91 results
Continuous approximations for optimizing allele trajectories
-
- Journal:
- Genetics Research / Volume 92 / Issue 2 / April 2010
- Published online by Cambridge University Press:
- 01 June 2010, pp. 157-166
-
- Article
-
- You have access
- HTML
- Export citation
Characterisation of linkage disequilibrium and subsequent estimation of effective population size in Thoroughbred horses using single nucleotide polymorphism markers
-
- Journal:
- Advances in Animal Biosciences / Volume 1 / Issue 1 / April 2010
- Published online by Cambridge University Press:
- 29 November 2010, p. 58
- Print publication:
- April 2010
-
- Article
-
- You have access
- Export citation
Genetic evaluation of sport horses for performance in Eventing competitions in Great Britain
-
- Journal:
- Advances in Animal Biosciences / Volume 1 / Issue 1 / April 2010
- Published online by Cambridge University Press:
- 29 November 2010, p. 59
- Print publication:
- April 2010
-
- Article
-
- You have access
- Export citation
Prediction of long-term contributions and inbreeding in populations undergoing mass selection
-
- Journal:
- Genetical Research / Volume 62 / Issue 3 / December 1993
- Published online by Cambridge University Press:
- 14 April 2009, pp. 231-242
-
- Article
-
- You have access
- Export citation
Genetic evaluation of UK sport horses for dressage competition
-
- Journal:
- Proceedings of the British Society of Animal Science / Volume 2009 / April 2009
- Published online by Cambridge University Press:
- 22 November 2017, p. 3
- Print publication:
- April 2009
-
- Article
- Export citation
Breed differences in copper metabolism in sheep
-
- Journal:
- The Journal of Agricultural Science / Volume 91 / Issue 2 / October 1978
- Published online by Cambridge University Press:
- 27 March 2009, pp. 433-441
-
- Article
- Export citation
The copper content of wool in relation to breed and the concentrations of copper in the liver and plasma
-
- Journal:
- The Journal of Agricultural Science / Volume 100 / Issue 2 / April 1983
- Published online by Cambridge University Press:
- 27 March 2009, pp. 505-507
-
- Article
- Export citation
The long-term accumulation and depletion of copper in the liver of different breeds of sheep fed diets of differing copper content
-
- Journal:
- The Journal of Agricultural Science / Volume 100 / Issue 2 / April 1983
- Published online by Cambridge University Press:
- 27 March 2009, pp. 441-449
-
- Article
- Export citation
The effect of dietary intake of calcium and dry matter on the absorption and excretion of calcium and phosphorus by growing lambs
-
- Journal:
- The Journal of Agricultural Science / Volume 105 / Issue 2 / October 1985
- Published online by Cambridge University Press:
- 27 March 2009, pp. 237-243
-
- Article
- Export citation
The effect of dietary intake of calcium and phosphorus on the absorption and excretion of phosphorus in chimaera-derived sheep
-
- Journal:
- The Journal of Agricultural Science / Volume 101 / Issue 3 / December 1983
- Published online by Cambridge University Press:
- 27 March 2009, pp. 597-602
-
- Article
- Export citation
Analysis of factors affecting superovulatory responses in ruminants
-
- Journal:
- The Journal of Agricultural Science / Volume 124 / Issue 1 / February 1995
- Published online by Cambridge University Press:
- 27 March 2009, pp. 61-70
-
- Article
- Export citation
Animal and dietary variation in the absorption and metabolism of phosphorus by sheep
-
- Journal:
- The Journal of Agricultural Science / Volume 103 / Issue 2 / October 1984
- Published online by Cambridge University Press:
- 27 March 2009, pp. 283-291
-
- Article
- Export citation
Conservation of animal genetic resources: approaches and technologies for in situ and ex situ conservation
-
- Journal:
- Animal Genetic Resources Information / Volume 42 / April 2008
- Published online by Cambridge University Press:
- 01 August 2011, pp. 71-85
- Print publication:
- April 2008
-
- Article
- Export citation
The UK National Action Plan on Farm Animal Genetic Resources
-
- Journal:
- Proceedings of the British Society of Animal Science / Volume 2007 / April 2007
- Published online by Cambridge University Press:
- 23 November 2017, p. 251
- Print publication:
- April 2007
-
- Article
- Export citation
Science of conservation for populations at risk
-
- Journal:
- Proceedings of the British Society of Animal Science / Volume 2007 / April 2007
- Published online by Cambridge University Press:
- 23 November 2017, p. 252
- Print publication:
- April 2007
-
- Article
- Export citation
Genetic correlations between the cross country, dressage and showjumping phases of eventing competition
-
- Journal:
- Proceedings of the British Society of Animal Science / Volume 2007 / April 2007
- Published online by Cambridge University Press:
- 23 November 2017, p. 61
- Print publication:
- April 2007
-
- Article
- Export citation

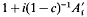 (where
(where 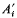 is one half the breeding value of the ancestor for the trait under selection standardized by the phenotypic standard deviation,
is one half the breeding value of the ancestor for the trait under selection standardized by the phenotypic standard deviation, 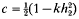 is the competitiveness which is defined by
is the competitiveness which is defined by 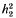 the heritability in generation 2 and
the heritability in generation 2 and