. On the one hand, the occurrence of AIT could be induced by iodine excess (IE). Excessive iodine consumption is a risk factor for the development of thyroid autoimmunity(Reference Luo, Kawashima and Ishido2). On the other hand, iodine supplementation in the areas of iodine deficiency increases the prevalence of AIT, and a cohort study has indicated that the prevalence of AIT elevated after iodine fortification (IF) in iodine-deficient areas(Reference Pedersen, Knudsen and Carlé3). For the genetic factors, the role of epigenetics in thyroid diseases has attracted increasing attention. DNA methylation is the most widely studied epigenetic mechanism(Reference Law and Holland4). Studies found that aberrant DNA methylation patterns in the genes such as ICAM-1 and PTPN22 have been associated with AIT(Reference Shalaby, Mackawy and Atef5,Reference Kyrgios, Giza and Fragou6) , and DNA methylation variations may affect the development and prognosis of AIT(Reference Hashimoto, Watanabe and Inoue7,Reference Guo, Wu and Yu8) . Therefore, whether iodine nutrition, as an environmental factor, can affect the methylation of certain genes contributing to AIT pathogenesis must be studied.
Apoptosis is a biological process in which cells actively destroy themselves. Genetic programmes can control apoptosis, which leads to the destruction of DNA and changes in cell morphology(Reference Wyllie, Morris and Smith9). Increasing evidence suggests that apoptosis plays an important role in AIT(Reference Wang and Baker10,Reference Lin11) . Apoptosis can affect the homeostasis of thyroid cells and participate in the destructive mechanism of AIT. The percentage of apoptotic thyrocytes increased in Hashimoto’s thyroiditis (HT), and apoptotic cells were mainly distributed in disrupted follicles and on the periphery of infiltrating lymphoid cells(Reference Hammond, Lowdell and Cerrano12). Studies have found intrinsic apoptotic signalling pathway is involved in the pathogenesis of AIT(Reference Lin11). Among the intrinsic apoptotic signalling pathways, the fate of thyrocytes was determined by the activity of the anti-apoptotic and apoptotic genes. For instance, in HT, the BCL2 gene is elevated in lymphocytes but decreased in thyrocytes, and this suggests that the relationship between BCL2 family expression and thyrocyte destruction is considered to be a key factor in regulating thyrocyte survival(Reference Stassi and De Maria13). However, the relationship between intrinsic apoptosis-associated gene methylation and AIT risk has not been reported.
Therefore, in this study, we investigated the effects of different levels of iodine on intrinsic apoptosis-associated gene methylation in AIT. We also investigated the relationships between intrinsic apoptosis-associated gene methylation in the whole-blood DNA and the environmental factor of iodine level, as well as their addictive and multiplicative interactions in AIT.
Materials and methods
Survey areas and participants
Based on the recent data(Reference Wang, Wan and Liu14), participants from three survey areas with different levels of iodine (water iodine content and the use of iodised salt) were chosen to conduct an epidemiological study in Shandong province. The details were as follows: 1. IF, with water iodine content ≤ 10 μg/l and > 90 % coverage rates of qualified iodised salt, including Dongtan and Qianlv villages. These areas were iodine-deficient areas until the implementation of universal salt iodisation in 1995; 2. iodine adequate (IA), with 40 μg/l < water iodine content < 100 μg/l and non-iodised salt supplied, including Liuxiangzhuang and Dongding villages; and 3. IE, with water iodine content ≥ 100 μg/l and non-iodised salt supplied, including Xieyuanji village. In total, 176 pairs of AIT cases and controls were selected (IF: 89 pairs, IA: 40 pairs and IE: 47 pairs). For the AIT group, the inclusion criteria were 1. serum thyroid peroxidase antibodies (TPOAb), thyroglobulin antibodies (TgAb) or double antibody positivity(Reference Weetman15); 2. Thyroid ultrasound examination has the following conditions: goitre or echo heterogenicity or multiple hypoechoic areas; 3. absence of hyperthyroidism and subclinical hyperthyroidism. For the control group, the inclusion criteria were 1. normal healthy participants matched to the case group based on sex, age, place of residence and BMI; 2. no family or personal history of autoimmune diseases or other thyroid diseases, no chronic or acute diseases, no use of long-term thyroid drugs or hormones and no pregnancy; 3. no goitre; negative TgAb and TPOAb; normal findings for other thyroid function indicators and no thyroid ultrasound abnormalities. This study was consistent with the Declaration of Helsinki and was approved by the Harbin Medical University Ethics Review Committee (No. hrbmuecdc20200320). Participants in the study all signed informed consent forms.
Laboratory testing
Venous blood was collected from the participants after an 8-hour fast. The levels of FT3, FT4, TSH, TGAb and TPOAb were measured using chemiluminescence immunoassays (Siemens Inc.). The reference ranges were as follows: 11·5 pmol/l < FT4 < 22·7 pmol/l; 3·1 pmol/l < FT3 < 6·8 pmol/l; 0·27 mIU/l < TSH < 4·20 mIU/l. Isolated positive TPOAb was defined as having TPOAb ≥ 34 μg/ml and TGAb < 115 μg/ml.; Isolated positive TGAb was defined as having TGAb ≥115 μg/ml and TPOAb < 34 μg/ml; TPOAb(+)&TGAb(+) were defined as TPOAb ≥ 34 μg/ml and TGAb ≥ 115 μg/ml. The water iodine content and urinary iodine concentrations (UIC) were determined with the As3+-Ce4+ catalytic spectrophotometry method(Reference Krzanowski16). Certified reference materials (GBW09108, GBW9109 and GBW9110) from the NRLIDD in China were used to control measurement quality, and the target values were 69·5 ± 9·0 μg/l, 134 ± 10 μg/l and 239 ± 15 μg/l. The intra-assay CV was 2·7, 1·4 and 2·3 %, and the inter-assay CV was 2·3, 2·5 and 2·4 %.
Determination of candidate genes and DNA methylation measurements
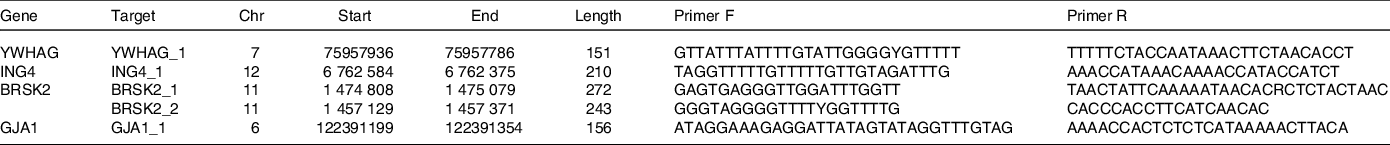
In our previous study(Reference Wan, Liu and Ren17), an Illumina Methylation 850K BeadChip was used to detect genome-wide DNA methylation levels in whole blood samples from the case and control groups. A total of 853 307 CpG sites were detected, and a total of 312 differential methylation sites in 257 differentially methylated genes were screened according to methylation differences and P values. To further investigate the key biological processes associated with these distinct genes, we assessed the significant key biological processes associated with AIT according to the GO databases, including intrinsic apoptosis. The genes associated with apoptosis are shown in Supplementary Table 1. YWHAG, ING4, BRSK2 and GJA1 genes were enriched in the intrinsic apoptosis process and were closely associated with AIT. DNA was extracted by the TIANGEN Extraction Kit (TIANGEN). DNA methylation was evaluated with MethylTarget performed by Genesky Biotechnologies Inc.. Table 1 lists the primers of selected genes. Genomic DNA was transformed into bisulphite by EZ DNA methylation kit (Zymo). According to the guidelines, all the samples were amplified, barcoded and sequenced (Illumina).
Table 1. Primer sequences in MethylTargetTM assay
Quantitative real-time PCR (qRT-PCR)
Total RNA was extracted from whole blood with RNAiso Plus (Takara). NanoDrop 2000C (Thermo Fisher) was used to determine the RNA concentrations. The value of OD 260/280 between 1·8 and 2·0 was thought to be sufficient quality. Each sample was performed with a QuantStudioTM5 real-time PCR (Thermo Fisher). The amplification reaction system was as follows: 5·0 μl SYBR Green (Roche), 1·0 μl cDNA, 0·5 μl upstream and downstream primer and 3·0 μl ddH2O. The reaction steps were as follows: 1. hold stage, 95°C 10 min; 2. PCR stage (40 cycles): 95°C 15 s and 60°C 1 min; 3. melt curve stage: 95°C 15 s, 60°C 1 min and 95°C 15 s. Specific primers were as follows: YWHAG-F: 5′-CGG CAA TGA GAA GAA GAT TGA G-3′; YWHAG-R: 5′-GCT GCA ATT CTT GAT CAG GTA G-3′; ING4-F: 5′-AGT TGG CCA CTG AGT ATA TGA G-3′; ING4-R: 5′-TGT GTT TGT CCA CCA TCT CAT A-3′; BRSK2-F: 5′-TTC CAC ATG CCG CAC TTT ATC-3′; BRSK2-R: 5′-GTG TTT CTG AAT GTG CTC TAG C-3′; β-actin-F: 5′-CCT TCC TGG GCA TGG AGT CCT G-3′; β-actin-R: 5′-GGA GCA ATG ATC TTG ATC TTC-3′. The 2–ΔΔCt method was used to analyse the levels of mRNA of the target genes.
Statistical analysis
In this study, we used SPSS 23·0 for statistical analysis. Normally distributed data are characterised by the mean ± standard deviation (sd) and were analysed with the Student’s t test and one-way ANOVA. Non-normally distributed data are described with median and 25th and 75th percentiles and were analysed with the Kruskal–Wallis H test and Mann–Whitney U test. The chi-square (χ2) test was used to compare the rates of the subjects with different thyroid antibody groups. The correlations between relative mRNA expression and the methylation levels of candidate genes were analysed with Pearson correlation analysis. Interactions between different levels of iodine and methylation levels of candidate genes on AIT were evaluated with multiplicative and addictive interaction models. The methylation levels in target regions were transformed into a bicategorical variable of hypomethylation and hypermethylation, the different levels of iodine were divided into IF, IA and IE and two risk factors (methylation levels and different levels of iodine) and their interaction terms were included in a logistic regression model(Reference Li, He and Wei18). GMDR is an alternative linear or logistic regression to non-parametric and gene-free models for detecting and characterising non-linear interactions between environmental attributes and discrete genetics(Reference Xu, Xu and Hou19). We used this method to analyse the high-dimensional interactions among three candidate genes (YWHAG, ING4 and BRSK2) and different levels of iodine on AIT. Age, sex, BMI, smoking, drinking, thyroid function, UIC and family history of thyroid disease were chosen as covariates. Among the candidate models, the most suitable models were those with a sign test P < 0·05, and the highest cross-validation consistency, training balanced accuracy and testing balanced accuracy(Reference Zhao, Yu and Lv20).
Results
Demographic characteristics and DNA methylation levels in participants
The distribution of basic demographic characteristics and thyroid function is shown in Table 2. According to different levels of iodine, a total of 352 participants were divided into three groups. No significant differences were found between AIT and control groups in terms of sex, age, UIC and BMI. In IF and IE, the levels of TSH in the AIT groups were significantly higher than that in the control groups (both P < 0·050). The prevalence of i-TGAb(+) and TPOAb(+) and TGAb(+) are significantly different among the different levels of iodine (both P < 0·050). In Table 3, we detected the methylation levels of five target regions in four genes. The AIT group exhibited higher methylation in the target regions of YWHAG_1 and BRSK2_1 than the control group (both P < 0·010). BRSK2_2 methylation level was not detected in the control group; for DNA methylation in the target regions of ING4_1 and GJA1_1, no significant differences were found between the AIT and control groups (P > 0·050). To find the details of the methylation status in these genes, all CpG sites on the targets were calculated. In Fig. 1, there are a total of thirty-two CpG sites in five target regions in four genes that were analysed. For YWHAG_1, six CpG sites exhibited higher methylation. There are eight hypermethylation CpG sites in BRSK2_1 and one hypomethylation CpG site in ING4_1 in AIT patients compared with controls (all P < 0·050).
Table 2. The demographic data in case and control groups with different levels of iodine
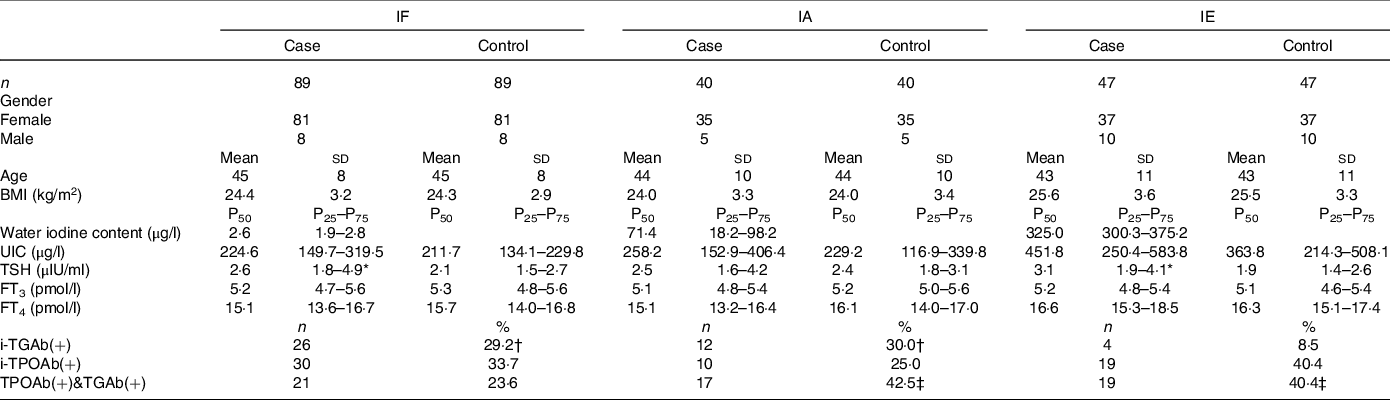
IF, iodine fortification; IA, iodine adequate; IE, iodine excess; UIC, urinary iodine concentrations; TSH, thyroid stimulating hormone; FT3, free triiodothyronine; FT4, free thyroxine; i-TPOAb(+), isolated positive TPOAb; i-TGAb(+), isolated positive TGAb; TPOAb(+)&TGAb(+), double positive TPOAb and TGAb.
* Significant differences between case and control groups.
† Significant differences compare to IE.
‡ Significant differences compare to IF. P < 0·05.
Table 3. DNA methylation levels of regions between cases and controls (mean ± sd)
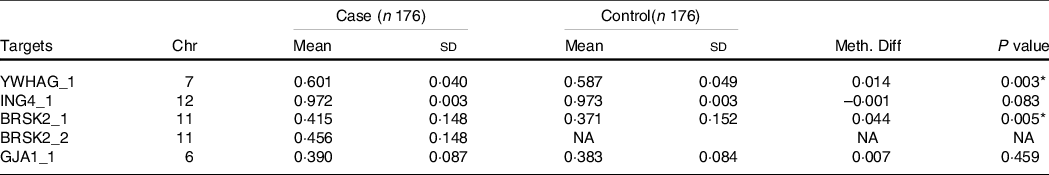
Chr, Chromosome; Meth. Diff = The methylation level of case-The methylation level of control; NA, not applicable.
* P < 0·01.
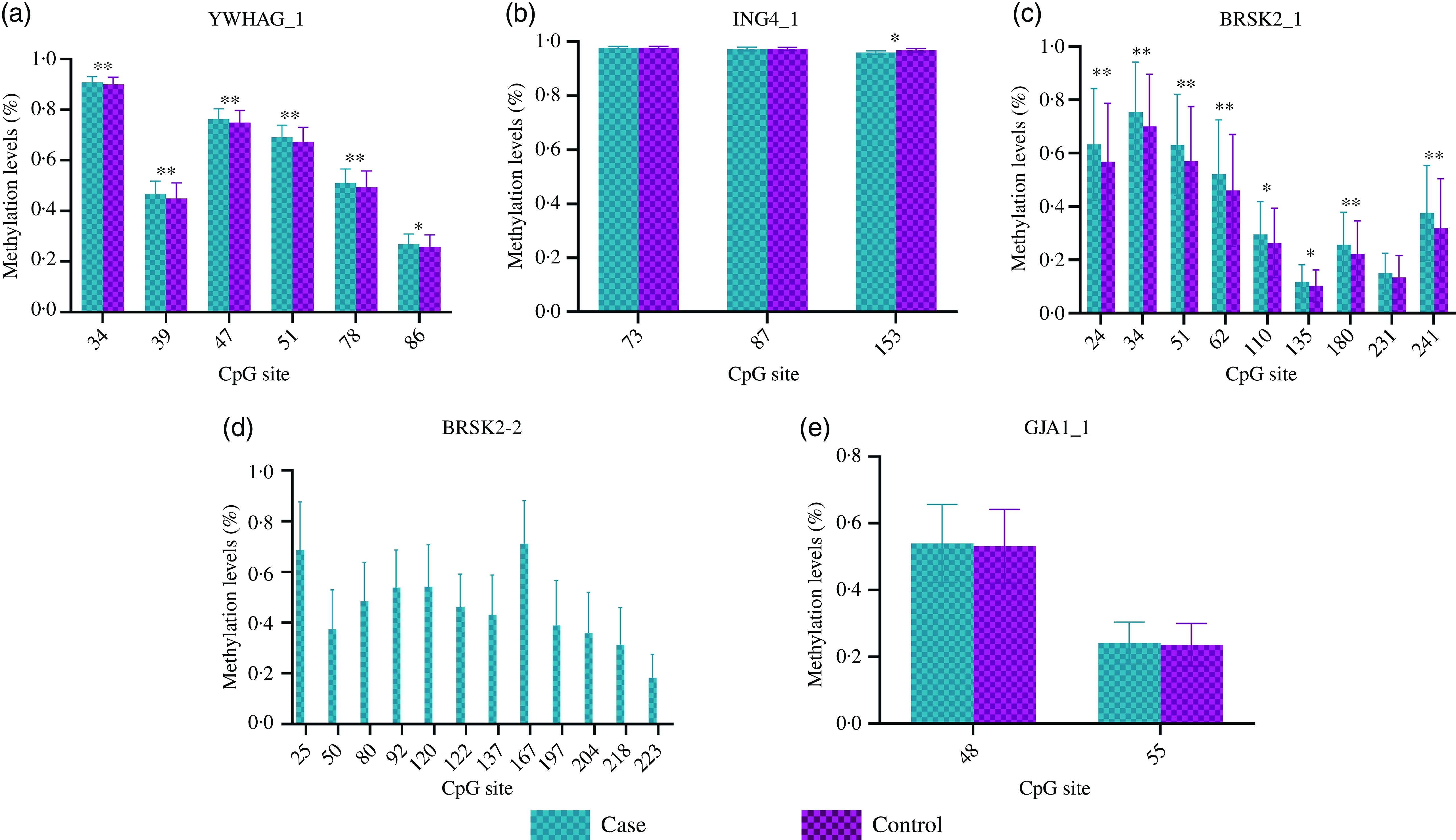
Fig. 1. DNA methylation levels of CpG sites between cases and controls. (a) YWHAG_1; (b) ING4_1; (c) BRSK2_1; (d) BRSK2_2; (e) GJA1_1; The case group compared with the control group, * P < 0·05, ** P < 0·001.
DNA methylation status of participants with different levels of iodine
In Table 4, based on the different levels of iodine, a stratified analysis was performed. In IF, compared with the healthy controls, the AIT patients exhibited higher methylation of YWHAG_1 and BRSK2_1 (both P < 0·050), but lower methylation of ING4_1 (P = 0·049). In IA and IE, we observed no significant differences in DNA methylation in the target regions for all candidate genes (all P > 0·050). As shown in Table 5, in IF, there were six hypermethylation CpG sites in YWHAG_1 and nine hypermethylation CpG sites in BRSK2_1, one CpG site exhibited lower methylation in ING4_1 in patients with AIT than in controls (all P < 0·05). In IA, six CpG sites exhibited higher methylation in BRSK2_1 in patients with AIT than in controls (all P < 0·050). In IE, no CpG site in the four genes exhibit a difference between AIT and the control group (all P > 0·050).
Table 4. DNA methylation levels of targets between cases and controls with different levels of iodine
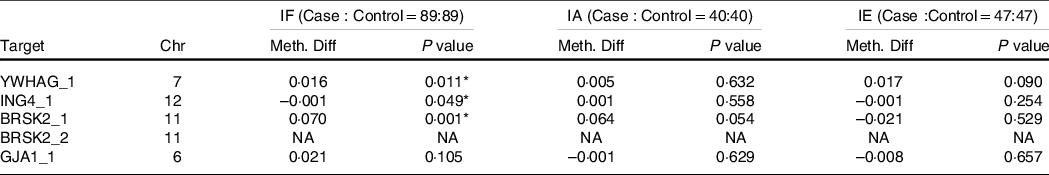
IF, iodine fortification; IA, iodine adequate; IE, iodine excessive; Chr, Chromosome; Meth. Diff = The methylation level of case-The methylation level of control; NA, not applicable.
* P < 0·05.
Table 5. DNA methylation levels of CpG sites between cases and controls with different levels of iodine
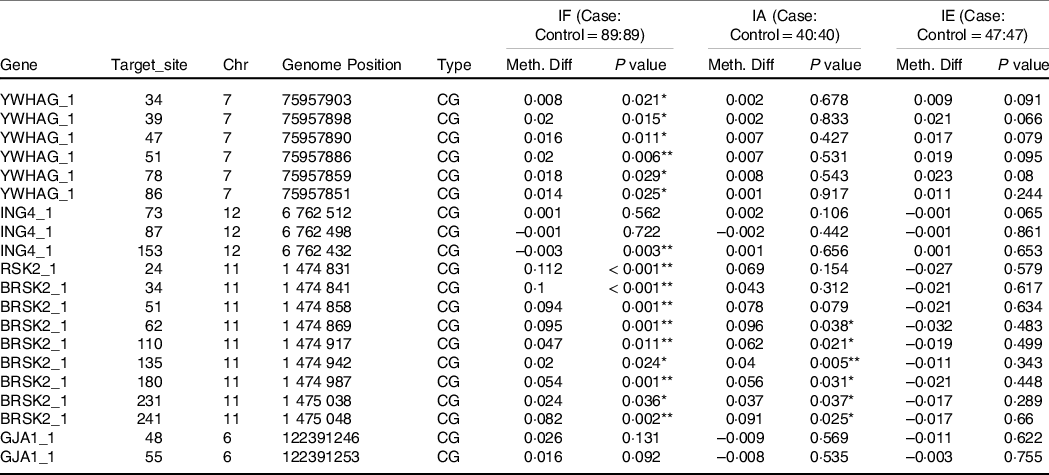
IF, iodine fortification; IA, iodine adequate; IE, iodine excess; Chr, Chromosome; Meth. Diff = The methylation level of case-The methylation level of control.
* P < 0·05, ** P < 0·01.
Correlations among intrinsic apoptosis-associated genes’ DNA methylation levels, age, UIC, thyroid function and thyroid antibodies
As shown in Fig. 2, the associations between the methylation levels of target regions and age were analysed with Pearson correlation. The associations between the UIC, thyroid function and the methylation levels in target regions in patients with AIT were analysed with Spearman’s rank correlation. Positive correlations were found between the methylation levels of YWHAG_1 and BRSK2_1 and age (both P < 0·001). The levels of TSH were positively correlated with the DNA methylation levels of YWHAG_1 and BRSK2_1 (both P < 0·050), but for UIC and FT4, the correlation was negative (all P < 0·050) in patients with AIT. Positive correlations were found between the methylation level of GJA1_1 and FT4 (P = 0·021). No correlations were observed between FT3 and target regions’ methylation. In Supplementary Table 2, we analysed the relationship between YWHAG, ING4 and BRSK2 gene methylation and serum TPOAb and TGAb levels. Among different thyroid antibody groups, the methylation levels of BRSK2_1 are significantly different (P = 0·039). TPOAb and TGAb were divided into three groups according to different levels of titre, and we further analysed the relationship between thyroid antibodies’ titre and DNA methylation levels. Among three different titres of TPOAb, we observed a significant difference in DNA methylation of the ING4 gene (P = 0·045) and BRSK2 gene (P = 0·033).

Fig. 2. Heat map of correlation between DNA methylation status of target region, age, UIC and thyroid function in AIT patients. *P < 0·05.
Combined and interactive effects between methylation levels in target regions and different levels of iodine in autoimmune thyroiditis
As shown in Tables 6 and 7, we performed a cross-over analysis to find the combined effect between different levels of iodine and methylation levels in AIT. Combinations of IF, YWHAG_1 hypermethylation (OR = 2·63, 95 % CI: 1·22, 5·69, P = 0·014) and BRSK2_1 hypermethylation (OR = 4·48, 95 % CI: 2·22, 10·59, P < 0·001) were significantly associated with elevated AIT risk, as compared with that in the reference group. Meanwhile, the interactive effects were analysed. We observed no significant multiplicative effects between methylation levels in target regions and IF in AIT. Moreover, we did not find any combined and multiplicative effects between IE and methylation levels in target regions in AIT. Because no difference was found between the methylation levels of the GJA1 gene between the AIT patients and the controls at different iodine levels, we did not analyse the interaction between the iodine level and GJA1.
Table 6. Combined and interactive effects between methylation levels in the target region and IF in AIT
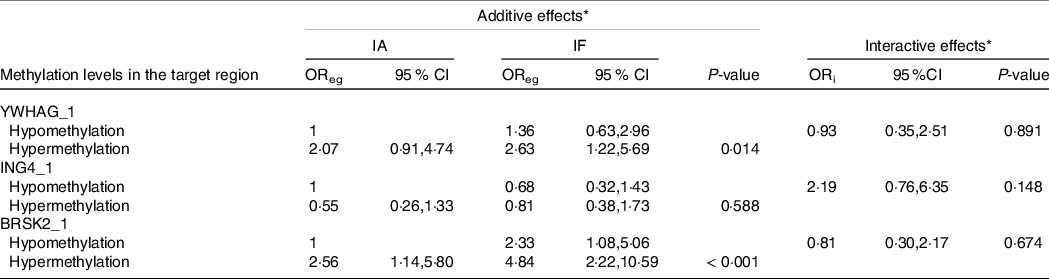
IA, iodine adequate; IF, iodine fortification; OReg: OR genetic&environment, additive effects of methylation and IF; ORi: OR interaction, multiplicative effects of methylation and IF.
* Adjustment: age, gender, BMI, smoking, drinking thyroid function and family history of thyroid disease.
Table 7. Combined and interactive effects between methylation levels in target region and IE in AIT
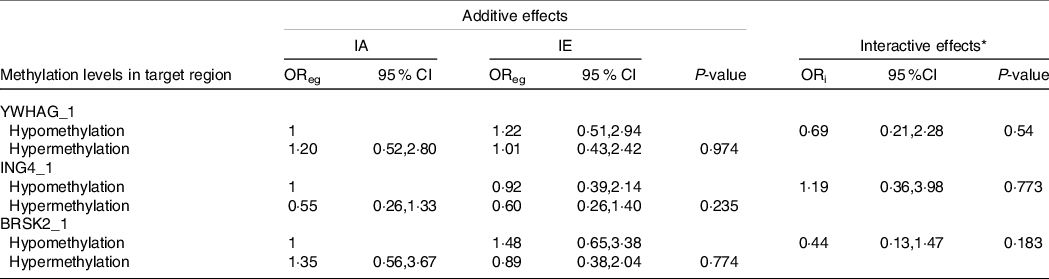
IA, iodine adequate; IE, iodine excessive OReg: OR genetic&environment, additive effects of methylation and IE; ORi: OR interaction, multiplicative effects of methylation and IE.
* Adjustment: age, gender, BMI, smoking, drinking thyroid function and family history of thyroid disease.
GMDR analysis of gene–environment interactions
Table 8 presents the GMDR gene–environment interaction model. The four-locus model (YWHAG_1 × ING4_1 × BRSK2_1 × iodine level) was the best model, with the highest training balanced accuracy (0·661) and testing balanced accuracy (0·606). This model also had the greatest cross-validation consistency (10/10; sign test P = 0·010). Therefore, this was the best model for assessing the interactions between different levels of iodine exposure and candidate gene DNA methylation. Figure 3 demonstrate the specific distribution of score in the best gene–environment interaction model. In different grids of Fig. 3, the scores for AIT groups are varied, suggesting the patterns of risk differed across multi-locus dimensions.
Table 8. GMDR analysis for the best gene–environment interaction models
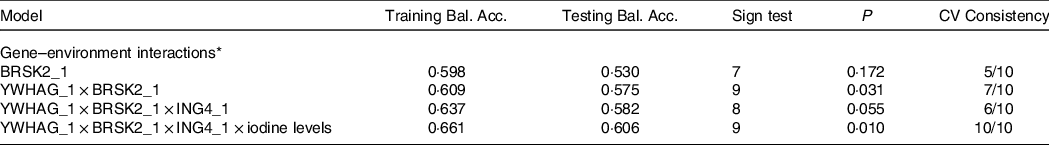
Training Bal. Acc. Training balanced accuracy; Testing Bal. Acc. Testing balanced accuracy, CV consistency cross-validation consistency.
* Adjustment: age, gender, BMI, smoking, drinking thyroid function and family history of thyroid disease.
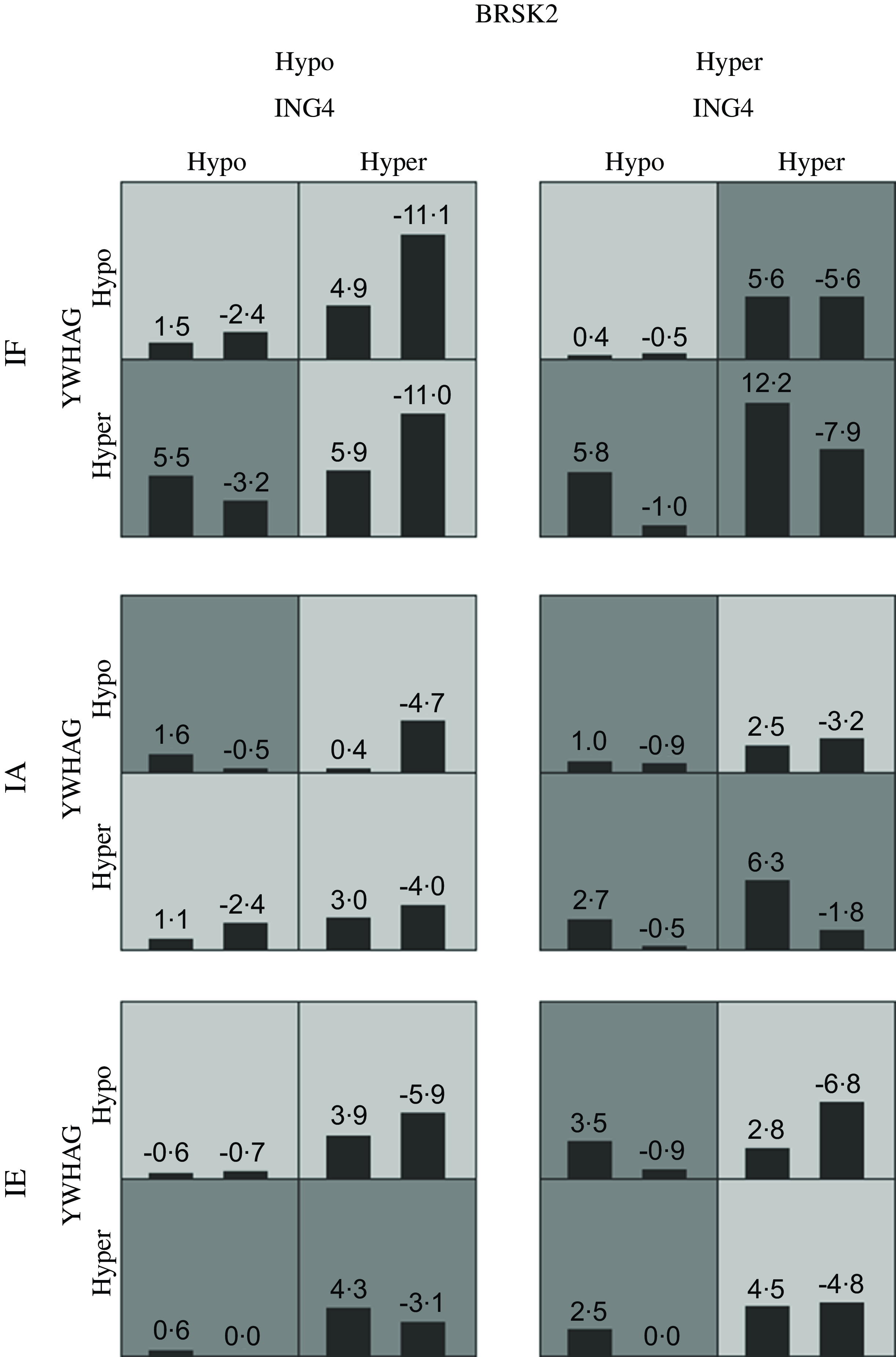
Fig. 3. The best adjusted GMDR model for gene–environment interaction. Hyper indicates hypermethylation; Hypo indicates hypomethylation; IF, iodine fortification; IA, iodine adequate; IE, iodine excessive; The adjusted covariates included age, gender, BMI, smoking, drinking thyroid function and family history of thyroid disease. The best model is composed of iodine levels, YWHAG_1, ING4_1 and BRSK2_1. In each cell, the left bar represents a positive score, and the right bar represents a negative score. High-risk cells are indicated by dark shading. The pattern of high-risk and low-risk cells differs across each of the different multilocus dimensions, presenting evidence of epistasis.
Relationships between DNA methylation and mRNA expression of YWHAG, ING4 and BRSK2 genes
As shown in Fig. 4, the mRNA expression levels of the YWHAG and BRSK2 genes in AIT patients were significantly lower than in controls (0·599 ± 0·215 v. 1·004 ± 0·003; 0·604 ± 0·248 v. 1·007 ± 0·004, both P < 0·001), but that for ING4 were higher (1·483 ± 0·568 v. 1·006 ± 0·003, P < 0·001) than those in healthy controls. A negative correlation was observed between DNA methylation levels and expression of the YWHAG gene (r = −0·111, P = 0·038). In supplementary Table 3, the mRNA expression of the YWHAG and BRSK2 genes in AIT patients was lower than controls in different levels of iodine, but for ING4 was higher (all P < 0·001).
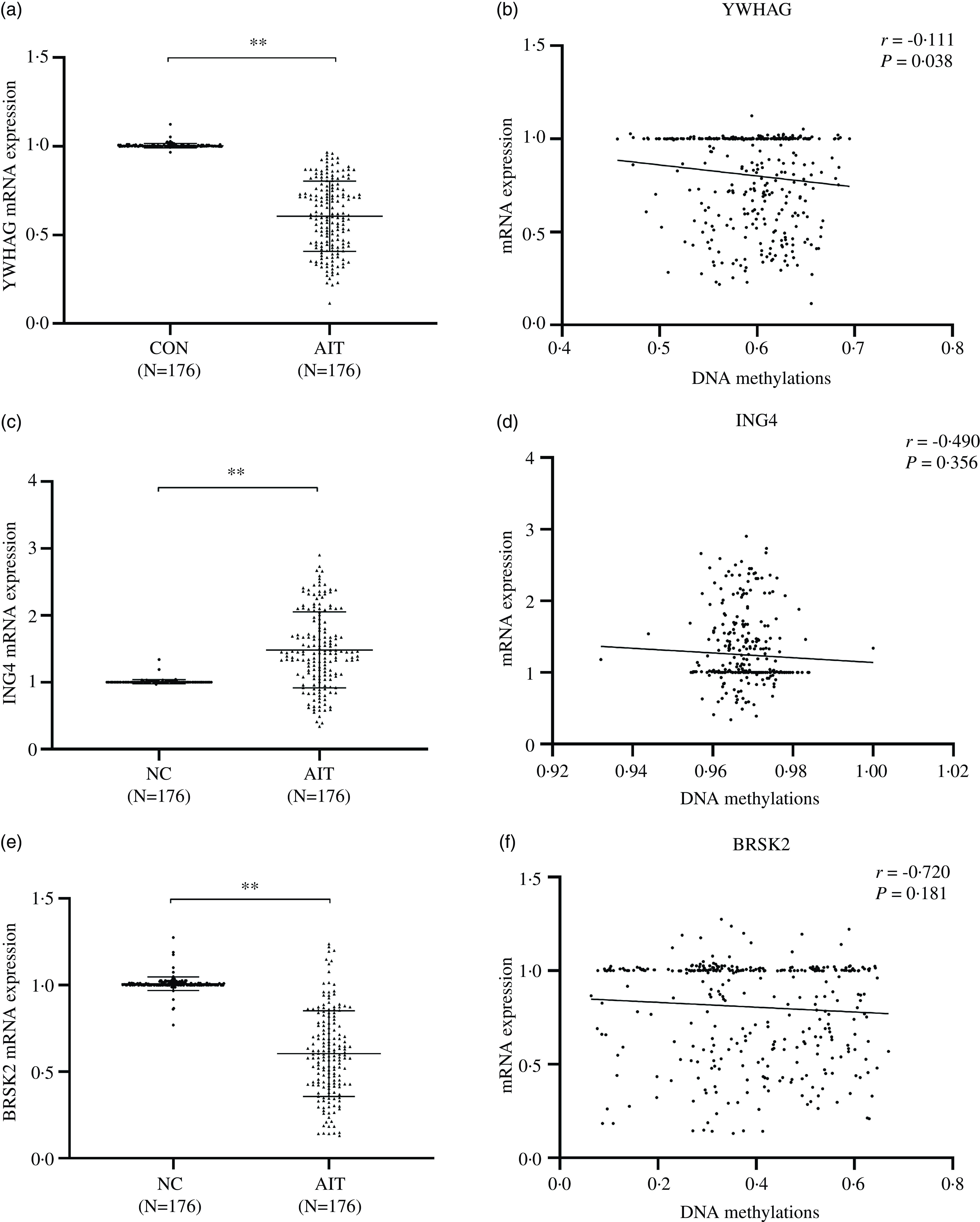
Fig. 4. Correlation analysis between DNA methylation and relative mRNA expression of YWHAG, ING4 and BRSK2 genes. (a) YWHAG mRNA expression; (b) Correlation between DNA methylation and relative mRNA expression of YWHAG_1; (c) ING4 mRNA expression; (d) Correlation between DNA methylation and relative mRNA expression of ING4_1; (e) BRSK2 mRNA expression; (f) Correlation between DNA methylation and relative mRNA expression of BRSK2_1.
Discussion
In this study, firstly, we revealed the status of DNA methylation of apoptosis-associated genes in AIT patients and analysed the relationship between mRNA expression and DNA methylation. Secondly, we investigated whether iodine nutrition could affect DNA methylation and plays a potential role in the development of AIT. Our study not only demonstrated that iodine as a nutrient could induce epigenetic changes in the body and affect thyroid health but also screened out potential candidate genes and CpG sites that may be affected by iodine. Our research provides a theoretical basis for future nutrition research.
YWHAG (14–3–3γ) regulates diverse cellular processes, including apoptosis and cell proliferation(Reference Aitken21). Two interaction points exist between the YWHAG gene and the apoptotic machinery. Signals arriving from growth factor receptors activate kinases (e.g., PI3K and Akt), which phosphorylate several intrinsic proapoptotic factors (e.g., Bad and FOXO1), and consequently promote the binding of YWHAG proteins and inhibit intrinsic apoptosis. In contrast, pro-death signals activate the kinase JNK, among others, which phosphorylates YWHAG isoforms and induces the release of intrinsic proapoptotic client proteins (e.g., Bax and Bim), thereby triggering intrinsic apoptosis(Reference Jin, Smith and Stark22,Reference Kuzelová, Grebenová and Pluskalová23) . In our study, the YWHAG gene was hypermethylated, and the expression of mRNA was diminished in patients with AIT. We speculate that in AIT patients, the high methylation levels of the target region and CpG sites of the YWHAG gene led to the lower expression, thus affecting the combination of YWHAG and the intrinsic proapoptotic factors, and ultimately intrinsic apoptosis. Nevertheless, in the future, more studies are needed to confirm this process.
Some research has shown that ING4 plays important roles in many biological processes, including DNA damage, cell proliferation and apoptosis(Reference Garkavtsev, Kozin and Chernova24). It is a well-defined tumour suppressor in cancers such as lung cancer, breast cancer and glioma(Reference Wang, Li and Zhang25,Reference Byron, Min and Thal26) . Researchers treated thyroid cancer cells with recombinant ING4 protein and the results indicate that the rate of intrinsic apoptosis in thyroid cancer cells increased significantly, while the migration ability of thyroid cancer cells was inhibited(Reference Wang, Yang and Luo27). ING4 is also thought to enhance the intrinsic apoptosis of human lung adenocarcinoma cells by activating the mitochondrial apoptotic pathway(Reference Li, Zhang and Cai28). To our knowledge, little research explores the association between ING4 gene methylation and AIT. Even so, the results of our study are consistent with results on the role of ING4 in thyroid cancer cells and the pathogenesis of other tumours. Our results showed that the methylation level of CpG site 153 in ING4 significantly differed between cases and controls. According to the biological role of ING4 in intrinsic apoptosis, our results suggest that the ING4 gene may affect AIT by intrinsic apoptosis pathway; however, the related details must be further studied.
BRSK2 is a serine/threonine kinase in the AMPK family(Reference Terai, Hiramoto and Masaki29). Endoplasmic reticulum stress can regulate the levels of BRSK2 protein and then participates in intrinsic apoptosis induced by endoplasmic reticulum stress(Reference Wang, Wan and Li30). Under the conditions of AIT, the function of the endoplasmic reticulum in sustaining proteostasis is perturbed, thus, leading to endoplasmic reticulum stress and intrinsic apoptosis. When BRSK2 is knocked down, the expression of CHOP and caspase-3 increases, thus, eventually enhancing endoplasmic reticulum stress-mediated intrinsic apoptosis in cells. These findings suggest that the effects of BRSK2 on intrinsic apoptosis under endoplasmic reticulum stress may be mediated through caspase-3. Therefore, this study is the first to investigate the correlation between BRSK2 methylation and AIT. The results showed that DNA methylation levels of the target region BRSK2_1 and eight CpG sites in AIT patients were significantly higher than those in controls, while the mRNA expression levels were significantly lower than those in controls. We hypothesised that BRSK2 hypermethylation might silence BRSK2 gene expression, thereby affecting intrinsic apoptosis induced by endoplasmic reticulum stress in AIT. However, all these hypotheses must be further studied.
We illustrate the effects of the methylation status of YWHAG, ING4 and BRSK2 on various signalling pathways related to apoptosis in Fig. 5. Signals from growth factor receptors activate PKA and Akt. PKA phosphorylates BRSK2. On the one hand, the phosphorylated BRSK2 can be combined with YWHAG and fixed in the cytoplasm(Reference Bright, Thornton and Carling31). On the other hand, both endoplasmic reticulum and BRSK2 hypermethylation can significantly downregulate the expression of BRSK2, followed by up-regulation of CHOP and ultimately leading to the up-regulation of Caspase 3 and apoptosis(Reference Wang, Wan and Li30). PI3K-Akt can phosphorylate Bad protein. When YWHAG is normally expressed, it will be combined with the phosphorylated Bad and inactivate the Bad. In our study, YWHAG is hypermethylated, which leads to a decrease in its expression level and its ability to combine with Bad. After that, uncombined Bad will play a proapoptotic role on the mitochondrial membrane(Reference Bretz and Baker32). Meanwhile, the low expression of YWHAG could not be effectively combined with the phosphorylated FOXO1, so FOXO1 could continue to play a proapoptotic role in the following process(Reference Yan, Lavin and Moser33). The death signal activates JNK and then breaks up the combination of phosphorylated BAX and YWHAG. After the separation, BAX is dephosphorylated, and the YWHAG is phosphorylated. The dephosphorylated BAX promotes cytochrome C release at the mitochondrial membrane and promotes apoptosis(Reference Nomura, Shimizu and Sugiyama34,Reference Tsuruta, Sunayama and Mori35) . JNK can also activate p53. When ING4 is hypomethylated, the overexpression of ING4 will combine with p53 and cause apoptosis. The overexpression of ING4 can also promote the expression of Bax and inhibit the expression of BCL2, resulting in intrinsic apoptosis(Reference Cai, Li and Zheng36). The above mechanism will be verified by relevant molecular biology experiments in our follow-up work.
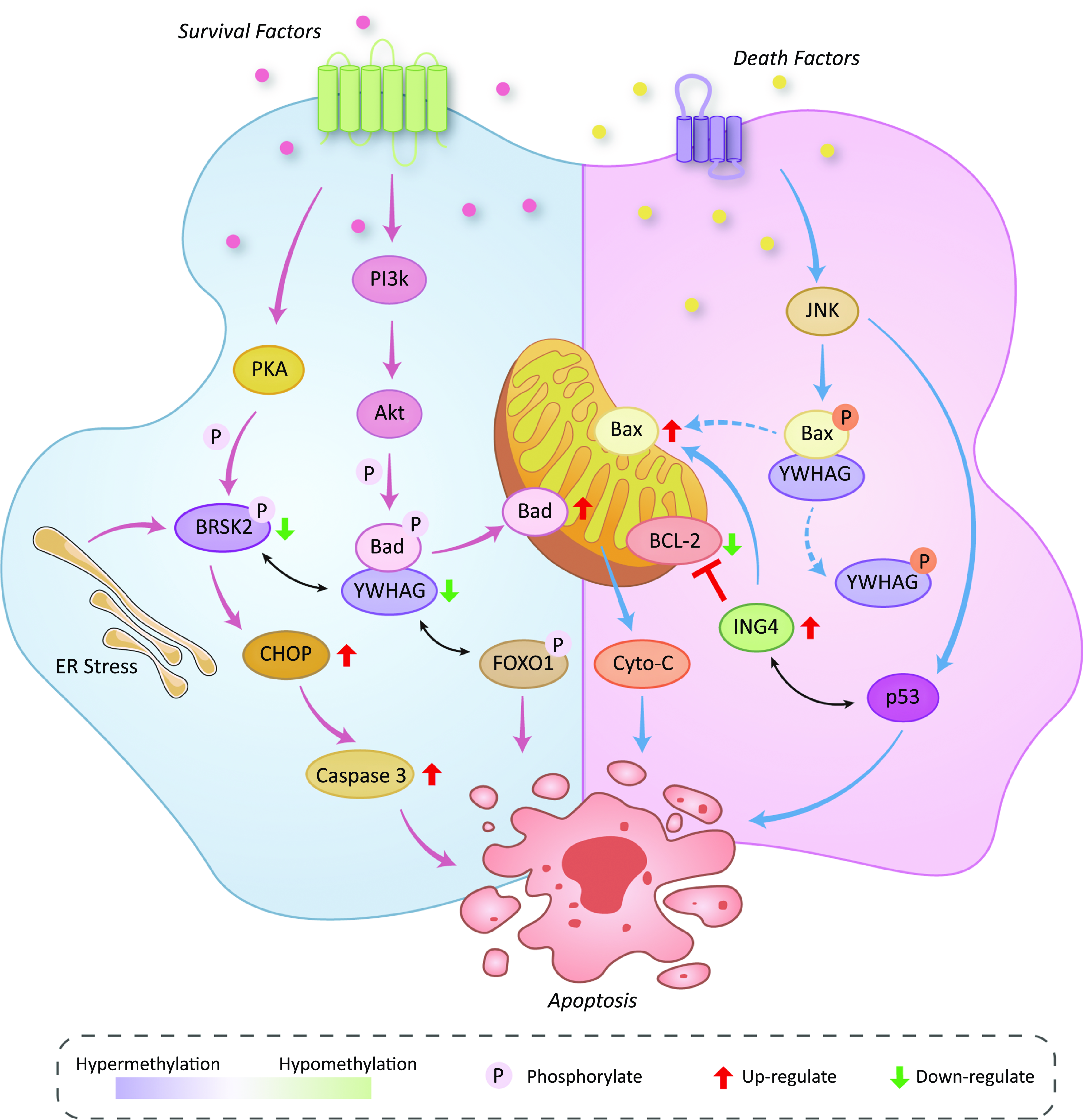
Fig. 5. Effects of the methylation status of YWHAG, ING4 and BRSK2 on various signaling pathways related to apoptosis.
Above, we analysed the relationship between methylation of our candidate genes and AIT and elaborated on the related mechanism. Next, we will discuss the effects of different iodine levels on the DNA methylation of candidate genes. In this study, the iodine levels mainly refer to the water iodine content and the use of iodised salt in the population because these are the main source of iodine nutrition for Chinese residents, dietary iodine and biomarker iodine were not discussed in this study. We found that according to recommendations from WHO/UNICEF/IGN, the iodine nutritional status of IF and IA were both adequate, but the difference in DNA methylation levels of the candidate genes (target and site) between the AIT cases and the control mainly occurred with the status of IF but not IA. Many studies have shown that after IF(Reference Pedersen, Knudsen and Jørgensen37,Reference Wang, Li and Teng38) , the status of thyroid autoimmune among some residents could change. Our results indicated that IF after iodine deficiency is more likely to alter the methylation levels of YWHAG, BRSK2 and ING4. At the same time, the universal salt iodisation could be further fine-tuned and start with small doses of iodine in the future to reduce the impact of IF after iodine deficiency on epigenetics. Patients in IE were exposed to high water iodine content for a long time, and according to recommendations from WHO/UNICEF/IGN, their iodine nutritional status was IE. But there were no differences in the methylation levels of candidate genes between AIT cases and controls. Our results indicated that the effect of IE on these candidate genes is relatively weak. In addition, we also explored the effects of iodine nutrition on the DNA methylation levels of target regions through UIC and found a negative correlation between DNA methylation levels of YWHAG, BRSK2 and UIC, and this further indicates that the methylation levels of YWHAG and BRSK2 are more likely to be high when the iodine nutritional status is deficient. Over recent years, there has been growing evidence showing that the effects of nutrition on health could be mediated by DNA methylation(Reference McKay and Mathers39). On the one hand, nutrients can change the level of global DNA methylation. For example, folic acid and vitamin B12 could significantly increase the levels of global DNA methylation, whereas selenium, bioflavonoids and green tee polyphenols could significantly decrease the levels of global DNA methylation(Reference Davis, Uthus and Finley40,Reference Pufulete, Al-Ghnaniem and Khushal41) . On the other hand, many studies have found that some specific methylation sites are closely related to the nutritional status of the body; for example, the use of folic acid in the perimenopausal period can significantly increase methylation levels in the promoter region of the IGF2 gene(Reference Steegers-Theunissen, Obermann-Borst and Kremer42). There are few studies on iodine nutrition and DNA methylation. One study in the elderly showed that iodine could not modulate the global DNA methylation profile of leukocytes(Reference Passador, Toffoli and Fernandes43), and another study in animals showed that IE did not change the global methylation levels in lymphocytes(Reference Guo, Wu and Fan44). Our previous studies have shown that different iodine levels could affect the methylation patterns of DAPK1 and ITGA6 (Reference Qu, Wan and Wu45,Reference Ren, Wan and Wu46) . This study further confirms that iodine status could affect the methylation levels of intrinsic apoptosis genes.
Based on the above findings, we further investigated whether combined and interactive effects might exist among different iodine levels and the DNA methylation levels of these genes, to explore the potential interaction between iodine nutrition and epigenetics. In this study, IF combined with YWHAG_1 hypermethylation and BRSK2_1 hypermethylation significantly increased the risk of AIT. These findings suggest that IF not only influences the methylation pattern of intrinsic apoptosis-associated genes but also interacts with the methylation levels of these genes and may ultimately increase the risk of AIT. In the occurrence or development of AIT, this IF–methylation interaction model may play an important role. We analysed the single interactions between each gene and different levels of iodine, but the four genes are all associated with intrinsic apoptosis regulation, and the result of one gene–iodine level interaction could be influenced by other genes. So, we structured gene–environment interaction models by the GMDR method. We found a four-order high–dimensional interaction among YWHAG, ING4 and BRSK2 methylation and iodine levels, which strongly contributed to the risk of AIT. These epidemiological findings suggest that the methylation of the intrinsic apoptosis gene is associated with AIT.
Studies have shown that there is a positive relationship between the prevalence of thyroid antibodies and age(Reference Zhou, Liu and Jin47). Methylation has also been shown to be associated with ageing processes. However, few studies have assessed the relationship between intrinsic apoptosis-associated gene methylation and age in patients with AIT. In this study, we explored whether changes in the methylation of intrinsic apoptosis-associated genes might be correlated with age. Older age was associated with increased methylation levels of YWHAG and BRSK2. Thus, in older AIT patients, the changes in methylation in these two genes deserve more attention. In addition, we discussed the relationship between intrinsic apoptosis-associated gene methylation levels and thyroid function. Some studies have found that interindividual variations in FT4, FT3 and TSH are influenced by genetic factors(Reference Samollow, Perez and Kammerer48). Animal studies have shown that thyroid hormones affect DNA methylation, such as T3, which leads to a significant reduction in DNA methylation(Reference Buisine, Grimaldi and Jonchere49). An Australian cohort study has described six CpG sites associated with increased levels of FT3 and two CpG sites associated with increased levels of TSH(Reference Lafontaine, Campbell and Castillo-Fernandez50). There are few studies on the association between the levels of DNA methylation in intrinsic apoptosis-associated genes and thyroid function in AIT patients. A positive correlation was found between the methylation level of YWHAG_1 and BRSK2_1 and TSH and a negative correlation with FT4. We hypothesised that the methylation levels of intrinsic apoptosis-associated genes might affect thyroid function and predict changes in thyroid function in AIT. However, these possibilities require further verification.
There are some following strengths and limitations in our study. First, we not only analysed levels of DNA methylation in intrinsic apoptosis-associated genes in different levels of iodine, which have been rarely studied in the past but also comprehensively analysed gene–environment interactions and proposed the calculational model for assessing the interactions between iodine exposure and DNA methylation. Second, the intrinsic apoptosis-associated genes in our study were novel differentially methylated genes, which have rarely been reported before. However, the present study also had several limitations. First, this study was a case–control study, and we could not get the past methylation data of the participants, especially those who have experienced iodine supplementation. Second, the relationship between intrinsic apoptosis gene methylation and increased risk of AIT was based on the results of population epidemiological studies, but we did not study the molecular functional mechanism in depth because of the difficulty in obtaining thyroid tissues from patients with AIT. Thus, more experiments in animals, such as the NOD. H2 h4 model and more molecular mechanism research are needed to extend the results of this study in the future.
In conclusion, our study showed that changes in methylation levels and mRNA expression in the YWHAG, ING4 and BRSK2 genes are associated with AIT. IF not only affects the methylation levels of YWHAG and BRSK2 but also interacts with the methylation levels of these genes and may ultimately increase the risk of AIT. A four-order high–dimensional interaction among YWHAG, ING4 and BRSK2 methylation and iodine levels appear to contribute to the risk of AIT strongly. These findings suggest a mechanism in which intrinsic apoptosis-associated gene methylation plays a critical role in AIT.
Acknowledgments
We would like to thank the Institute of Endemic Disease Control of Shandong for their assistance in collecting samples and data. Thanks to all the volunteers who participated and donated samples to this study.
The original data generated during the current study are available in the SequenceRead Archive (SRA) repository, the Accession ID is PRJNA876118. https://www.ncbi.nlm.nih.gov/bioproject/PRJNA876118.
This study was supported by grants from the National Natural Science Foundation of China (82073490).
The contribution of each author is as follows: H. S. designed the study; H. S., Z. Z., M. J., B. L., Y. H., L. L., B. R., J. L., F. L., J. L., Y. C., S. W. conducted the research. Z. Z. analysed the data. Z. Z. drafted the manuscript. All authors revised the report and approved the final version before submission.
The authors declare that no conflict of interest that could be perceived as prejudicing the impartiality of this study.
Supplementary material
For supplementary material/s referred to in this article, please visit https://doi.org/10.1017/S0007114523001216
















