At a population level, generic dietary advice is provided using a ‘one-size-fits-all’ approach on the basis of requirements for population groups( Reference de Roos 1 ), which ignores inter-individual differences, and therefore nutrient requirements. In addition, individuals’ responses to dietary interventions can be highly variable( Reference van Ommen, Keijer and Kleemann 2 – Reference Zeevi, Korem and Zmora 4 ). Demographic characteristics such as sex and age, and factors such as adiposity, physical activity, metabolic profile and genetic factors contribute to this variation( Reference Lampe, Navarro and Hullar 5 ). This phenomenon is well recognised in the medical field with a current emphasis on precision medicine( Reference Schork 6 ). Considering the reported variation in response to dietary interventions, there is now an emerging recognition that this should be considered in the development of personalised or precision nutrition( Reference Brennan 7 , Reference Kaput and Morine 8 ). Personalised nutrition or dietary advice that has been tailored for an individual offers the possibility of improving health and reducing risk of diet-related diseases( Reference Celis-Morales, Lara and Mathers 9 ). Many studies suggest that tailored dietary advice is more effective than generic advice, promoting greater improvements in dietary behaviours and related health outcomes such as body weight( Reference Celis-Morales, Lara and Mathers 9 , Reference Curtis, Adamson and Mathers 10 ). A recent meta-analysis reported that personalised interventions were more effective than non-personalised advice, with participants receiving the personalised intervention reducing body weight by 1·8 kg more on average than those receiving the non-personalised advice( Reference Celis-Morales, Lara and Mathers 9 ). However, these studies have not taken individual variability into account, and in the long term the effectiveness of the personalised dietary advice will depend on the ability to tailor advice taking into account knowledge about an individual’s potential response to the intervention( Reference Ryan, O’Donovan and Forster 11 ).
The concept of using metabolic profiles to identify responders to dietary interventions is relatively new( Reference Brennan 7 ). However, a number of examples exist in the literature demonstrating the potential of such an approach. O’Sullivan et al. ( Reference O’Sullivan, Gibney and Connor 12 ) used k-means cluster analysis to identify responders and non-responders to a vitamin D intervention. van Bochove et al. ( Reference van Bochove, van Schalkwijk and Parnell 13 ) applied k-means clustering to lipoprotein profiles and identified three clusters, two of which responded positively to fenofibrate, whereas Elnenaei et al.( Reference Elnenaei, Chandra and Mangion 14 ) identified responders and non-responders to vitamin D and Ca supplementation, on the basis of a baseline metabolomic profile. Metabolomic and transcriptomic profiles have also been used to discriminate between responders and non-responders to an n-3 PUFA supplementation( Reference Rudkowska, Paradis and Thifault 15 ). The objective of this study was to investigate differences in the phenotype and in particular blood lipids of responders and non-responders to personalised nutrition, with a specific focus on changes in circulating cholesterol levels. Using data from the Food4Me personalised dietary intervention study, individuals with borderline high baseline total cholesterol (>5 mmol/l) were examined for factors that predict their response to the intervention.
Methods
Subjects were participants of the Food4Me study – a 6-month, web-based randomised controlled trial conducted in seven European countries. The aim of the present study was to determine whether providing personalised dietary advice leads to improvements in dietary intakes and health outcomes relative to population-based public health messages. The 1607 adult subjects were randomly assigned to one of four intervention treatment groups – level 0 (standard non-personalised dietary and physical activity guidelines), level 1 (personalised advice based on current diet and physical activity), level 2 (personalised advice based on current diet, physical activity and phenotype) and level 3 (personalised advice based on current diet, physical activity, phenotype and genotype)( Reference Celis-Morales, Livingstone and Marsaux 16 ). The control group received conventional, non-personalised advice, and therefore were not considered for this analysis. The study protocol is detailed in the study by Celis-Morales et al. ( Reference Celis-Morales, Livingstone and Marsaux 16 ).
All data were collected remotely following standardised operating procedures. At baseline, participants received study kits by post containing all necessary materials to perform measurements at home. Printed instructions were included, and demonstration videos were available on the Food4Me website (http://www.food4me.org). Following measurements at baseline and 3 months, participants received a personalised report. The personalised feedback provided was based on a predefined set of algorithms, including anthropometric, physical activity (levels 1–3), phenotypic (levels 2 and 3) and genotypic (level 3 only) data( Reference Celis-Morales, Livingstone and Marsaux 16 ).
Demographic characteristics
The measurement of characteristics including age, country and sex and have been described elsewhere( Reference Celis-Morales, Livingstone and Marsaux 16 ). Having excluded the control group and those with normal total cholesterol levels at baseline (total cholesterol<5 mmol/l), there were 151 males and 162 females, with a mean age of 46·8 years from seven European countries – Germany (n 67), Greece (n 48), Ireland (n 39), the Netherlands (n 54), Poland (n 30), Spain (n 43) and the UK (n 32). Subjects were classified as responders and non-responders on the basis of the change in blood cholesterol from baseline to month 6. To achieve this, the subjects were first stratified into quartiles on the basis of cholesterol response; two of the groups, the lower and upper quartiles, were defined as responders and non-responders, respectively. This resulted in seventy-eight responders and seventy-nine non-responders.
Anthropometric measurements
Body weight, height and waist circumference were self-measured and self-reported by participants via the Internet, as described previously( Reference Celis-Morales, Livingstone and Marsaux 16 ). They were provided with clear instructions in text and video format to facilitate accurate measurements, and a validation study demonstrated the reliability of these Internet-based, self-reported anthropometric data( Reference Celis-Morales, Livingstone and Woolhead 17 ). Waist circumference was measured at the mid-point between the lower rib and the iliac crest using the same tape measure. Physical activity was self-reported using the Baecke questionnaire online( Reference Baecke, Burema and Frijters 18 , Reference Marsaux, Celis-Morales and Livingstone 19 ) on the basis of physical activity during the last month. Physical activity level scores were calculated at baseline and month 6, according to the questionnaire protocol.
Dietary intake measurements
Habitual dietary intake was quantified using an online FFQ including food items frequently consumed in each of the seven recruitment sites. The Food4me FFQ has been compared with a paper-based FFQ( Reference Forster, Fallaize and Gallagher 20 ) and 4-d weighed food record( Reference Fallaize, Forster and Macready 21 ) for both food group and nutrient intakes. The Bland–Altman analysis showed good agreement between the online and the paper-based FFQ for both nutrient and food groups level. Cross-classification into exact plus adjacent quartiles ranged from 77 to 97 % at the nutrient level and 77 to 99 % at the food group level. For comparison with the weighed food record, the mean cross-classification into exact agreement plus adjacent was 80 and 78 % for nutrient and food groups, respectively. Importantly, the energy intake estimated by the FFQ was in agreement with the weighed food record, indicating that overall the online FFQ was a suitable tool for assessing dietary intake.
Fatty acid and carotenoid profiles
Finger-prick blood samples were collected by participants using a test kit provided by Vitas Ltd, as described previously( Reference Hoeller, Baur and Roos 22 ). Each participant filled two Dry Blood Spot cards (equivalent to five drops of blood or 150 μl of blood per card) at each collection time point. The samples were sent to Vitas (Vitas Ltd) for measurements of total cholesterol, carotenoids and thirty-two fatty acids (FA). The n-3 FA index was calculated as the sum of EPA (20 : 5n-3) and DHA (22 : 6n-3). The Δ5 desaturase index (D5D) and Δ6 desaturase index (D6D) were calculated on the basis of key enzymes in the metabolism of PUFA. The D5D was calculated as the ratio of arachidonic acid (20 : 4n-6):dihomo-γ-linoleic acid (20 : 3n-6), and the D6D was calculated as the ratio of dihomo-γ-linoleic acid (20 : 3n-6):linoleic acid (18 : 2n-6).
Ethics
This study was conducted according to the guidelines laid down in the Declaration of Helsinki. The Research Ethical Committees at each participating centre granted ethics approval for the study( Reference Celis-Morales, Livingstone and Marsaux 16 ).
Statistical analysis
The baseline demographic and phenotypic characteristics of responders and non-responders were compared using generalised linear models (GLM). Models were fitted using the GLM (for continuous variables) and GENMOD (a procedure for fitting GLM; for categorical variables) procedures in SAS 9.3 (SAS Institute). To account for multiple comparisons, false discovery rate-adjusted P values are presented for FA profile data.
To assess whether baseline demographic or phenotypic characteristics can discriminate between responders and non-responders, a step-wise logistic regression procedure was applied in four stages. First, only anthropometric characteristics were included (model 1). Next, baseline cholesterol value was added to the model (model 2). Third, dietary intake data were added to the analysis (model 3), and, finally, all demographic, anthropometric, dietary intake and biochemical characteristics were included (model 4). At each stage, the step-wise procedure selected the characteristics that best discriminated between the two groups. Variables were tested using a bootstrapping approach to correct for over-optimism in model fitting. The ability of the models to classify responders and non-responders was assessed using area under the ROC curves. ROC comparisons were performed using a contrast matrix to calculate differences of the areas under the empirical ROC curves.
Results
Characteristics of responders and non-responders
Demographic characteristics did not differ significantly between responder and non-responder groups by country (χ 6 2=5·0, P=0·544, Table 1), sex (χ 1 2=0·16, P=0·693, Table 1) or age (P=0·082, Table 1). There was also little difference between responder and non-responder groups with respect to anthropometric characteristics measured at baseline (Table 1).
Table 1 Demographic and phenotypic profiles of responders and non-respondersFootnote † (Numbers and percentages; measurements at baseline and mean change (Δ) between baseline and month 6 are presented as means with their standard errors)

* P values are significant at the 5 % level.
† P values were obtained from generalised linear models including the responder group as a factor.
During the intervention period, both groups significantly reduced BMI, weight and waist circumference, with both groups exhibiting similar effect sizes (Table 1). The responders significantly increased their blood omega-3 index while the non-responders did not (mean change Δ=0·31 v. 0·14, P<0·001).
At baseline, the responders and non-responders had similar dietary intakes of most food groups, with the exception of alcohol (Table 2), for which the responders had lower intakes (170 v. 258 g/d, P=0·035). After intervention, the responders reported reduced intake of dairy products (Δ=−59 g/d, Table 2), and both responders and non-responders reported significantly reduced red meat intake (Δ=−31 and −28 g/d, respectively).
Table 2 Baseline dietary intake (g/d) and change from baseline to month 6 for responders and non-respondersFootnote † (Dietary intake at baseline and mean change (Δ) between baseline and month 6 are presented as means with their standard errors)
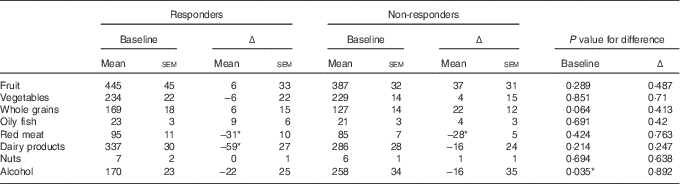
* Mean changes are significant at the 5 % level.
† P values were obtained from generalised linear models including responder group as a factor.
The percentage of participants receiving dietary advice for specific target nutrients was broadly similar (online Supplementary Table S1). The most common nutrient targeted at baseline was salt (73 % of responders and 59 % of non-responders). There was no difference in the percentage of responders and non-responders receiving a dietary message specifically targeted at cholesterol (24 v. 23 %, P=0·816), although a greater number of non-responders received a message to increase physical activity (56 % of responders v. 73 % of non-responders, P=0·027). Although the responders had a significant reduction in cholesterol, there was no significant change in physical activity during the intervention period for either group.
At baseline, the responders had higher total cholesterol levels than the non-responders (6·09 v. 5·54 mmol/l, P<0·001, Table 1). The FA profiles differed between the responders and the non-responders at baseline (Table 3). There was no difference between the groups for total SFA (P=0·203), but the responders had lower palmitic acid (16 : 0, P=0·009). At baseline, the responders had significantly lower total MUFA (P=0·016), particularly lower palmitoleic acid (16 : 1n-7, P=0·012) and cis-vaccenic acid (18 : 1n-7, P=0·001). At baseline, the responders had higher total PUFA (P=0·008), particularly linoleic acid (18 : 2n-6, P=0·011), eicosadienoic acid (20 : 2n-6, P=0·006) and DPA (22 : 5n-3, P=0·014). At baseline, both groups had similar carotenoids profiles (Table 4).
Table 3 Mean percentage of blood total fatty acid at baseline for responders and non-responders and mean change from baseline to month 6Footnote † (Fatty acid percentage at baseline and mean change (Δ) between baseline and month 6 are presented as mean values with their standard errors)

ALA, α-linolenic; GLA, γ-linolenic acid; DGLA, dihomo-γ-linolenic; ARA, arachidonic acid; FDR, false discovery rate.
* P values are significant at the FDR 5 % level.
† P values were obtained from generalised linear models including the responder group as a factor. FDR-adjusted P values control for FDR. The Δ5 desaturase was calculated as the ratio of arachidonic acid (20 : 4n-6):dihomo-γ-linoleic acid (20 : 3n-6). The Δ6 desaturase was calculated as the ratio of dihomo-γ-linoleic acid (20 : 3n-6):linoleic acid (18 : 2n-6).
Table 4 Mean blood carotenoid levels (μmol/l) for responders and non-responders at baselineFootnote † (Carotenoid levels at baseline and mean change (Δ) between baseline and month 6 are presented as means with their standard errors)
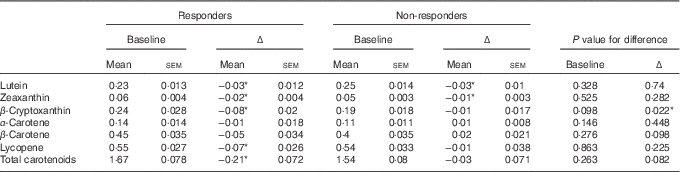
* P values are significant at the 5 % level.
† P values were obtained from generalised linear models containing the responder group as a factor.
Discriminating between responders and non-responders
When the step-wise logistic regression model was applied using demographic and anthropometric data, age and weight were selected as being important factors in discriminating responders from non-responders (model 1, Table 5). The classification accuracy (as measured by the area under the ROC curve, Fig. 1) was 0·61, indicating that the demographic and anthropometric data do not provide sufficient discriminatory power. As expected, classification accuracy improved when the model was adjusted for baseline cholesterol (model 2, AUC=0·76, Table 5, Fig. 1). Including dietary intake data (model 3) did not improve the discriminatory power, with none of the food groups being selected when tested in the step-wise model. When the additional biochemical data were added to the model (model 4), the key variables selected were baseline levels of cholesterol, glucose, stearic acid, DPA and eicosenoic acid, each with significant positive coefficients, and EPA and trans-FA, each with significant negative coefficients. Alcohol intake also had a significant negative coefficient in this model that included the biochemical variables. The coefficients of the final logistic regression discriminant model are detailed in Table 6. Increases in the variables with positive or negative coefficients were associated with increased or decreased probability of being a responder, respectively. The additional biochemical data significantly improved the classification accuracy (model 4 AUC=0·90, Table 5, Fig. 1), with increases in the true-positive rate (sensitivity) resulting in only a small trade-off with the false-positive rate (1 – sensitivity). For example, to achieve a sensitivity of 80 % in model 3, the false-positive rate was only 10 %. This compares with 67 % for model 1 and 44 % for model 2 (Fig. 1). Furthermore, it is also worth noting that the intervention group was not selected as a discriminant variable, indicating that it did not contribute to classification as a responder or non-responder.
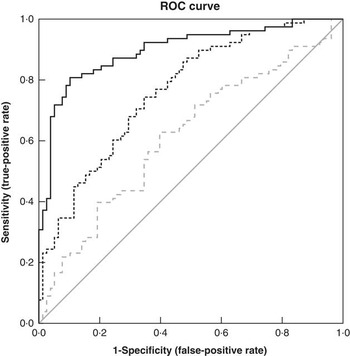
Figure 1 Area under the ROC curves illustrating the performance of models M1, M2 and M4 at discriminating responders from non-responders. The selected variables in M3 were identical to M2, and therefore have not been included. The diagonal reference line represents random discrimination, with points above the line indicating discrimination ability. ![]() , M1: anthropometric;
, M1: anthropometric; ![]() , M2: M1+baseline cholesterol;
, M2: M1+baseline cholesterol; ![]() , M4: M2+biochemical;
, M4: M2+biochemical; ![]() , reference line.
, reference line.
Table 5 Examining the ability to classify responders and non-respondersFootnote * (Area under the ROC curves with their standard errors)
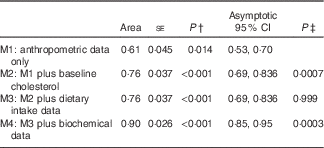
AUC, area under the ROC curve.
* The area measures the accuracy, or discrimination ability, to classify responders and non-responders.
† Null hypothesis: true area=0·5.
‡ P value for comparison of C-statistic v. previous model.
Table 6 List of discriminating parametersFootnote * (Estimates and standard errors)
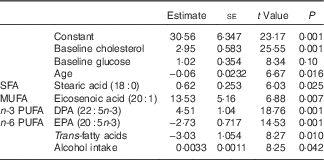
* Step-wise logistic regression discriminant analysis. Estimates are on the logit scale. This is the final model selected using step-wise selection procedure including all demographic, anthropometric and dietary intake data as well as fatty acid and carotenoid intakes as potential predictors. The logistic regression model estimates the probability of being a responder. A positive coefficient for an independent variable implies an increased probability of being a responder with increasing values of the variable.
Discussion
Identification of sub-phenotypes that respond differently to dietary interventions has the possibility to significantly enhance delivery of personalised nutrition. In the present study, a baseline phenotype characterised by age, alcohol intake, and levels of stearic acid, DPA, EPA, eicosenoic acid and trans-FA was identified, which could discriminate responders and non-responders in 90 % of cases. Discriminant analysis has previously been used in dietary intervention studies to test whether metabolic profiles may be used to identify responders and non-responders. In a choline-depletion study, analysis of the baseline metabolomics profile predicted that participants developed liver dysfunction when deprived of dietary choline( Reference Sha, da Costa and Fischer 23 ). Mutch et al.( Reference Mutch, Temanni and Henegar 24 ) classified responders and non-responders to dietary intervention using linear discriminant analysis on a gene expression snapshot. In this study, we used a step-wise logistic regression model to select the individual factors that best classified the probability of being a responder. Incorporation of such information into dietary advice strategies has the potential to significantly enhance the success of interventions.
Wide inter-individual variation has been observed in the response of total, LDL- and HDL-cholesterol to dietary changes( Reference Masson, McNeill and Avenell 25 – Reference Jacobs, Anderson and Hannan 27 ), with little alterations in blood cholesterol for some participants despite significant changes in dietary FA pattern and cholesterol intake( Reference Cox, Mann and Sutherland 28 ). This means that, although the population response to a diet can be estimated, the responsiveness a single individual will have as a result of dietary change is difficult to determine( Reference Denke, Adams-Huet and Nguyen 29 ). Lefevre et al. ( Reference Lefevre, Champagne and Tulley 30 ) observed that variability in the change of serum was related to anthropometric measurements including BMI, waist circumference and body fat percentage. Furthermore, there is a large body of evidence to support the genetic influence on response of plasma cholesterol to dietary interventions( Reference Masson, McNeill and Avenell 25 , Reference Qi, Durst and Schwarzfuchs 31 – Reference Wallace, Mann and Sutherland 33 ). The present study determined a profile that was responsive to dietary advice in terms of lowering cholesterol levels. Overall, this study in conjunction with the literature provides compelling evidence that individual variation and response to interventions have to be incorporated into dietary advice strategies.
The most marked differences between the responder and non-responder phenotypes were found in their baseline FA profiles. The responders had a lower mean percentage of trans-FA at baseline. Trans-FA have been found to increase LDL-cholesterol and decrease HDL-cholesterol levels( Reference Hunter 34 ). Although the responders and non-responders did not differ in their total percentage of SFA, contributions of different SFA differed. The responders had lower palmitic acid (16 : 0) and higher stearic acid (18 : 0) than the non-responders. A review comparing the risk factors for stearic acid with other SFA( Reference Hunter, Zhang and Kris-Etherton 35 ) reported that diets high in stearic acid have favourable effects on LDL-cholesterol compared with palmitic acid. However, it has also been reported that stearic acid itself has no cholesterol-enhancing effect in clinically very well controlled exchange of single FA, whereas palmitic, myristic and lauric acids have strong cholesterol-raising effects( Reference Müller, Kirkhus and Pedersen 36 ).
The responder group had lower total MUFA, particularly palmitoleic acid (16 : 1n-7) and cis-vaccenic acid (18 : 1n-7). A meta-analysis investigating the effects of MUFA on cardiovascular and diabetic risk factors observed no consistent evidence for a relationship between MUFA and total cholesterol( Reference Müller, Kirkhus and Pedersen 36 , Reference Schwingshackl and Hoffmann 37 ). The PUFA profiles differed between the responders and the non-responders, with a more marked difference in the n-6 PUFA. The responders had higher levels of linoleic acid (18 : 2n-6) and eicosadienoic acid (20 : 2n-6) compared with the non-responders at baseline. Linoleic acid, the primary n-6 PUFA, has been shown to have a cholesterol-lowering effect( Reference Müller, Kirkhus and Pedersen 36 , Reference Harris, Mozaffarian and Rimm 38 ), and a recent meta-analysis reported a lower risk of CHD events and deaths with increasing linoleic acid intake( Reference Farvid, Ding and Pan 39 ). Although the total n-3 PUFA did not differ between the two groups, the responders had a higher percentage of DPA (22 : 5n-3). Higher levels of DPA in human blood have been shown to be correlated with lower cholesterol( Reference Byelashov, Sinclair and Kaur 40 ). Overall, the data support the growing evidence that FA patterns as opposed to single individual FA are important in determining health. Moreover, it supports the importance of adequate intake of PUFA.
The demographic profiles of the responders and non-responders did not differ, and at baseline the groups also had similar anthropometric characteristics. Dietary intake at baseline was similar across the two groups, with only alcohol intake differing. As this was a study of the effects of personalised nutrition, the dietary advice given to the participants differed between individuals. However, for all the participants, the percentage of subjects receiving dietary advice for specific target nutrients was generally similar. The strengths of this study included the fact that it was a multi-country group with multiple time points allowing analysis of change in response to the intervention. Furthermore, the participants were well phenotyped. A limitation of the present study is the unique study design involving personalised nutrition advice, which makes replication and prospective analysis in an independent cohort difficult.
An objective of this study was to investigate whether the different types of data were useful in classifying whether an individual will respond to the dietary intervention. Our study has shown that baseline phenotypic data provided more classification power than anthropometric or dietary intake data in classifying responsiveness to personalised dietary advice. While the work identified particular predictive characteristics, it was not our aim to establish causative relationships between the variables. Our study has shown that, in principle, we can predict a priori whether an individual’s health status will improve in response to the consumption of a given food/diet. This strengthens the evidence base for the concept that intervention and dietary advice can be personalised with more confidence. Future work should examine the optimal method for incorporation of such data into dietary advice and should pave the way for precision nutrition.
Acknowledgements
This project was supported by the European Commission under the Food, Agriculture, Fisheries and Biotechnology Theme of the 7th Framework Programme for Research and Technological Development, grant no. 265494.
The authors’ contributions were as follows: L. K., L. B., E. R. G. and M. C. W. derived the research question for this manuscript, drafted the manuscript and conducted statistical analysis; J. C. M. was the study director of the proof-of-principle study of Food4Me; H. D., I. T., C. A. D., M. G., J. A. L., Y. M., J. A. M. and W. H. M. S. contributed to the design of the proof-of-principle study and were principal investigators for their respective research centre; L. B., R. F., H. F., E. R. G., M. G., S. K., K. M. L., C. F. M. M., C. C.-M., G. M., S. N.-C., C. B. O.’D., A. S., M. C. W. and C. W. contributed to the study design and execution at the research centres. All authors read and approved the final version of the manuscript.
C. A. D. is the founder, stock owner, board member and consultant for Vitas Ltd, Oslo, Norway. The other authors have no potential financial or personal conflicts of interest to declare.
Supplementary material
To view supplementary material for this article, please visit https://doi.org/10.1017/S0007114516004256










