Acinetobacter baumannii today plays a significant role worldwide as a cause of outbreaks of infections among hospitalized patients, especially those in critical-care environments. The carbapenems, imipenem and meropenem, have been widely used to treat infections caused by multidrug-resistant A. baumannii clinical isolates but carbapenem resistance among these strains has increased in recent years [Reference McGowan1]. Resistance is mediated through various combined mechanisms that include target inaccessibility or drug inactivation by β-lactamases with carbapenemase properties such as class B metallo-β-lactamases (MBLs) and several class D OXA-type enzymes [Reference Livermore2]. The metallo-enzymes are rarely detected, while the OXA-type carbapenemases are widespread among A. baumannii from several geographic regions [Reference Brown and Amyes3–Reference Tsakris5]. The latter enzymes initially formed two subgroups, namely OXA-23 and OXA-24 [Reference Brown and Amyes3], but recently two novel subgroups of class D oxacillinases with carbapenemase properties, formed by OXA-51 variants and OXA-58, have been additionally described in A. baumannii [Reference Brown, Young and Amyes6, Reference Poirel7].
The OXA-51 subgroup of enzymes exhibits relatively weak hydrolytic activities, compared with other carbapenemases [Reference Héritier8], but has been associated with carbapenem resistance in isolates with the insertion sequence ISAba1 upstream of the oxacillinase gene [Reference Turton9]. bla OXA-51-like alleles are chromosomally located and it has been confirmed that these sequences are intrinsic in most or all A. baumannii strains [Reference Coelho4, Reference Merkier and Centrón10, Reference Turton11]. To date in the same cluster of enzymes, a large number of closely related variants have been described that include OXA-64 to -71, OXA-75 to -80, OXA-82 to -84, OXA-86 to -89, OXA-91, OXA-92, OXA-94 and OXA-95 (www.lahey.org/studies/other.asp#table1 [Reference Turton9–Reference Vahaboglu14]). A. baumannii is a predominant pathogen in Greece, particularly among immunocompromised patients and the OXA-51 subgroup has been commonly detected among selected clinical isolates with various susceptibility profiles to carbapenems [Reference Tsakris5, Reference Pournaras15]. The purpose of the present study was to investigate the geographical distribution and the diversity of the bla OXA-51-like alleles among A. baumannii clones in Greece.
In total, 150 isolates of A. baumannii were included randomly, all of which were collected between 2000 and 2005 from clinical samples of patients hospitalized in five hospitals in three major Greek regions (Athens, Thessaloniki and Larissa). Several of these isolates were retrieved from collections stored during previous studies [Reference Tsakris5, Reference Pournaras15, Reference Tsakris16]. The isolates were identified to species level by the Vitek 2 automated system (bioMérieux, Marcy l'Etoile, France), the API 20NE system (bioMérieux) and conventional biochemical tests.
The minimum inhibitory concentrations (MICs) of imipenem and meropenem were determined using an agar dilution method [17] with agar plates containing serially diluted antibiotics ranging from 0·25 to 512 mg/l respectively for each agent; the inoculum was 104 c.f.u. per spot. The isolates were also tested by E-test MBL (AB Biodisk, Solna, Sweden) as well as the imipenem-EDTA double-disc synergy test (DDST) for MBL production [Reference Oh18]. Pseudomonas aeruginosa ATCC 27853 was used as control in susceptibility testing and a VIM-type carbapenemase-producing A. baumannii strain [Reference Tsakris5] was used as control for E-test MBL.
PFGE of ApaI-digested genomic DNA of A. baumannii isolates was performed with a CHEF-DRIII system (Bio-Rad, Hemel Hempstead, UK) according to previously described methods [Reference Kaufmann, Pitt and Chart19]. The interpreting criteria were described by Tenover et al. [Reference Tenover20]. ApaI macrorestriction patterns were digitized and analysed using the Quantity One Software (Bio-Rad Laboratories Inc., Hercules, CA, USA) to calculate Dice coefficients of correlation and to generate a dendrogram by the unweighted pair-group method using arithmetic averages (UPGMA) clustering.
The isolates were screened by PCR for the bla OXA-51-like genes using primers OXA-69A and OXA-69B that amplify a 975-bp product including the whole coding sequence of the gene [Reference Héritier8], in order to discriminate the alleles by sequence analysis. Isolates that did not give a PCR product with these external primers were screened with the partially degenerate primers published previously [Reference Pournaras15] that were designed to amplify the 825-bp ORF of all bla OXA-51-like alleles. Amplicons were purified using ExoSAP-IT reagent (USB Corporation, Cleveland, OH, USA) and both strands were sequenced using the standard dideoxynucleotide method in an ABI Prism 377 DNA sequencer (PerkinElmer, Applied Biosystems Division, Foster City, CA, USA).
The UPGMA dendrogram of ApaI-digested genomic DNA grouped as many as 112 of the isolates into two major clusters; four additional clusters were detected each containing from two to 19 isolates, while the remaining seven isolates exhibited unique PFGE types (Fig.). The characteristics of the A. baumannii isolates of the study are presented in the Table. The imipenem and meropenem MICs ranged from 0·5 to 512 mg/l and from 0·25 to 256 mg/l, respectively. Both E-test MBL and DDST were negative in all isolates. PCR screening for bla OXA-51-like gene with primers OXA-69A and OXA-69B was positive in all but seven of the 150 acinetobacters. The seven isolates that did not give a PCR product with these external primers, gave a positive signal with the degenerate primers. Sequencing analysis of the 150 bla OXA-51-like amplicons revealed alleles that were identical to bla OXA-66 in all isolates from the regions of Thessaloniki (n=47) and Larissa (n=63), while bla OXA-66 (n=20) along with bla OXA-69 (n=18) and bla OXA-65 (n=2) alleles were detected in isolates from the Athens region. In total, bla OXA-66 was detected in 130 isolates that grouped into ten genotypes, bla OXA-69 in 18 isolates that grouped into three genotypes and bla OXA-65 in two isolates, each one belonging to an individual genotype.
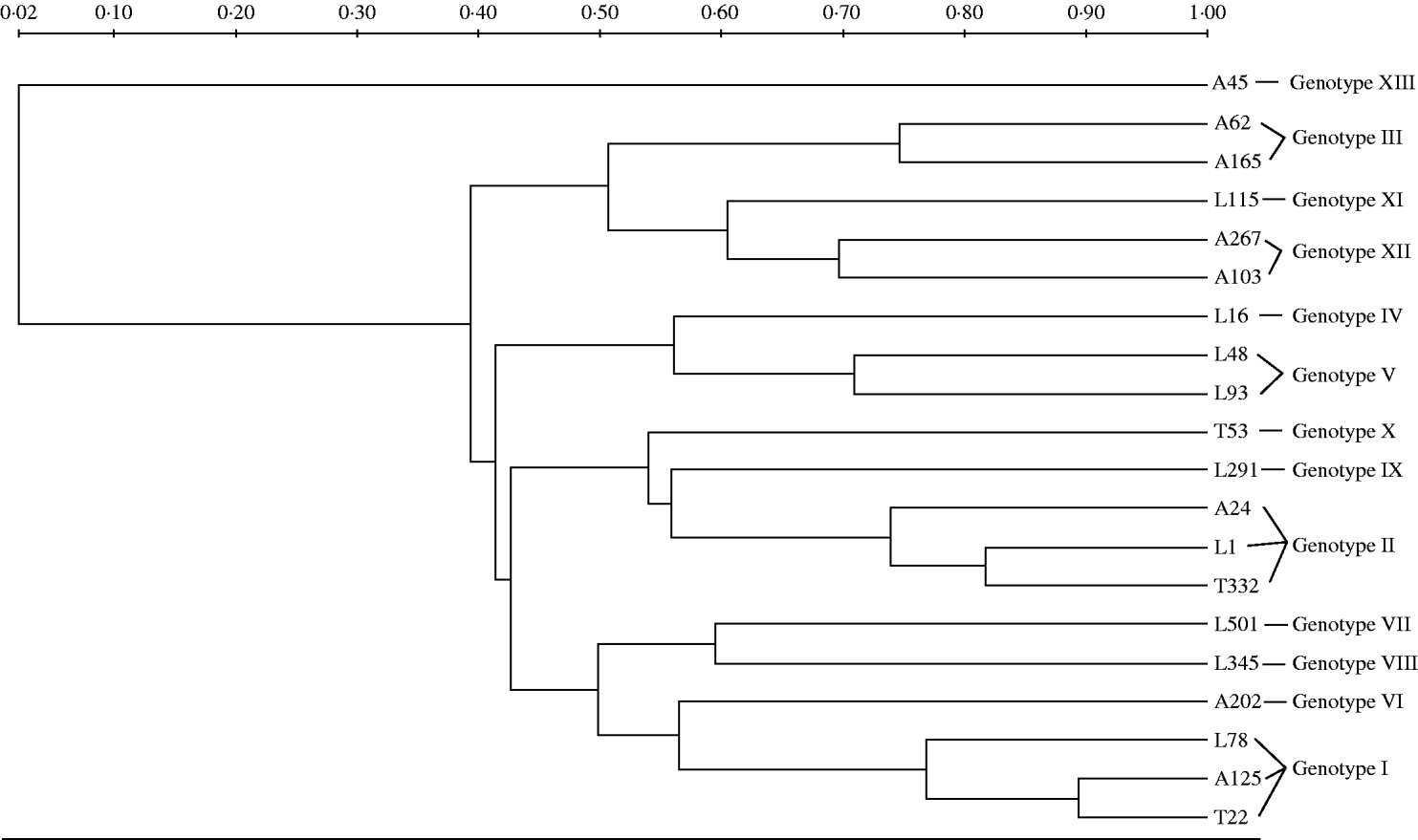
Fig. UPGMA dendrogram of ApaI-digested genomic DNA of representative isolates corresponding to all 13 genotypes included in the study. Clusters of possibly related isolates were defined at 70% similarity. Tags of each branch stand for the origin (A, Athens; L, Larissa; T, Thessaloniki) and the numbering of each isolate.
Table. Genotypes, region of isolation, range of imipenem and meropenem minimum inhibitory concentrations (MICs) and blaOXA-51-like alleles among the 150 A. baumannii isolates

* Region of isolation: A, Athens (n=40); T, Thessaloniki (n=47); L, Larissa (n=63). The + sign refers to the region of isolation of each genotype.
Carbapenem-hydrolysing β-lactamases that belong to class D have been widely described over the last years [Reference Walther-Rasmussen and Høiby13]. The subgroup of class D carbapenemases formed by OXA-51 variants was first detected in distinct clones of A. baumannii from Argentina [Reference Brown, Young and Amyes6]; it shares very weak identities with other known oxacillinases and thus comprises a novel phylum among the OXA-type carbapenemases [Reference Brown and Amyes3]. It is of interest that the OXA-51 cluster, in contrast with other class D carbapenemases, does not possess the tyrosine-to-phenylalanine substitution in the conserved Y-G-N motif of the class D protein [Reference Brown and Amyes12]. So far, more than 20 class D β-lactamases, all of which belonged to the same OXA-51 subgroup, have been identified among A. baumannii clinical isolates from centres in Argentina, South Africa, Hong Kong, Spain, Singapore and Turkey [Reference Turton9–Reference Vahaboglu14]. The number of alleles in this subgroup seems to have extended constantly through silent or amino-acid shift mutations. The members of the cluster diverge by 1–15 amino-acid modifications and these intrinsic enzymes seem to represent the largest and most diverse collection of class D carbapenemases in A. baumannii [Reference Héritier8, Reference Walther-Rasmussen and Høiby13, Reference Woodford21].
In the present study three bla OXA-51 variants were detected in different parts of the country. Interestingly, the bla OXA-66 allele was exclusive for acinetobacters that exhibited various levels of susceptibility to carbapenems and were recovered in two regions. It was also the prevalent bla OXA-51-like allele among A. baumannii in the third region of the study. The bla OXA-66 sequence was integrated in almost all isolates of the two major PFGE-defined clones, while the same sequence was additionally detected in eight of the remaining 12 clonal types; less frequently detected were other bla OXA-51-like alleles such bla OXA-69 and bla OXA-65. It is of interest that the three bla OXA-51-like alleles detected here are the most closely related at the genetic level within the OXA-51 subgroup [Reference Brown and Amyes12, Reference Walther-Rasmussen and Høiby13]. Initially, the bla OXA-66 allele was identified in individual isolates from Spain, Hong Kong and Singapore, while the bla OXA-69 allele was found in two isolates from Singapore and one from Turkey and the bla OXA-65 allele was found in clonal isolates from Argentina [Reference Brown and Amyes12]. In a previous study conducted in Turkey, a higher diversity of bla OXA-51-like alleles occurred among A. baumannii isolated in distinct hospitals [Reference Vahaboglu14]; the original bla OXA-51 sequence was the most frequently detected while bla OXA-66 was present only among imipenem-susceptible A. baumannii clones. In a French collection of A. baumannii strains, the latter sequence was identified along with six additional variants among imipenem-susceptible and -resistant isolates [Reference Héritier8], and recently in Argentina seven bla OXA-51-like alleles were found among representative A. baumannii isolates [Reference Merkier and Centrón10]. Our study brings more evidence to support the hypothesis that the bla OXA-51-like gene is intrinsic to A. baumannii. It also highlights the high prevalence and spread of a specific allele, bla OXA-66, which possibly indicates that our A. baumannii clones might have originated from a common ancestor and have undergone, as yet, limited dissemination. It might be expected that after a lengthy time period a common ancestor isolate could have evolved to different macrorestriction patterns following the accumulation of spontaneous mutations. However, the possibility that these few bla OXA-51-like variants, with bla OXA-66 being predominant, are widely disseminated in several unrelated A. baumannii strains in Greece cannot be excluded and warrants further investigation.
DECLARATION OF INTEREST
None.






