Schizophrenia is a major cause of global disability with a lifetime risk of approximately 1%. Perhaps because of its complex genetics, identifying the genetic variation responsible for disease risk has proven difficult, despite schizophrenia’s substantial heritability (∼80%). Reference Risch1-Reference Sullivan, Kendler and Neale4 Genome-wide association studies (GWAS) have identified a small number (∼12) of common risk variants for schizophrenia. Reference O'Donovan, Craddock, Norton, Williams, Peirce and Moskvina5,Reference Stefansson, Ophoff, Steinberg, Andreassen, Cichon and Rujescu6 Recently, the Psychiatric GWAS Consortium ‘mega-analysis’ of over 51 000 cases and controls identified seven risk loci, including five novel findings. Reference Ripke, Sanders, Kendler, Levinson, Sklar and Holmans7 Of the novel loci identified, one of these (on 10q 34.33) was unique in harbouring two genome-wide significant variants, including the rs7914558 single nucleotide polymorphism (SNP) located within an intron of the cyclin M2 gene (CNNM2, OMIM: 607803 (this gene was previously known as ‘ancient conserved domain protein 2’, ACDP2)). Since, CNNM2 has been associated with both increased risk for schizophrenia in a family-based study and increased risk for psychiatric disorders generally. Reference Aberg, Liu, Bukszár, McClay, Khachane and Andreassen8,9
The cyclin M2 gene is implicated in the transport of magnesium (Mg2+), which plays a crucial role in many biological processes, including neuronal transmission. Despite the extensive evidence for unique mammalian Mg2+ transporters, CNNM2 is one of only a few proteins biochemically identified as fulfilling this role. Reference Goytain and Quamme10 Sequence variation within CNNM2 increases hypertension risk and alters serum Mg2+ blood levels, a risk factor for a variety of vascular diseases. Reference Meyer, Verwoert, Hwang, Glazer, Smith and van Rooij11 Furthermore, CNNM2 mutations are associated with hypomagnesaemia risk. Reference Stuiver, Lainez, Will, Terryn, Gunzel and Debaix12 Whether or how this biological role in Mg2+ regulation is relevant to CNNM2’s association with increased risk for schizophrenia is uncertain.
Approaches to elucidating the biological role of novel schizophrenia genetic susceptibility factors include delineating their effects on ‘intermediate’ or ‘endo-’phenotypes (i.e. discrete aspects of behaviour or brain structure and function that mediate genetic effects on the broader phenotype). Reference Tan, Callicott and Weinberger13-Reference Walters and Owen15 Neurocognitive intermediate phenotypes widely used in schizophrenia genetics include general cognitive ability, long-term and working memory, and attentional control. In addition to these more ‘traditional’ neuropsychological phenotypes, social cognition, which includes functions such as emotional recognition and perception, theory of mind and attributional style, has been a recent focus of schizophrenia gene studies. Measures of social cognition predict social and occupational function in schizophrenia independently of neurocognitive task performance and may mediate the effects of neurocognition on functional outcome. Reference Allen, Strauss, Donohue and van Kammen16 Moreover, there are data to suggest that social cognitive functions may be critically mediated by genetic factors (see Skuse & Gallagher for a review Reference Skuse and Gallagher17 ). Therefore, studying the effects of schizophrenia risk variants on social cognition may index aspects of disability distinct from traditional neurocognitive tasks. For example, although ZNF804A has not been reliably associated with general cognitive deficits, it has been associated with poorer performance on indices of social cognition, including cortical activation during a theory of mind task and increased biases in social attributions. Reference Walter, Schnell, Erk, Arnold, Kirsch and Esslinger18,Reference Hargreaves, Morris, Rose, Fahey, Moore and Cummings19 Our study considers the effects of SNP rs7914558 at CNNM2 on indices of neurocognition and social cognition in both patients and healthy individuals. We focused solely on rs7914558 because the frequency of the second schizophrenia-associated SNP at this locus (rs11191580, located within the neighbouring gene, NT5C2, with a risk allele frequency of 0.91) yielded insufficient power to examine its neurocognitive effects in our samples. We tested the hypothesis that the increased risk for schizophrenia associated with rs7914558 was mediated via deleterious effects on both these aspects of cognition. Furthermore, we sought to investigate the impact of rs7914558 on brain structure (i.e. grey and white matter volume), via delineating the mediating impact of this SNP on brain structure in regions underlying associated functions.
Method
Neuropsychological study
We assessed cognition in 400 clinically stable patients with a DSM-IV diagnosis 20 of schizophrenia or schizoaffective disorder and 160 healthy individuals (online Table DS1). Participants were recruited from five sites across Ireland, and ethics approval was obtained from local ethics committees. Written informed consent was obtained from all participants. Participants were aged 18-65 years and had confirmed Irish lineage (i.e. Irish grandparents on both sides of the family). Healthy individuals had no history of significant neurological or psychiatric conditions and no first-degree relatives with a history of psychosis. These samples represent a subset of the Irish samples included in the original study (Ripke et al Reference Ripke, Sanders, Kendler, Levinson, Sklar and Holmans7 ) identifying CNNM2 as a risk variant.
Neuropsychological test measures
Participants completed a neuropsychological battery that included measures of IQ (i.e. Wechsler Adult Intelligence Scale (WAIS-III) Reference Wechsler21 ), working and long-term memory (i.e. letter number sequencing and logical memory subtests from the Wechsler Memory Scale (WMS-III) Reference Wechsler22 ) and the Cambridge Neuropsychological Battery (CANTAB) Reference Robbins, James, Owen, Sahakian, McInees and Rabbitt23 spatial working memory task and attentional control (i.e. the CANTAB intradimensional/extradimensional set-shifting task).
Social cognition measures
Social cognitive function was assessed using the: (a) Hinting Task; Reference Corcoran, Mercer and Frith24 (b) Reading the Mind in the Eyes (‘Eyes’) Task; Reference Baron-Cohen, Wheelwright, Hill, Raste and Plumb25 and (c) Internal, Personal and Situational Attributions Questionnaire (IPSAQ). Reference Kinderman and Bentall26 The Hinting and Eyes tasks target theory of mind processes. The IPSAQ, on the other hand, is concerned with attributional style and uses subscale scores that delineate the extent of internal and external attributions for hypothetical events to calculate two indices of cognitive bias, i.e. externalising bias and personalising bias. Externalising bias scores index an individual’s tendency to attribute more positive than negative events to oneself. As this bias might be expected to have a positive effect on one’s sense of self it is often described as a ‘self-serving’ bias. Personalising bias is indicative of the tendency to credit external attributions to personal (i.e. actions or omissions of another) as opposed to situational factors (i.e. circumstance or chance). These data were collected as part of an ongoing multisite project; consequently all assessments were not available in the entire cohort (IPSAQ: n = 387 (256 patients and 131 controls); Hinting Task: n = 408 (276 patients and 132 controls); Eyes Task: n = 201 (151 patients and 50 controls)).
Structural imaging study
Irish imaging sample
Structural magnetic resonance imaging (sMRI) data were acquired from healthy individuals as part of the Trinity College Institute for Neuroscience (TCIN) biobank project (see Rose et al Reference Rose, Greene, Kelly, Morris, Robertson and Fahey27 for details). Participants were right-handed and had no history of significant neurological, psychiatric or other major medical health problems, and had no other contraindication for magnetic resonance imaging (MRI). In accordance with local ethics committee approval, written, informed consent was obtained for the use of sMRI data and participants provided a saliva sample (Oragene DNA self-collection kits: DNA Genotek, Ontario, Canada) for genetics analysis. Individuals (n = 159) were selected for imaging analysis based on sMRI data quality and successful rs7914558 genotyping. The majority were recorded as ‘Caucasian Irish’ or ‘other European’ (n = 129). Lineage was not documented in 30 participants; however, the relative homogeneity of the local population (92% of Dublin city’s population are of ‘Caucasian Irish’ background; www.cso.ie/census) suggests that these were also likely to be Caucasian Irish. Furthermore, this subgroup did not differ from other imaging participants in gender ratio, age and years of education or minor allele frequency.
Italian imaging sample
Patients with a confirmed diagnosis of schizophrenia (n = 66) were recruited consecutively from two out-patient clinics in central Italy. Patient exclusion criteria included a history of: (a) DSM-IV Axis I or II diagnosis except schizophrenia; (b) traumatic head injury with a loss of consciousness; (c) epilepsy, seizures or other relevant neurological or medical illness (for example cerebrovascular disease); and/or (d) substance misuse in the 6 months pre-participation. Healthy participants (n = 37) were recruited from the same geographical area as patients and screened for current or past diagnosis of any DSM-IV Axis I or II disorder using the SCID-I and SCID-II. Reference First, Spitzer, Gibbon and Williams28,Reference First, Gibbon, Spitzer, Williams and Benjamin29 In addition to the exclusion criteria outlined for patients, individuals with a history of schizophrenia or any other psychiatric disorder diagnosis among first-degree relatives were also excluded. All participants were of White Italian ancestry.
sMRI parameters
Irish structural images were acquired on a Philips Intera Achieva 3T MRI system, and involved the acquisition of a 180 slice T 1-weighted image using a turbo field echo gradient echo pulse sequence (repetition time TR) = 8.4 ms; echo time (TE) = 3.8 ms; flip angle, 8°; slice thickness, 0.9 mm; voxel size, 0.9 mm3; 180 slices; duration, 6 min). The Italian sMRI was conducted using a 1.5T whole-body Siemens Vision Magnetom scanner and involved the acquisition of T 1-weighted images using a magnetisation-prepared rapid gradient echo (MPRAGE) sequence (TR = 11.4 ms; TE = 4.4 ms; flip angle, 151°; slice thickness, 1 mm, no interslice gap; voxel size, 1 mm3).
Genotyping
Genetics analysis was carried out using DNA extracted from blood or saliva samples. The rs7914558 SNP was genotyped using a Taqman SNP genotyping assay on a 7900HT sequence detection system. The call rate for the Taqman genotyping was >95% and the samples were in Hardy-Weinberg equilibrium (P>0.05). Along with these samples a small number of HapMap CEU DNA samples (www.hapmap.org) were genotyped for rs7914558 for quality control purposes and were found to be concordant with available HapMap data for this SNP.
Statistical analyses
Neuropsychological data
Behavioural data were analysed in SPSS (Release 16 for Windows PC). For each measure, univariate ANOVA considered both group (patient v. control) and rs7914558 genotype (AA v. AG. v. GG) as fixed effects, and included age, gender and IQ as covariates.
Voxel-based morphometry (VBM) Reference Ashburner and Friston30
Structural MRI analysis was performed within SPM5 (www.fil.ion.ucl.ac.uk/spm) running under Matlab (v7.8; The MathWorks, Cambridge, UK) and utilised the VBM toolbox (v5.1, http://dbm.neuro.uni-hen.de/vbm). Volumes passing initial data quality control (i.e. visual inspection for scanner artifacts and gross anatomical abnormalities) were segmented into grey matter, white matter and cerebrospinal fluid, without tissue priors and using a Hidden Markov Random Field weighting of 0.15. Segmented images were normalised using DARTEL, Reference Ashburner31 in which grey matter and white matter templates were created using standard parameters. Jacobian scaled (‘modulated’) warped tissue classes were subsequently created for both grey matter and white matter for each participant and the resultant images smoothed with an 8 mm3 Gaussian kernel.
Structural MRI data were analysed using full factorial ANOVA models in SPM5. Separate analyses were carried out for the Irish and Italian imaging samples. Although the imaging sequences used at each site would be expected to produce T 1-weighted images that were relatively comparable, methodological concerns related to differences in the imaging parameters precluded combining the data-sets for a single analysis. Modelling included genotype as a fixed factor with either three (AA v. AG. v. GG) or two (i.e. risk homozygotes v. non-risk carriers) levels plus age and gender as covariates. Although the three-group analysis was designed to replicate modelling in our neuropsychological analyses, we also carried out the two-group analysis in order to address potential issues in accurately modelling the data because of the relatively small number of individuals in the homozygous minor allele group (i.e. ‘GG’). To account for normal variation in brain volume, total volume was also included as a covariate. Analysis of the Italian data-set also included group (patient v. control) as a fixed factor. In addition to preliminary whole brain analyses of grey matter and white matter, based on the outcomes of our cognitive analyses (see Results) post hoc region-of-interest (ROI) analyses considering grey matter variation in regions that support social cognitive functions (i.e. anterior cingulate cortex/BA32; superior temporal sulcus/BA22; and temporal pole/BA38) Reference Gallagher and Frith32-Reference Ochsner34 were also carried out.
Since the non-uniform smoothness of VBM data can influence the interpretation of these types of analysis, Reference Ashburner and Friston30,Reference Worsley, Andermann, Koulis, MacDonald and Evans35 prior to the determination of significance a non-stationarity cluster extent correction utilising the random field theory version of cluster inference under non-stationarity Reference Hayasaka, Phan, Liberzon, Worsley and Nichols36 was implemented using the NS toolbox for SPM5 (http://fmri.wfubmc.edu/cms/NS-General).
Results
The effects of CNNM2 rs7914558 on demographic and clinical measures
There was no genotype-related variability in gender ratio, age or years of education in any of the samples. Moreover, in patient samples, rs7914558 was not associated with age at onset, illness duration or medication (i.e. chlorpromazine equivalents) and there was no difference in genotype frequency between patients and controls. There was, however, a significant difference in age and years of education between patients and healthy participants, such that patients were older (Italian VBM sample: F (1,97) = 10.79, P = 0.001; Irish neurocognition sample: F (1,537) = 4.86, P<0.05) and had less years of education (Italian VBM sample: F (1,81) = 21.59, P<0.001; neurocognition sample: F (1,537) = 105.61, P<0.001) (Table DS1).
The effects of CNNM2 rs7914558 on cognition
General cognition
Although patients performed worse than controls on all measures of general cognitive function (P⩽0.001), there was no genotype-related variability on indices of IQ, memory or attentional control (online Table DS2).
Social cognition
In tests of theory of mind, patients performed significantly worse than healthy participants on the Eyes Task (F (2,121) = 8.11, P<0.05) but not the Hinting Task. Performance was unaffected by genotype for both measures. In contrast, differences in attributional style as measured by the IPSAQ scores were observed between genotype groups. Specifically, we observed a main effect of genotype for externalising bias scores, in the absence of any differences between patients and controls (Fig. 1). This effect was because of the linear impact of genotype on externalising bias (F (1,389) = 10.14, P<0.001), such that the non-risk ‘GG’ individuals scored highest (i.e. had greater externalising bias) on this subscale, followed by ‘AG’ and then ‘AA’ individuals (P<0.05). Although personalising bias scores differed between patients and controls (F (2,375) = 6.12, P<0.05), a main effect of genotype on this index was not observed.
The effects of CNNM2 rs7914558 on brain volume
Irish sMRI/VBM sample analysis of grey matter
There were no effects of genotype that survived correction for multiple comparisons at the whole brain level in either the three (AA v. AG v. GG) or two (AA v. AG/GG) groups. Conversely, post hoc anatomical ROI analyses showed a main effect of rs7914558 status (Fig. 2). In the three-group analysis, there was a main effect of genotype in the right temporal pole/BA38 (Montreal Neurological Institute (MNI): 40, 16, –33; F (2,154) = 7.88, number of voxels/cluster extent (K E) = 217, P corrected for multiple comparisons <0.05) and anterior cingulate cortex/BA32 (MNI: 10, 39, 19; F (2,154) = 7.26, K E = 50, P corrected<0.05). In both clusters, post hoc t-tests indicated that this main effect was because of relatively greater grey matter volume in homozygous ‘A’ individuals compared with heterozygotes (t (154) = 3.95 and t (154) = 3.77, P corrected<0.05). Similarly, the two-group analysis (AA v. AG/GG) was also indicative of a significant main effect of genotype in the right temporal pole (MNI: 39, 15, –33; F (1,155) = 14.92, K E = 948, P corrected<0.05) and right anterior cingulate cortex (MNI: 13, 41, 18; F (1,155) = 11.47, K E = 118, P corrected<0.05). Again this was because of relatively greater volume in ‘A’ homozygous individuals (AA v. AG/GG: t (155) = 3.39 and t (155) = 3.86, P corrected<0.05).
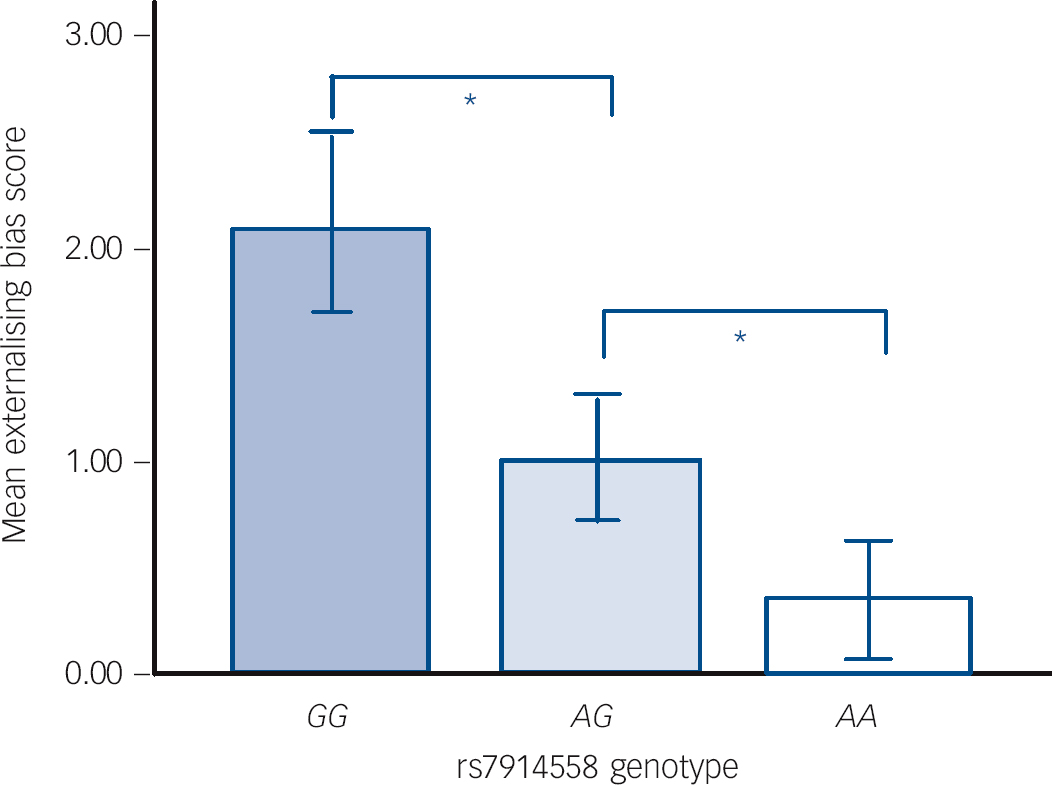
Fig. 1 Main effect of rs7914558 genotype on Internal, Personal and Situational Attributions Questionnaire (IPSAQ) externalising bias score.
Error bars show ±1 standard error. * P<0.05.
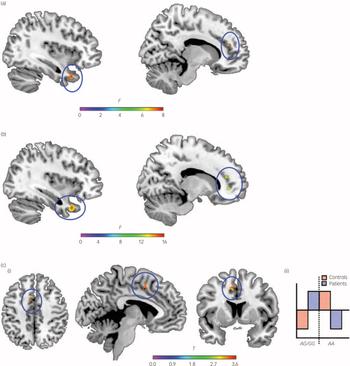
Fig. 2 The impact of rs7914558 genotype on grey matter volume in post hoc anatomical region-of-interest (ROI) analysis in regions supporting social cognitive function (anterior cingulate cortex, superior temporal gyrus and temporal pole).
(a) Irish healthy individuals (AA v. AG v. GG); (b) Irish healthy individuals (AA v. AG/GG); and (c) Italian patients and healthy individuals (AA v. AG/GG) (i) 3-plane view of grey matter volume differences, (ii) statistical parametric mapping (SPM) contrast. Note: clusters showing an impact of genotype are shown rendered on the ch2better brain template from MRIcron (www.nitrc.org/projects/mricron).
Irish sMRI/VBM sample analysis of white matter
Following correction for multiple comparisons at the whole brain level there were no white matter regions where volume varied as a function of genotype, irrespective of whether rs7914558 was considered as a fixed factor with three or two levels.
Italian sMRI/VBM sample analysis of grey matter
Both two- and three-genotype group analyses were indicative of a main effect of group (P corrected<0.05) on grey matter volume in a wide range of regions (online Table DS3). Whole brain analysis modelling genotype as a fixed factor with two or three levels failed to note effects of genotype on grey matter volume (either main effects or interactions with group). Similarly, post hoc ROI analyses that considered all three genotype groups did not reveal any impact of genotype on grey matter. However, when ‘A’ homozygotes were compared with the combined group of hetero- and homozygous carriers of the rs7491558 non-risk ‘G’ allele in ROI analyses, there was a significant interaction between group and genotype in the left anterior cingulate (MNI: –7, 7, 52; t (97) = 3.80, K E = 135, P corrected<0.05). Within this cluster, patients who were homozygous for the risk ‘A’ allele showed reduced grey matter volume (t (97) = 3.49, K E = 101, P corrected<0.05), whereas control homozygous risk carriers had greater grey matter volume (t (97) = 3.79, K E = 69, P corrected<0.05), compared with carriers of the ‘G’ allele.
Italian sMRI/VBM sample analysis of white matter
As with the grey matter analyses, there was a main effect of group on white matter volume in a number of clusters (P corrected<0.05; Table DS3), but no impact of rs7149558 at CNNM2 genotype on white matter volume in either three- or two-genotype group whole brain analyses.
Discussion
This study sought to ascertain the effects of the putative schizophrenia risk variant rs7914558 at CNNM2 on measures of neuropsychological performance, social cognition and brain volume. In Irish patients and healthy participants the risk ‘A’ allele was associated with response variation on a measure of attributional style, in the absence of genotype-related effects on neuropsychological performance. Specifically, carriers of the risk allele showed a lower externalising bias than other genotype groups. The same CNNM2 variant was also associated with variation in grey matter volume in putative ‘social cognition’ regions (i.e. anterior cingulate cortex and temporal pole), compared with non-risk individuals (AG and GG). Collectively, these data suggest that CNNM2 exerts significant but subtle effects on social cognition and its neural underpinnings.
CNNM2 rs7914558 and social cognition
Altered attributional style is commonly reported in psychiatric conditions. Instead of representing an impairment in social cognition such as mental state decoding or mental state reasoning, attributional style refers instead to an individual’s relatively stable tendency towards making inferences about the causes of social events as related to self (something I am responsible for) others (something another person intended) or situation (contextual factors). In paranoia, attributional style is characterised by the tendency to attribute negative events to external, global and stable factors. Reference Bentall, Kinderman and Kaney37 In depression, attributional style is often characterised by a tendency to attribute negative events internally. Reference Candido and Romney38 Such tendencies towards bias in social attributions and how they contribute to psychiatric disease phenotypes have been widely studied. For example, whereas the tendency to attribute external negative events to people rather than situations (i.e. personalising bias) has been associated with paranoia, Reference Jolley, Garety, Bebbington, Dunn, Freeman and Kuipers39 a predisposition towards attributing positive events to internal factors and attribute negative events to external factors (i.e. externalising bias/self-serving bias) is more typical of healthy individuals than of patients with paranoid or depressive symptoms. In the present study, we observed an association between CNNM2 and variation in externalising bias scores, suggesting that risk allele carriers were less inclined towards such self-serving attributions. These differences might be expected to reflect differences in symptomatology, for example increased depressive symptoms. Reference Seidel, Eickhoff, Kellermann, Schneider, Gur and Habel40 However, a post hoc comparison of symptom severity scores in our patients (i.e. Scale for Assessment of Positive Symptoms (SAPS) and Scale for Assessment of Negative Symptoms (SANS) scores; collected and analysed as previously reported) Reference Cummings, Donohoe, McDonald, Dinan, O'Neill and O'Callaghan41 revealed no association between clinical profiles (positive and negative symptoms) and rs7914558. This result combined with the observation that CNNM2’s effects on attributional style were consistent across patients and controls suggests that variability in attributional bias associated with CNNM2 is not necessarily indicative of clinical pathology. Rather, although CNNM2 may be associated with variation in how social attributions are formed, these effects may not be deleterious. Further studies that include measures more sensitive to depressive symptomatology (for example the Beck Depression Inventory Reference Beck42 or Hamilton Rating Scale for Depression Reference Hamilton43 ) will be helpful in elucidating this hypothesis further.
CNNM2 rs7914558 and brain structure
Anatomical ROI analyses of sMRI data were indicative of CNNM2-related variability in grey matter volume in putative social cognition regions (i.e. right anterior cingulate cortex and temporal pole) in both imaging samples. In the context of evidence of CNNM2’s influence on attributional style reported above, these findings are intriguing, although the mechanism by which these behavioural and volumetric findings relate to each other are unclear. A functional imaging investigation, which may speak to this issue, found that activity in dorsal anterior cingulate cortex was associated with self-serving bias in causal attribution. Reference Seidel, Eickhoff, Kellermann, Schneider, Gur and Habel40 This study is not designed to establish a causal relationship between brain structure and function in the anterior cingulate cortex, or between this genotype and brain function. However, dorsal anterior cingulate cortex volume has been consistently associated with cognitive reappraisal in emotion regulation, Reference Giuliani, Drabant and Gross44 and altered volume has consistently been reported in psychiatric disorders involving emotional dysregulation (including depression). Reference Giuliani, Drabant and Gross44-Reference Yucel, McKinnon, Chahal, Taylor, Macdonald and Joffe46 One possibility therefore is that the CNNM2 risk variant rs7914558 studied here may increase risk for schizophrenia and changes in attributional style via an impact on emotional processing mediated by alterations in grey matter volume in regions regulating these functions, such as the anterior cingulate cortex.
Although this notion is intriguing, it is curious that distinct genotype effects on grey matter were noted in healthy individuals v. patients. That similar effects were seen in Irish and Italian controls (for example AA>AG/GG in anterior cingulate cortex), despite the latter sample being relatively underpowered, suggests that in unaffected individuals the rs7914558 risk allele is reliably associated with increased volume. By comparison, patients who were homozygous carriers of the risk allele had relatively reduced grey matter volume in this same region. This finding was not hypothesised in our study, and may be the result of multiple factors, including CNNM2 interactions with other schizophrenia genetic risk factors and/or environmental influences that are specific to the disease phenotype. Furthermore, observed effects of CNNM2 did not indicate neurocognitive deficits, but rather variability in attributional style: whether these effects should be associated with relative increases or decreases in brain volume is unclear. To our knowledge, this is the first neurocognitive investigation of this variant to be reported, and these interpretations are inevitably speculative; further investigation of the influence of CNNM2 genotype on anterior cingulate cortex volume and function is clearly warranted to more fully ascertain its effects. In this context it is interesting to note precedents for differences between patients and controls with regard to other schizophrenia risk variants (for example ZNF804A, DARRP32), which have since been replicated.
Bioinformatic analysis of rs7914558
The SNP under consideration here, rs7914558, is located on chromosome 10q24 in a large region of high and complex linkage disequilibrium. Analysis of HapMap Phase II + III data (release 28 NCBI build 36) indicates that rs7914558, positioned at 104 765 898(hg18), is in high linkage disequilibrium (r 2>0.8) with 42 other SNPs in this region (including 22 SNPs at r 2 = 1). These SNPs span a 322 kb region from rs11191438 at position 104 627 854 to rs4307650 at position 104 949 842. This region contains four RefSeq genes: C10orf32, AS3MT, CNNM2 and NT5C2. Analysis of SNP×gene expression databases including brain (see online supplement) does not identify a direct functional link between rs7914558 (or proxy) and altered expression of any of the four genes in this region; due to long-range linkage disequilibrium in this region all four remain candidate schizophrenia risk genes. It is worth noting that new data identify a putative functional interaction between CNNM2 and ZNF804A, an established schizophrenia risk gene. Hill and colleagues Reference Hill, Jeffries, Dobson, Price and Bray47 recently reported that a reduction in CNNM2 expression was observed following small interfering RNA (siRNA) knockdown of the ZNF804A protein in human neural progenitor cells. This is interesting because ZNF804A is the only other gene - schizophrenia-implicated or otherwise - to have been associated with variation in social attributional styles, the phenotype implicated in our study. Reference Hargreaves, Morris, Rose, Fahey, Moore and Cummings19
Study limitations
Interpretation of these data is limited by a number of factors. First, little is known regarding the functional pathways of the rs7914558 SNP considered here, either in general or with regard to schizophrenia specifically. Certainly, our understanding of how or why this particular variant contributes to social cognitive processes involved in attributional style and variability in brain volume, and how this speaks to schizophrenia risk, will be significantly enhanced by the elucidation of the functional biology of this particular variant. Second, we are limited by the absence of patients with schizophrenia in the Irish imaging sample, and as such are unable to more fully determine patterns of similarity and/or difference between the Irish and Italian samples. Finally, the relatively small number of patients available for sMRI means that we are unable to accurately model other factors that might contribute to patterns of altered brain volume because of variability in the rs7914558 genotype. Although many neuroimaging genetics studies do not include patients, and notwithstanding that our total neuroimaging samples are well in excess of the average samples size of these studies, Reference Rose and Donohoe48 replication in further samples (including patient samples) will be important to confirm these results.
Implications
This study, which reports for the first time on the neurocognitive effects of CNNM2, provides evidence that rs7914558 has an effect on both social cognition (i.e. attributional style) and grey matter volume in regions previously implicated in the processing of social stimuli. These effects occur in the absence of an effect on neuropsychological performance as measured by IQ, memory and attentional control. Given the uncertainty regarding the functional significance of this variant and the biological mechanism by which it increases schizophrenia risk, these data suggest that this schizophrenia-associated variant may effect neural systems relevant to social cognition. Whether the effects observed here at the level of behaviour and cortical volumes related to social cognition represent pleiotropic effects, or may mediate the relationship between genotype and increased disease risk remains to be established.
Acknowledgements
We thank our collaborators in the TCIN Biobank Project (Drs Ian H. Robertson, Hugh Garavan, John O’Doherty, Fiona N. Newell, Jane McGrath, Arun Bokde and their staff and students) and those who volunteered to participate in these studies. We are also grateful to Professors John Waddington, Ted Dinan, Eadbhard O’Callaghan and Kieran Murphy, and Dr F. Anthony O’Neil for their role in the recruitment of the Irish neuropsychological patient sample. The authors also acknowledge data management support from the Trinity Centre for High Performance Computing. The data management system used for this work was BC|SNPmax v. 3.5-121 (Biocomputing Platforms Ltd, Finland).





eLetters
No eLetters have been published for this article.