Introduction
Although significant progress has been made in reducing the incidence and impact of Escherichia coli O157:H7, it remains the largest cause of post-diarrhoeal hemolytic uremic syndrome (HUS) [Reference Banatvala1]. HUS incidence varies by age, with the greatest burden among children <5-years-old [2–Reference Crim5].
Beyond age, pathogen characteristics are an important factor in determining progression to HUS. Shiga toxin (Stx), E. coli O157:H7's cardinal virulence factor, can be encoded by multiple genes (stx1 and stx2), with some genotypes more frequently associated with HUS than others [Reference Friedrich6–Reference Luna-Gierke9]. A study in 2008 identified a subtype of E. coli O157:H7, termed clade 8, associated with increased risk of HUS [Reference Manning10]. Although numerous studies have investigated virulence factor expression that may be responsible for this association [Reference Abu-Ali11–Reference Amigo14], studies confirming the association have been limited, suggesting an effect of varying strength and specificity [Reference Soderlund15–Reference Haugum17].
The phylogenetic definition of the E. coli O157:H7 serotype has advanced since the 2008 discovery of the putatively hypervirulent clade 8 and it is unknown whether branches of the updated phylogenetic tree are also associated with HUS. In contrast to the earlier tree, Bono et al. [Reference Bono18] reported a tree of phylogenetic lineages that drew on a large pool of systematically chosen single nucleotide polymorphisms (SNPs) and incorporated isolates from a diverse set of sources. This tree was further developed by Jung et al. [Reference Jung19].
In a population-based cohort of 1160 E. coli O157:H7 cases in Washington State, reported 2005–2014, we sought to use the updated lineages to confirm and refine the role of phylogenetics in HUS risk proposed by Manning et al. [Reference Manning10]. Given the higher incidence of HUS among young children and the preponderance of clade 8 strains isolated from children by Manning et al. [Reference Manning10], we also investigated the role of age in the association between phylogenetic lineage and HUS, evaluating it as a potential confounder and effect modifier.
Methods
Study setting and design
We conducted a population-based retrospective cohort study of all culture-confirmed E. coli O157:H7 cases reported to the Washington State Department of Health (DOH) from 2005 through 2014. Mandatory Shiga toxin-producing E. coli case reporting occurs primarily through diagnostic laboratories and healthcare providers. Local health jurisdiction personnel use a standardised DOH case report form to obtain demographic information, potential exposures and details of the course of illness.
We confirmed HUS status during a review of all reported, hospitalised, culture-confirmed E. coli O157:H7 cases from the study period. HUS was defined as a hematocrit <30%, platelet count <150 000/mm3 and serum creatinine concentration above the normal for age. All criteria needed to be met on the same day. Non-hospitalised cases were considered to not have HUS because all patients with HUS would be hospitalised due to the severity of disease.
The Washington State Institutional Review Board designated this study as exempt.
Isolate typing
All E. coli O157:H7 isolates were sent to DOH for microbiological confirmation and pulsed-field gel electrophoresis (PFGE) analysis. We obtained these isolates from DOH and determined their lineage according to the phylogenetic tree developed by Bono et al. [Reference Bono18] and expanded by Jung et al. [Reference Jung19]. The 48-plex SNP assay developed by Jung et al. [Reference Jung19] was used to type 793 of the 1160 isolates. Isolates that did not undergo SNP-typing were then assigned the lineage of a SNP-typed isolate with the same PFGE profile (Supplementary Material), for a total of 1121 isolates with an assigned lineage. Of the 39 excluded isolates, six were biochemically atypical E. coli O157:H7 and 33 were not available for typing (Supplementary Material).
Based on Jung et al.’s categorisation as clinical, bovine-biased, or sparsely represented lineages [Reference Jung19], we retained the clinical lineages Ib, IIa and IIb as separate categories. The bovine-biased and remaining lineages were grouped into a clinically rare category, reflecting the low frequency of isolating these groups from human cases.
A subset of 480 isolates was also typed using a 32-plex SNP assay to determine Manning clade [Reference Manning10]. We used PFGE type to infer Manning clade for an additional 422 untyped isolates (Supplementary Material). Distribution of Manning clades by Bono-Jung lineages is shown in Supplementary Table S1.
A subset of 453 isolates underwent Stx-encoding bacteriophage insertion (SBI) typing. The SBI typing methods, which use PCR to detect 12 targets including stx1, stx2a and stx2c, have been described [Reference Jung19, Reference Shringi20].
Statistical analysis
Case data were merged with isolate typing results using a unique identifier and the dataset was de-identified before analysis. Age [Reference Rogers21–Reference Reiss24] and sex [Reference Rowe25–Reference Al-Jader27] were considered a priori confounders. Distributions of other potential confounders were summarised in contingency tables. Aside from age, no examined variable was significantly associated with both lineage and HUS.
Logistic regression with generalised estimating equations (GEE) was used to estimate the association between lineage and HUS. Lineage was modeled as a categorical, group-level variable, with the most common lineage (Ib) as the reference category and groups defined by PFGE types. An exchangeable working correlation matrix was used for all analyses. Robust standard errors calculated using the sandwich estimator accounted for any potential misspecification of the correlation structure. HUS is sufficiently rare that the odds ratio (OR) calculated from model coefficients could be interpreted as the risk of HUS associated with a lineage (e.g. IIa) relative to lineage Ib. A 95% confidence interval (CI) was estimated for each OR. Age, modelled as a continuous variable, and sex were added as covariates in the adjusted model. To examine effect modification, the sex-adjusted GEE model was stratified by age group and the lineage OR estimates were compared across strata. Unadjusted, adjusted and stratified analyses were also conducted for the association of Manning clade and HUS (Supplementary Material).
To understand the impact of our use of multiple isolates per PFGE type, we simulated the method used in previous studies [Reference Manning10, Reference Iyoda16] for comparison with our results. One isolate per PFGE type was randomly drawn and used in a single-level adjusted logistic regression. For each of 10 000 repetitions of this process, the coefficient estimates for lineages IIa and IIb and the associated P-values were recorded. We examined the distribution of coefficient estimates by lineage and calculated the proportion that was statistically significant at P < 0.05.
The frequency of Shiga toxin subtypes was summarised by lineage and HUS status. Formal mediation analysis of the role of stx genotypes in the association between lineage and HUS was planned. However, most major stx genotypes were too highly correlated with lineage to differentiate the direct effect of lineage and that mediated by stx genotype.
R [28] was used for all analyses.
Results
There were 1160 culture-confirmed E. coli O157:H7 cases reported to DOH during the 10-year-period. Validated HUS status was available for 1082 cases of the 1121 cases with an assigned lineage; the HUS definition was met for 76 (7.0%). HUS status differed by age, with children <5-years-old constituting over half of HUS cases but less than one-fourth of non-HUS cases (Table 1). The case fatality was 3.9% among HUS cases and 0.4% among non-HUS cases.
Table 1. Frequency of case characteristics of patients reported in Washington State with confirmed E. coli O157:H7 infection, 2005–2014
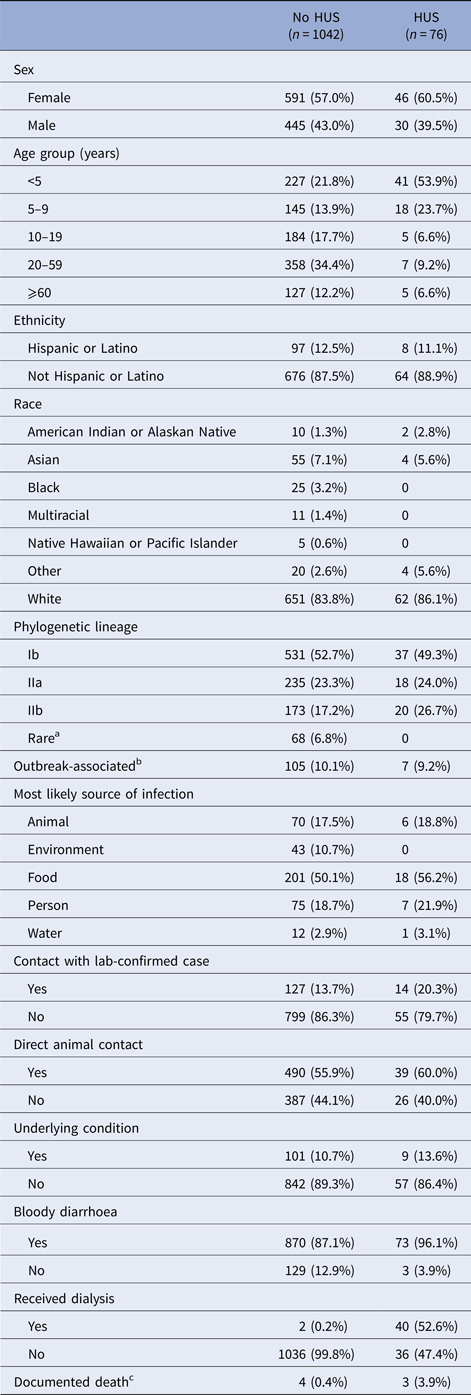
HUS, hemolytic uremic syndrome.
a ‘Rare’ lineages include 12 different lineages.
b Whether a case was associated with an outbreak was not reported for most cases, so only positive responses are shown.
c Death status was not reported in most cases. There were eight reported deaths. Only seven are shown in the table. The eighth was hospitalised, but the chart could not be abstracted to determine HUS status.
Phylogenetic association with HUS
In the unadjusted GEE model, lineage IIb was associated with increased risk of HUS relative to lineage Ib (OR = 1.65, 95% CI 1.05–2.60) (Table 2). There was no elevation in HUS risk among lineage IIa cases, compared with lineage Ib cases. No HUS cases occurred in the group of rare lineages; effect estimates for this group are not presented because of statistical instability. After adjustment for age and sex, the association between IIb and HUS was attenuated and no longer distinguishable from the null (OR = 1.43, 95% CI 0.90–2.25).
Table 2. Association of E. coli O157:H7 phylogenetic lineage and HUS
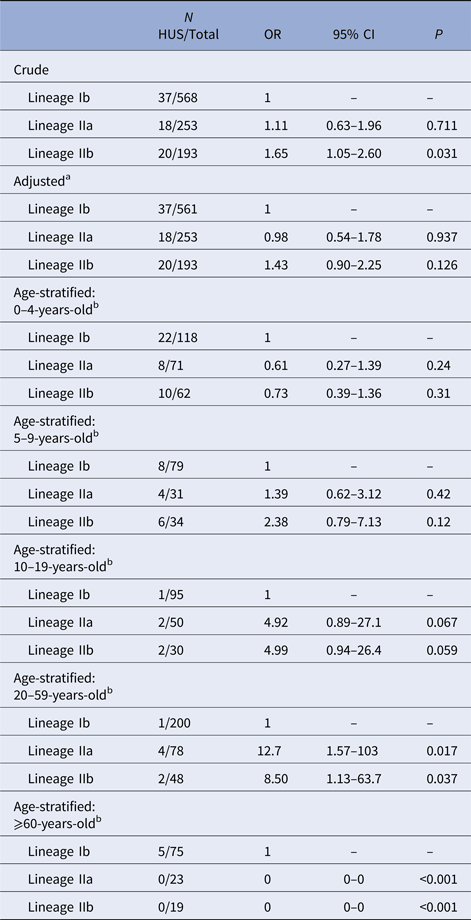
Logistic regression, using GEE, of HUS status on phylogenetic lineage. No HUS occurred in the group of cases infected with rare lineages, so results are not shown for this group.
a Model adjusted for age as a continuous variable and sex.
b Model adjusted for sex. CI, confidence interval; GEE, generalised estimating equations; HUS, hemolytic uremic syndrome; OR, odds ratio.
The proportion of E. coli O157:H7 infections caused by lineage IIb strains decreased with age and those caused by Ib strains increased. An effect of lineage on HUS risk could not be established in 0–4 or 5–9 year-olds. In 10–19 and 20–59-year-olds, lineages IIa and IIb were associated with an increased risk of HUS, relative to lineage Ib, with effect estimates highest in the latter group (IIa OR = 12.7, 95% CI 1.57–103; IIb OR = 8.50, 95% CI 1.13–63.7). There were no lineage IIa or IIb HUS cases among ⩾60-year-olds (Table 2).
Results of unadjusted, adjusted and stratified analyses assessing the association between Manning clade and HUS were consistent with those seen for lineages (Supplementary Table S2).
In simulations of selecting and analysing only one isolate per PFGE type, the distribution of effect sizes for the association between lineage IIa and HUS was centered near 0 (OR = 1) and 0.1% of estimates had P < 0.05 (Supplementary Fig. S1). This is consistent with our adjusted effect estimate for lineage IIa (Table 2). The distribution of effect sizes for lineage IIb was centred near 2 (OR > 7) with P < 0.05 for 25% of simulations (Supplementary Fig. S1). This is not consistent with our results for lineage IIb using all isolates (OR = 1.43, P = 0.13) (Table 2).
Shiga toxin genotype
Shiga toxin genotype was determined for 469 cases, 453 of which also had a validated HUS status. Distribution of stx genotypes by lineage showed that 92% of isolates contained stx2a, whether alone or in combination with another stx gene (Table 3). Lineage Ib isolates were dominated by the stx1-stx2a genotype (90%). Lineage IIa isolates were predominantly the stx2a-stx2c genotype (84%). Most lineage IIb isolates (94%) had only the stx2a gene. Six isolates had none of the three probed stx genes at the time of typing. Relative to the frequency of HUS among cases infected with stx1-stx2a and stx2a-stx2c strains (11% and 12%, respectively), cases infected with stx2a-only strains had a higher frequency of HUS (21%).
Table 3. Distribution of Shiga toxin genotypes by phylogenetic lineages
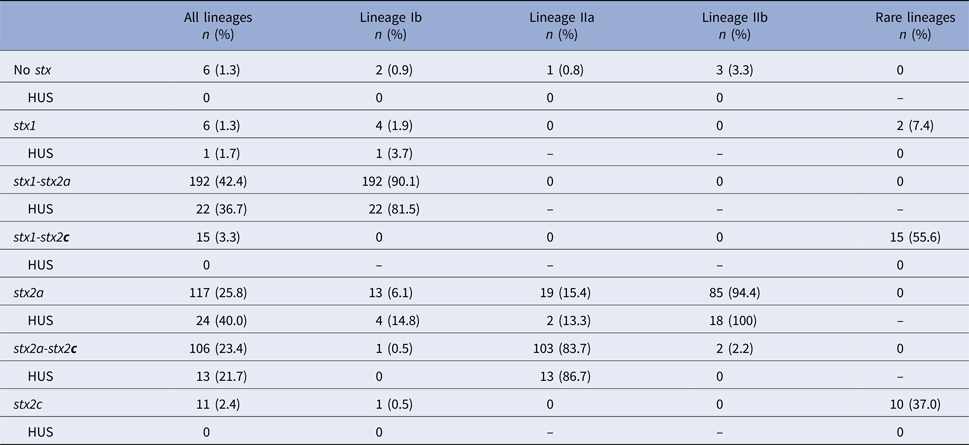
HUS, hemolytic uremic syndrome; stx, Shiga toxin gene.
No isolates were observed with the stx1-stx2a-stx2c genotype.
Discussion
The results of this study do not support an association between phylogenetic lineage (or clade) and HUS for children <10, the age group with the greatest burden of E. coli O157:H7 and HUS. While lineage IIb was associated with increased risk of HUS in unadjusted analysis, stratifying by age indicated an increased risk of HUS associated with lineages IIa and IIb, relative to lineage Ib, only among 10–59-year-olds. In the eldest group, lineage Ib conferred greater HUS risk than either lineage IIa or IIb. Our analysis of the risk associated with Manning clade 8 was consistent with our lineage IIa/IIb results.
Age has long been considered the strongest predictor of progression to HUS among those with E. coli O157:H7 infection and our results are consistent with that: 15.9% of children <10-years-old progressed to HUS, compared to 2.5% of individuals ⩾10. Similarly, the incidence of HUS in our study was 6.79 per 100 000 <10-year-olds, compared with 0.29 per 100 000 ⩾10-year-olds. The lack of association we found between lineage and HUS among those aged <10 years suggests that differential infection by high virulence lineages does not explain why young children are more likely to progress to HUS. However, our findings show that lineage IIb strains disproportionately establish disease in young children, driving the observed unadjusted association between lineage IIb and HUS and suggesting that there is a difference in either exposure or early disease manifestation that leads to more IIb-infected cases being reported in this age group than cases infected with other lineages.
Among those aged 10 and over, we observed substantially more reported cases infected by lineage Ib than lineages IIa and IIb, which may indicate less exposure to lineage IIa and IIb strains or greater difficulty for these strains in establishing disease. However, if IIa or IIb strains are successful in establishing disease, they appear more likely to cause HUS than the more common lineage Ib strains. The eldest group, ⩾60-year-olds, appears to be an exception, with higher risk associated with lineage Ib strains. Individuals ⩾60-years-old have a slightly higher incidence of HUS (0.58 per 100 000) than 10–59-year-olds (0.26 per 100 000), and E. coli O157:H7 outbreaks have occurred in nursing homes [Reference Reiss24], making this age group of particular interest. The reversal of the association in the eldest group is curious and with a low number of cases among older children and adults, urges caution in interpreting our results in 10–59 and ⩾60-year-olds.
In a 2008 study of 333 Michigan cases with unique PFGE fingerprints, Manning et al. [Reference Manning10] identified a sevenfold increased odds of HUS among patients infected with E. coli O157:H7 clade 8 strains after adjustment for age (0–18 vs. 19–64), sex and symptoms. Subsequent studies have also suggested an association of varying magnitudes between clade 8 and HUS [Reference Soderlund15, Reference Iyoda16]. Lineages IIa and IIb in the present study overlap with clade 8 and show an elevation in risk of HUS only among 10–59-year-olds. There are multiple reasons why our results may have differed from those of others. First, some previous studies have either not adjusted for age [Reference Soderlund15] or adjusted by large age groups [Reference Manning10], increasing the potential for residual confounding. Only one previous study stratified by age [Reference Iyoda16]. In our analysis, sensitive age groups defined based on the epidemiology of the disease were critical in better understanding the association.
Second, both Manning et al. [Reference Manning10] and Iyoda et al. [Reference Iyoda16] used one representative isolate from each outbreak or PFGE-defined strain. We demonstrated through simulation that studies using only one isolate per strain had an average effect estimate higher than that obtained using all isolates for lineage IIb and that for lineage IIb, 25% of analyses would appear statistically significant merely by chance. This finding emphasises the importance of incorporating the complete data.
Third, previous studies relying on logistic regression [Reference Manning10, Reference Iyoda16] appear to have modelled each clade as an independent variable, interpreting estimates as the odds of HUS in one clade vs. all other clades. This method introduces perfect multicollinearity, which can induce large, unpredictable biases in point estimates and standard errors [Reference Tu, Clerehugh and Gilthorpe29, Reference Vatcheva30]. Perfect multicollinearity also gives the OR dubious interpretability, because, by definition, you cannot hold the other clades constant (at 0 or 1) and change the clade of interest from 0 to 1. To avoid this pitfall, we modelled lineage (and clade) as a categorical value in which the most common lineage Ib (clade 2/3) was used as the reference category.
Finally, the only other study to consider effect modification by age, Iyoda et al. [Reference Iyoda16] reported an OR for clade 8 of 6.1 for 0–9-year-olds and 3.1 for children and adults ⩾10 years. Their results are in contrast to those we report here, potentially because of their use of asymptomatic controls. We estimated the odds of HUS for ill E. coli O157:H7 cases, thus estimating virulence, the probability of progressing from non-severe to severe disease. Comparing HUS cases with asymptomatic carriers mixes virulence with pathogenicity, the probability of becoming ill if infected.
We observed a very close correlation of lineage and stx genotype, which is similar to previous studies [Reference Jung19, Reference Mellor31]. This may be suggestive of a major role of stx genotype in the association between lineage and HUS. Other cohorts, including one of <10-year-olds, have shown stx2a-only and, to a lesser degree, stx2a-stx2c genotypes associated with progression to HUS [Reference Friedrich6, Reference Persson7, Reference Jelacic32]. These are also the most common genotypes among clade 8 isolates [Reference Manning10, Reference Iyoda16, Reference Haugum17] and studies of clade 8 isolates have described the potential for high Stx2 production [Reference Neupane12, Reference Ogura33, Reference Amigo34]. Our analysis, which shows that most lineage IIa strains carry stx2a-stx2c and most IIb strains carry only stx2a, is consistent with these studies.
Our study was limited to reported cases, which are likely more severe than unreported cases. Our results can therefore not be extended to unreported cases. However, it is unlikely that any HUS cases went unreported due to the severity of the condition. Our study also included cases from only Washington State, potentially limiting its generalisability to areas with differing E. coli O157:H7 populations. Indeed, previous work has suggested local E. coli O157:H7 circulation in Washington [Reference Tarr35], emphasising the importance of small geographic areas in the bacteria's population dynamics. The strains composing each lineage may differ in other geographic regions and those within-lineage differences could alter the association observed with HUS. However, it is reassuring that a large number of isolates in our study are from the most commonly isolated PFGE types in the USA.
We were also not able to assign phylogenetic lineage to 39 isolates, one of which was identified as a HUS case. These isolates tended to be from earlier in the study period, indicating that they are not missing completely at random. The composition of the bacterial population shifted slightly during the study period [Reference Tarr35], with lineage Ib more dominant early in the period. However, the small number of untyped isolates relative to the whole sample likely did not alter our results.
Over 75% of our HUS cases were in children <10-years-old, giving us limited precision to estimate the effect of lineage on HUS in older children and adults. This is reflected in the large confidence intervals around estimates for the 10–19 and 20–59 age groups. A larger sample of cases ⩾10-years-old would provide a better estimate of the true effect of lineage on HUS in this age group. However, we are confident in our estimates for the effect in young children, the age group with the highest incidence of both E. coli O157:H7 and HUS.
The lack of sufficient variability of most stx genotypes in a single lineage precluded formal mediation analysis. It is possible that with a much larger sample one could differentiate the direct effect of lineage on progression to HUS from the effect mediated by stx genotype. Ideally, mediation should be examined stratified by age group, to reflect the apparent effect modification of the overall association.
This study benefited from over 1100 E. coli O157:H7 cases, including 76 HUS cases. HUS outcomes were validated with hospital records using a standardised definition to ensure the comparability of our outcome. By employing correlated data methods, we were able to incorporate data from the entire cohort instead of limiting the study to representative isolates from each PFGE type, which our simulation study showed is an important step in accurately estimating the association. By using a consistent reference group (lineage Ib), we were also able to avoid the perfect multicollinearity of previous studies, reducing bias and allowing meaningful interpretation of our effect estimates. Applying these methods to the Bono-Jung lineages and Manning clades produced consistent results.
This study demonstrates that E. coli O157:H7 phylogenetic lineage likely only contributes to HUS risk among older children and adults. Further studies are needed to confirm this association, given the rarity of the disease among adults. In young children, the proportion of infections caused by lineage IIb strains was higher than in older groups. It will be important to determine whether this is driven by differences between age groups in exposure, transmission and/or early disease development. Additionally, given the close correspondence of lineage and stx genotype, learning how exposure and early illness differ across lineages may translate to prevention opportunities for the strains that tend to carry more virulent stx genotypes.
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1017/S0950268818001632
Disclaimer
The content is solely the responsibility of the authors and does not necessarily represent the official views of the funding agencies.
Acknowledgements
This work was supported by the National Institute of Environmental Health Sciences of the National Institutes of Health (G.A.M.T., grant number T32ES015459); the National Institute of Allergy and Infectious Disease of the National Institutes of Health (G.A.M.T., grant number F31AI126834); and the US Department of Agriculture National Institute of Food and Agriculture (T.E.B., grant numbers 2009-04248, 2010-04487). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health or US Department of Agriculture.
Conflict of interest
None.





