101 results
Temporal decreases in pathogen colonization and infection among hospitalized neonates following routine skin antisepsis with chlorhexidine gluconate: Botswana 2022 – 2023
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 4 / Issue S1 / July 2024
- Published online by Cambridge University Press:
- 16 September 2024, p. s12
-
- Article
-
- You have access
- Open access
- Export citation
Improving antibiotic use for community acquired pneumonia in hospitalized children through electronic feedback reports
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 4 / Issue S1 / July 2024
- Published online by Cambridge University Press:
- 16 September 2024, pp. s48-s50
-
- Article
-
- You have access
- Open access
- Export citation
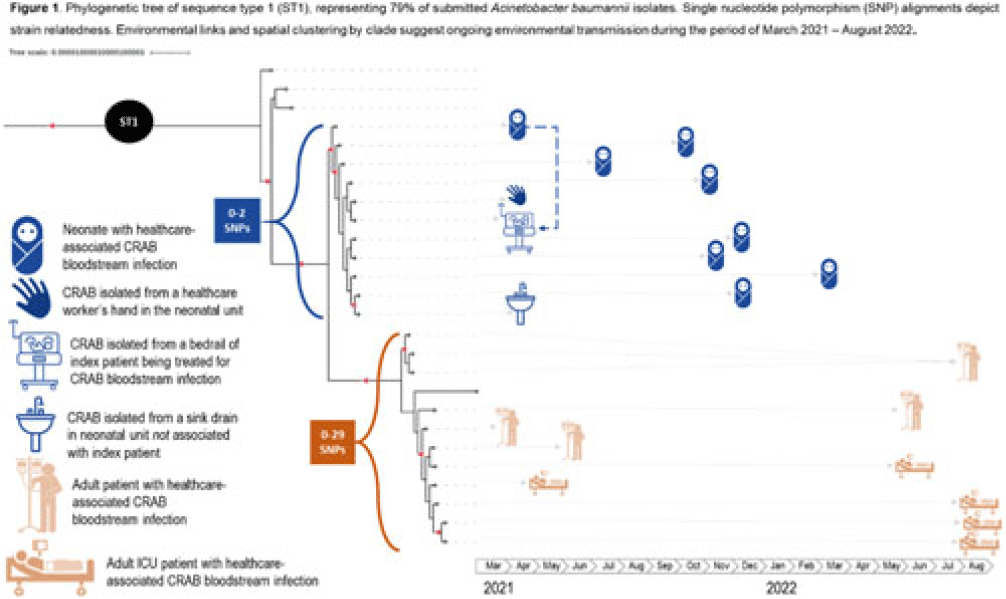
Hyperendemic carbapenem-resistant Acinetobacter baumannii at a hospital in Botswana: Insights from whole-genome sequencing
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 3 / Issue S2 / June 2023
- Published online by Cambridge University Press:
- 29 September 2023, pp. s115-s116
-
- Article
-
- You have access
- Open access
- Export citation
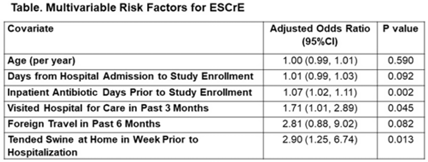
Colonization with extended-spectrum cephalosporin-resistant Enterobacterales (ESCrE) in hospitalized patients in Botswana
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 3 / Issue S2 / June 2023
- Published online by Cambridge University Press:
- 29 September 2023, p. s81
-
- Article
-
- You have access
- Open access
- Export citation
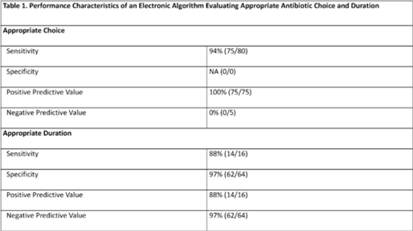
Validation of an electronic algorithm to identify appropriate antibiotic use for community-acquired pneumonia in children
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 3 / Issue S2 / June 2023
- Published online by Cambridge University Press:
- 29 September 2023, p. s106
-
- Article
-
- You have access
- Open access
- Export citation
Characterization of resistance to newer antimicrobials among carbapenem-resistant Klebsiella pneumoniae in the post–acute-care setting
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 44 / Issue 7 / July 2023
- Published online by Cambridge University Press:
- 28 July 2022, pp. 1159-1162
- Print publication:
- July 2023
-
- Article
- Export citation
Spatial and temporal effects on severe acute respiratory coronavirus virus 2 (SARS-CoV-2) contamination of the healthcare environment
- Part of
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 43 / Issue 12 / December 2022
- Published online by Cambridge University Press:
- 27 December 2021, pp. 1773-1778
- Print publication:
- December 2022
-
- Article
- Export citation
Severe acute respiratory coronavirus virus 2 (SARS-CoV-2) surface contamination in staff common areas and impact on healthcare worker infection: Prospective surveillance during the coronavirus disease 2019 (COVID-19) pandemic
- Part of
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 44 / Issue 1 / January 2023
- Published online by Cambridge University Press:
- 15 November 2021, pp. 110-113
- Print publication:
- January 2023
-
- Article
- Export citation
Healthcare microenvironments define multidrug-resistant organism persistence
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 43 / Issue 9 / September 2022
- Published online by Cambridge University Press:
- 24 August 2021, pp. 1135-1141
- Print publication:
- September 2022
-
- Article
- Export citation
The Impact of Social Role Identity on Communication in Hospital-Based Antimicrobial Stewardship
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 1 / Issue S1 / July 2021
- Published online by Cambridge University Press:
- 29 July 2021, p. s14
-
- Article
-
- You have access
- Open access
- Export citation
Urinary tract infection stewardship: A urinary antibiogram and electronic medical record alert nudging narrower-spectrum antibiotics for urinary tract infections
- Part of
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 1 / Issue 1 / 2021
- Published online by Cambridge University Press:
- 29 June 2021, e8
-
- Article
-
- You have access
- Open access
- HTML
- Export citation
Development and validation of antibiotic stewardship metrics for outpatient respiratory tract diagnoses and association of provider characteristics with inappropriate prescribing
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 43 / Issue 1 / January 2022
- Published online by Cambridge University Press:
- 02 June 2021, pp. 56-63
- Print publication:
- January 2022
-
- Article
- Export citation
Pediatric research priorities in healthcare-associated infections and antimicrobial stewardship
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 42 / Issue 5 / May 2021
- Published online by Cambridge University Press:
- 26 November 2020, pp. 519-522
- Print publication:
- May 2021
-
- Article
- Export citation
Enhanced Environmental Cleaning to Reduce Rates of Clostridioides difficile Infection on Oncology Units
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, p. s213
- Print publication:
- October 2020
-
- Article
-
- You have access
- Export citation
Impact of Diagnosed and Undiagnosed Respiratory Pseudomonas on VAP and VAE During Long-Term Acute Care
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s258-s259
- Print publication:
- October 2020
-
- Article
-
- You have access
- Export citation
Development of an Electronic Algorithm to Identify Inappropriate Antibiotic Prescribing for Pediatric Pharyngitis
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s188-s189
- Print publication:
- October 2020
-
- Article
-
- You have access
- Export citation
Impact of Removal of Automatic 7-Day Stop Orders for Inpatient Antimicrobials
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s264-s265
- Print publication:
- October 2020
-
- Article
-
- You have access
- Export citation
Development of an Electronic Algorithm to Target Outpatient Antimicrobial Stewardship Efforts for Acute Bronchitis
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, p. s32
- Print publication:
- October 2020
-
- Article
-
- You have access
- Export citation
An Interactive Sociotechnical Analysis of the Implementation of Electronic Decision Support in Antimicrobial Stewardship
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s115-s116
- Print publication:
- October 2020
-
- Article
-
- You have access
- Export citation
Comparison of Respiratory Microbiome Disruption Indices to Predict VAP and VAE risk at LTACH Admission
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s179-s180
- Print publication:
- October 2020
-
- Article
-
- You have access
- Export citation