Implications
Fertilization is an intricate cascade of events that irreversibly alter the participating male and female gamete and ultimately lead to the union of paternal and maternal genomes in the zygote. Proper functioning, recycling and remodeling of gamete structures at fertilization is aided by the ubiquitin–proteasome system (UPS), the universal substrate-specific protein recycling pathway present in bovine and other mammalian oocytes and spermatozoa. Research focused on the UPS-regulated aspects of the fertilization process has a potential to impact the management of bull fertility, the optimization of artificial insemination, genomic selection for production and fertility-related traits, and the improvement of sperm viability after sexing and cryopreservation in cattle.
Introduction: prelude to sperm–oocyte interactions
However, for the semen to exert its effect on the eggs, it must enter their little bodies, as it seems unlikely that it could animate them by the mere contact with their skin (Spallanzani, Reference Zimmerman, Manandhar, Yi, Gupta, Sutovsky, Odhiambo, Powell, Miller and Sutovsky 1780 ).
Quoted above, Italian pioneer of reproductive research and father of fertilization biology Lazzaro Spallanzani (1729–99), known for conducting the first recorded experiments with artificial insemination in frogs and canines, captured the essence of the fertilization process long before his successors were able to observe the actual union of a spermatozoon and an oocyte and decipher the molecular machineries that guide it. To paraphrase the words of another luminary of fertilization research, C.R. Moore, fertilization is a precisely orchestrated cascade of irreversible events that starts long before the male and female gametes meet. This intricate wedding dance starts with the mature gametes reaching the site of fertilization within the female oviduct. Spermatozoa have to bypass multiple hurdles crossing from the site of deposition though the female cervix (in vaginal depositors), uterus and uterotubal junction, to finally reach the site of oviductal sperm reservoir and bind, transiently, to its apical epithelia surfaces (Suarez, Reference Song, Yi, Sutovsky, Meyers and Sutovsky2015). In bulls, sperm transport is believed to take between 4 and 16 h. The sperm–oviductal epithelium binding is mediated by acrosomal surface ligands and the fucose-containing glycans of the plasma membrane annexins, expressed by the oviductal epithelial cells. The sperm release upon capacitation requires proteolytic cleavage and shedding of the acrosomal binder of sperm proteins, particularly binder of sperm protein 1 (BSP1) (Suarez, Reference Song, Yi, Sutovsky, Meyers and Sutovsky2015).
Upon ovulation, signals from the ovary, oocyte–cumulus complex and its accompanying follicular fluid may be responsible for the capacitation and hyperactivation of spermatozoa bound to the oviductal sperm reservoir. Capacitation endows the reservoir-bound spermatozoa with fertilizing ability and helps them detach from the reservoir epithelium. As in other species, bull sperm capacitation encompasses cholesterol efflux, calcium influx, pH increase, remodeling of plasma membrane, and increase in tyrosine phosphorylation of sperm head and sperm tail proteins (Parrish, Reference Pan, Ju, Huang, Zhang, Qi, Gao, Zhou, Li, Wang, Zhong, Liu and Wang2013). Such structural and molecular remodeling brings about hyperactivation, manifested by an increased amplitude and frequency of flagellar movement. Chemoattractants issued from the oocyte and its surrounding cumulus cells are likely responsible for gamete recognition in mammals. Although the exact identity of the attractants involved in bovine gamete recognition is not known, progesterone from cumulus cells has been shown to fulfill such a role in other species (Oren-Benaroya et al., Reference Oko and Sutovsky2008).
Ubiquitin–proteasome system and fertilization
Ubiquitin–proteasome system (Figure 1) is likely involved in all steps of the reproductive process, including gametogenesis, fertilization, zygotic and preimplantation embryonic development, implantation, and beyond (Sutovsky, Reference Susor, Liskova, Toralova, Pavlok, Pivonkova, Karabinova, Lopatarova, Sutovsky and Kubelka2003). Protein ubiquitination is a stable, covalent post-translational modification by which the substrate proteins can be targeted for degradation by 26S proteasome, lysosome or autophagosome (Cohen-Kaplan et al., Reference Cohen-Kaplan, Livneh, Avni, Cohen-Rosenzweig and Ciechanover2016). Alternatively, protein ubiquitination, which is reversible by deubiquitinases, can serve a regulatory function, for example, through receptor endocytosis and histone code modification. Ubiquitination is a substrate-specific process that requires an energy donor (ATP) and a set of specific activator, carrier and ligase enzymes; it starts with the activation of single ubiquitin molecules by the ubiquitin-activating enzyme E1 (HUGO acronym UBA1). The activated ubiquitin is passed onto one of several carrier enzymes (E2 or UBE2) and ligated to an internal lysine residue of a substrate protein by substrate-specific ubiquitin ligases (E3 enzymes or UBE3), of which there is a great variety. Additional ubiquitin molecules are linked to the substrate-bound monoubiquitin by E3 enzymes, sometimes assisted by chain elongation factors (E4 enzymes). Substrate proteins linked to multi-ubiquitin chains of four or more ubiquitin molecules are degraded by the 26S proteasome, a multi-subunit holoenzyme composed of a 20S proteolytic core capped with a 19S proteasomal regulatory complex responsible for the recognition and removal of the multi-ubiquitin chain, and for the priming of the substrate protein before its transport to the 20S core.
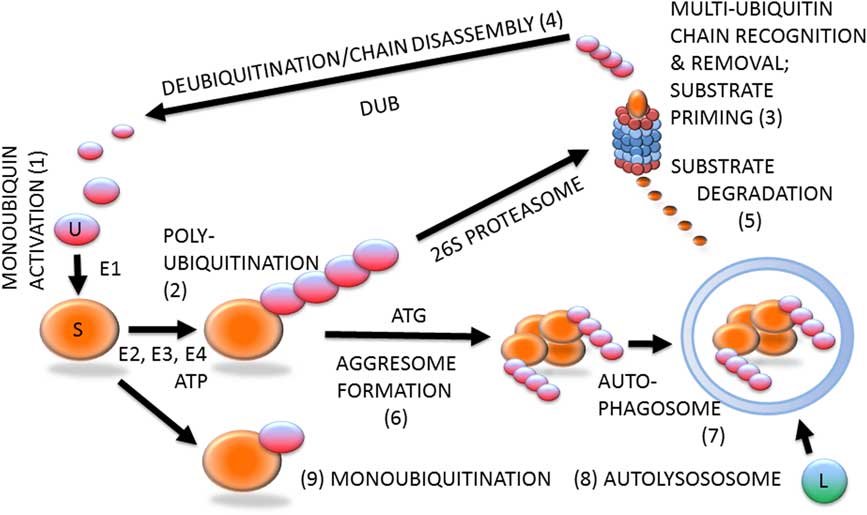
Figure 1 (Colour online) Protein ubiquitination and degradation. Unconjugated monoubiquitin (U; 1) binds covalently to its substrate (S) protein, catalyzed by ubiquitin-activating and -conjugating enzymes (E1, E2, E3) and fueled by ATP, to form a multi-ubiquitin chain (2), which is recognized by the 26S proteasome (3). The multi-ubiquitin chain is removed by the 19S proteasomal regulatory complex to be disassembled by deubiquitinating enzymes (DUB) and re-enter the pool of monoubiquitin available for conjugation (4). The substrate is degraded in the 20S proteasomal core (5). Proteasomal degradation has been implicated in sperm–zona pellucida binding, acrosomal exocytosis and sperm–zona penetration. Proteasomal proteolysis also regulates cell cycle progression, and pronuclear development and apposition in the zygote. During autophagy, which has been implicated in the regulation of early embryo development and post-fertilization sperm mitophagy, polybiquitinated substrate molecules are aggregated by autophagy-associated proteins (ATG) to form an aggresome (6). Aggresomes are recognized and engulfed by the autophagophore that encloses them to form an autophagosome (7), which upon fusion with a lysososome (L) becomes an autolysosome (8) capable of degrading the entire aggresome, or an ubiquitinated organelle such as sperm mitochondrion. The ubiquitin–proteasome system-regulated post-fertilization sperm mitophagy mediates clonal, maternal inheritance of mammalian mitochondrial genes. Monoubiquitination (9) is reversible and serves regulatory purposes relevant to gametogenesis, fertilization and early development, such as histone modification to establish the epigenetic histone code, or plasma membrane receptor internalization to change cell responsiveness to specific external stimuli/ligands.
While the proteasomes have been studied in spermatozoa of a number of mammalian species, as well as in other vertebrate and invertebrate animals (Sutovsky, Reference Sutovsky2011), little is known about the role of UPS in the early events of ungulate fertilization. Involvement of the sperm-borne proteasomes in sperm capacitation has been described in human and porcine spermatozoa (reviewed by Kerns et al., Reference Kerns, Morales and Sutovsky2016). A recent study details the role of proteasomal degradation of the A-kinase anchoring fibrous sheath protein AKAP3 (A-kinase anchoring protein 3) in bull sperm capacitation, which was correlated with capacitation and acrosomal exocytosis ability that varied among individual bulls (Hillman et al., Reference Hillman, Ickowicz, Vizel and Breitbart2013). More is known about the involvement of UPS in later events of fertilization, as will be discussed next.
Sperm–zona pellucida interactions
Bull and other mammals’ spermatozoa bind to oocyte zona pellucida (ZP) by the apical ridge of their acrosome, composed of the acrosomal matrix sandwiched between the inner and outer acrosomal membranes (IAM and OAM, respectively), with the IAM laid atop subacrosomal perinuclear theca (PT) (Oko and Sutovsky, Reference Ogushi, Palmieri, Fulka, Saitou, Miyano and Fulka2009). Bovine ZP is composed of three heavily glycosylated proteins, ZPA (homolog of human ZP2), ZPB (homolog of human ZP4/ZP3β) and ZPC (homolog of human ZP3/ZP3α), with the ZPC/ZP3 and ZPB/ZP4 being the predicted primary receptors for sperm binding and induction of acrosomal exocytosis (Yonezawa, Reference Yi, Manandhar, Sutovsky, Li, Jonakova, Oko, Park, Prather and Sutovsky2014). In this scenario, ZPA/ZP2 is proposed to be responsible for secondary/sustained sperm–zona binding (Gadella, Reference Gadella2010).
During fertilization, the spermatozoa primed by acrosomal membrane remodeling during capacitation bind to ZP surface, undergo acrosomal exocytosis enacting the loss of OAM and acrosomal matrix in all parts of the acrosome except the equatorial segment, and penetrate the ZP with only the IAM left intact after exocytosis (Gerton, Reference Gerton2002). The OAM and acrosomal matrix do not merely disperse from zona surface as previously thought; they remain associated with it in the form of acrosomal shroud. In addition to enabling sperm–zona penetration, acrosomal exocytosis prepares spermatozoa for fusion with the oolemma, by exposing oolemma-binding proteins on the sperm head equatorial segment (Cuasnicu et al., Reference Cuasnicu, Da Ros, Weigel Munoz and Cohen2016). Based on imaging of live mouse spermatozoa with green fluorescent acrosomes, it was proposed that acrosomal exocytosis is initiated before sperm contact with the oocyte, that is, during sperm migration along the female oviduct (La Spina et al., Reference La Spina, Puga Molina, Romarowski, Vitale, Falzone, Krapf, Hirohashi and Buffone2016). It is not clear whether such concept applies to bovine fertilization. Furthermore, one has to be cautious about interpreting such observations as the true acrosomal exocytosis, as opposed to the remodeling of acrosomal membranes induced by sperm capacitation.
The signaling cascade of acrosomal exocytosis, induced by the binding of sperm head ligands to ZPC, activates signaling through phospholipases, and both the tyrosine and serine/threonine protein kinases which leads to the influx of external calcium, depolymerization of the actin microfilaments scaffolding the acrosome, and the activation of membrane fusion pathways leading to the fusion and hybridizing vesiculation of plasma membrane and OAM, resulting in the formation of acrosomal shroud (Belmonte et al., Reference Belmonte, Mayorga and Tomes2016; Romarowski et al., Reference Rodgers, Morgan, Leu and Bale2016).
Bull spermatozoa display all three major proteasomal core enzymatic activities (Pizarro et al., Reference Perry, Wakayama, Cooke and Yanagimachi2004). Proteasomes are localized in all three layers of the mammalian sperm acrosome, including OAM, matrix and IAM (Sutovsky et al., Reference Sutovsky, Plummer, Baska, Peterman, Diehl and Sutovsky2004; Zimmerman et al., Reference Zhao, Wang, Hao, Li, Zhao, Yan, Wang, Du, Wang, Liu, Pang and Zhu2011). Acrosomal proteasomes may serve a dual purpose during sperm–ZP interactions, first as the mediators of acrosomal exocytosis and subsequently as proteases contributing to localized degradation of the zona matrix that results in the formation of the fertilization slit (Zimmerman et al., Reference Zhao, Wang, Hao, Li, Zhao, Yan, Wang, Du, Wang, Liu, Pang and Zhu2011; Miles et al., Reference Massicotte, Coenen, Mourot and Sirard2013). Bull spermatozoa capacitated in the presence of proteasomal inhibitor epoxomicin had reduced ability to undergo acrosomal exocytosis induced in vitro by progesterone and to fertilize oocytes in vitro (Sanchez et al., Reference Saldivar-Hernandez, Gonzalez-Gonzalez, Sanchez-Tusie, Maldonado-Rosas, Lopez, Trevino, Larrea and Chirinos2011). Addition of proteasomal inhibitor MG132 into in vitro fertilization (IVF) medium significantly reduced the rate of bovine fertilization (Rawe et al., Reference Rassoulzadegan, Grandjean, Gounon, Vincent, Gillot and Cuzin2008). Solubilized ZP protein degradation by bull sperm proteasomes has not been described, to our knowledge, but the degradation of solubilized ZP proteins by sperm proteasomes has been shown in humans (Saldivar-Hernandez et al., Reference Sakatani, Yamanaka, Balboula, Takenouchi and Takahashi2015) and Japanese quail (Sasanami et al., Reference Sanchez, Deppe, Schulz, Bravo, Villegas, Morales and Risopatron2012). In agreement with their proposed role in sperm–zona interactions, a subpopulation of proteasomes are exposed on the bull sperm surface and prominently represented in the sperm surface proteome (Byrne et al., Reference Byrne, Leahy, McCulloch, Colgrave and Holland2012), as has also been described in other mammals. It is thus possible that a third function could be assigned to them: that of secondary or alternative receptor for ZP. In agreement the proteasomes of human and porcine spermatozoa are part of a large multimeric zona-binding complex that also contains chaperonins, zona-binding receptors, acrosin and its binding protein, and seminal plasma proteins (Redgrove et al., Reference Rawe, Diaz, Abdelmassih, Wojcik, Morales, Sutovsky and Chemes2011; Kongmanas et al., Reference Kongmanas, Kruevaisayawan, Saewu, Sugeng, Fernandes, Souda, Angel, Faull, Aitken, Whitelegge, Hardy, Berger, Baker and Tanphaichitr2014). In summary, UPS is likely involved in multiple events during sperm interactions with oocyte ZP.
Gamete adhesion and fusion, and sperm incorporation
Commonly referred to as gamete fusion, the process by which the fertilizing spermatozoon contacts and passes through the oocyte plasma membrane, the oolema, is likely more complex than originally thought. It is thus prudent to distinguish between its adhesion and fusion phase (reviewed in Sutovsky, Reference Sutovsky2009). Moreover, important is to note that gamete adhesion and fusion is enabled by sperm capacitation and acrosomal exocytosis during which proteins involved in the sperm–oolemma interactions become exposed on the sperm head equatorial segment (e.g. equatorin/MN-9 (Manandhar and Toshimori, Reference Maddox-Hyttel, Svarcova and Laurincik2001), CD9 (Ito et al., Reference Ito, Yamatoya, Yoshida, Maekawa, Miyado and Toshimori2010) and IZUMO1 (Satouh et al., Reference Satouh, Nozawa and Ikawa2012)).
Early studies implicated sperm disintegrins and oolemma integrins in sperm–oolemma adhesion, though none such receptors have fusogenic properties. Gene knock-out studies demonstrated that these molecules are either non-essential or compensable in mouse fertilization (Primakoff and Myles, Reference Platts, Dix, Chemes, Thompson, Goodrich, Rockett, Rawe, Quintana, Diamond, Strader and Krawetz2007). Alternatively proposed sperm receptors on the oollema, tetraspanins CD9 and CD81 were knocked out, resulting in reduced fertility of single gene knock-out and complete infertility of double knock-out (Benammar et al., Reference Benammar, Ziyyat, Lefevre and Wolf2016). The most convincing evidence for gamete adhesion receptors came with the discovery of the sperm-specific immunoglobulin family cell adhesion protein IZUMO (Inoue et al., Reference Inoue, Ikawa, Isotani and Okabe2005) and its oocyte-binding partner, the folate-binding receptor JUNO (Bianchi et al., Reference Bianchi, Doe, Goulding and Wright2014). Mice null in either of these proteins are infertile. Both IZUMO (Fukuda et al., Reference Fukuda, Sakase, Fukushima and Harayama2016) and JUNO (Zhao et al., Reference Zhang, Zhang, Yang, Zhang, Ju, Wang, Jiang, Sun, Huang, Zhong and Wang2017) are present in bovine gametes, but there is not yet a direct proof of their involvement in bovine gamete adhesion. A plausible model for bovine/mammalian sperm–oolemma interaction is thus the tetraspanin web on the oolemma, in which said tetraspanins are the organizers of various receptors for sperm equatorial segment ligands, including integrins, JUNO and the downstream signaling proteins that relay the adhesion signals to oocyte cortical cytoskeleton (Sutovsky, Reference Sutovsky2009; Wright and Bianchi, Reference Tung, Hansen, Ban, Loktev, Summers, Adler and Jackson2015). Accordingly, microfilaments in the bovine oocyte microvilli (Sutovsky et al., Reference Sutovsky, Aarabi, Miranda-Vizuete and Oko1996) and cortex are required for physical incorporation of the sperm head and tail in the ooplasm and probably for signaling in response to the initial adhesion between sperm and oocyte plasma membranes, preceding their fusion, as exemplified by gamete adhesion induced signaling via oocyte proline-rich tyrosine kinase (PYK2) in zebra fish (Sharma and Kinsey, Reference Selvaraju, Parthipan, Somashekar, Kolte, Krishnan Binsila, Arangasamy and Ravindra2012). Consequently, microfilament disrupting drug cytochalasin B prevents sperm incorporation without interfering with sperm–oolema fusion or with the release of sperm PT proteins that trigger oocyte activation (Sutovsky et al., Reference Sutovsky, Aarabi, Miranda-Vizuete and Oko1996; Sutovsky et al., Reference Sutovsky, Neuber and Schatten1997; Sutovsky et al., Reference Sutovsky and Schatten2003).
Cortical actin cytoskeleton appears to be regulated in part by UPS. Interference with ubiquitin C-terminal hydrolase (UCHL1), enriched in the mammalian oocyte cortex can thus alter polyspermy rate in vitro (Yi et al., Reference Wu, Sutovsky, Xu, van der Spoel, Platt and Oko2007; Mtango et al., Reference Mtango, Sutovsky, Susor, Zhong, Latham and Sutovsky2011b). The expression of Uchl1 gene in bovine zygotes was altered by heat stress (Sakatani et al., Reference Romarowski, Luque, La Spina, Krapf and Buffone2015), wherein the UCHL1 inhibition impaired migration and exocytosis of cortical granules, the major line of defense against polyspermic fertilization (Susor et al., Reference Suarez2010). A related deubiquitinase, UCHL3, is associated with oocyte meiotic spindle and may regulate oocyte maturation (Mtango et al., Reference Miles, O’Gorman, Zhao, Samuel, Walters, Yi, Sutovsky, Prather, Wells and Sutovsky2014 and Reference Mtango, Latham and Sutovsky2011a). Altogether, it is likely that the UPS mainly regulates the later events of sperm–oocyte adhesion and fusion; in particular, sperm incorporation and activation of the anti-polyspermy defense.
Oocyte activation
While adhesion and fusion between oocyte and sperm plasma membranes initially occurs within the sperm head equatorial segment, it quickly extends distally to post-acrosomal sheath (Sutovsky et al., Reference Sutovsky and Schatten2003). The current paradigm of oocyte activation assumes that a soluble sperm factor(s) released from post-acrosomal PT triggers calcium release from oocyte endoplasmic reticulum that triggers a signaling cascade guiding the completion of oocyte meiosis, mobilization of anti-polyspermy defense, zygotic/pronuclear development and ultimately embryo cleavage (Oko et al., Reference Oko, Aarabi, Mao, Balakier and Sutovsky2017). Initial studies of this process in mammals were conducted in rodent and bovine IVF systems. It was demonstrated that freeze-thaw sperm extracts activate hamster oocytes (Swann, Reference Sutovsky, Manandhar, McCauley, Caamano, Sutovsky, Thompson and Day1990). The removal and solubilization of sperm PT was shown to occur during bovine IVF even when sperm incorporation was prevented by treatment with actin depolymerizing drug cytochalasin B (Sutovsky et al., Reference Sutovsky, Neuber and Schatten1997).
While the dissolution of PT can be hindered by protease inhibitors (Perry et al., Reference Parthipan, Selvaraju, Somashekar, Arangasamy, Sivaram and Ravindra2000), it is not clear whether oocyte proteasomes are directly involved. However, UPS clearly regulates zygotic cell cycle by mediating the oocyte spindle exit from metaphase II through degradation of EMI1 (synonym F-boax protein 5/FBOX5) protein that maintains the activity of oocyte cytostatic factor and by degrading the cyclin component of the M-phase promoting factor to assure successful metaphase–anaphase transition (Glotzer et al., Reference Glotzer, Murray and Kirschner1991; Tung et al., Reference Tovich and Oko2005; Karabinova et al., Reference Karabinova, Kubelka and Susor2011).
Stepwise extraction of PT from mouse sperm heads progressively eliminated their ability to trigger oocyte activation after intracytoplasmic sperm injection, and the activity was shown to be associated with specific fractions of PT that had differential extraction resistance (Kimura et al., Reference Kimura, Yanagimachi, Kuretake, Bortkiewicz, Perry and Yanagimachi1998; Perry et al., Reference Parthipan, Selvaraju, Somashekar, Arangasamy, Sivaram and Ravindra2000). Eventually, the currently favored sperm-borne oocyte factor (SOAF) candidate, phospoholipase C zeta 1 (PLCZ1) was identified in silico and shown to trigger oocyte calcium oscillations when a recombinant protein or its messenger RNA (mRNA) were injected in metaphase II oocytes (Saunders et al., Reference Satouh, Inoue, Ikawa and Okabe2002). Persistent concerns about localization of PLCZ outside of post-acrosomal sheath (PAS) and its promiscuity with regard to oocyte-activating sperm cytosol fractions prompted a search for alternative or synergistic sperm SOAF components, chief among them male germline-specific WW-domain-binding WBP2NL (ww-domain binding protein 2 N-terminal like) (alias post-acrosomal WW-domain binding protein (PAWP)) and its somatic cousin WBP2 (Wu et al., Reference Wu, Sutovsky, Manandhar, Xu, Katayama, Day, Park, Yi, Xi, Prather and Oko2007b; Oko et al., Reference Oko, Aarabi, Mao, Balakier and Sutovsky2017). As in other mammals, PAWP is prominent in bull sperm PT where it is transported from the cytoplasmic lobe along the caudal manchette in the elongating spermatid phase of spermiogenesis, coinciding with the spermatids’ acquisition of oocyte-activating ability (Wu et al., Reference Wright and Bianchi2007a). In the mouse, targeting neither the Plcz1 (Hachem et al., Reference Hachem, Godwin, Ruas, Lee, Ferrer Buitrago, Ardestani, Bassett, Fox, Navarrete, de Sutter, Heindryckx, Fissore and Parrington2017) nor the Wbp2nl (Satouh et al., Reference Sathananthan, Tatham, Dharmawardena, Grills, Lewis and Trounson2015) gene appears to eliminate male fertility. Intriguingly, increased expression of Wbp2nl gene is observed in the testis of Plcz1 null mice (Hachem et al., Reference Hachem, Godwin, Ruas, Lee, Ferrer Buitrago, Ardestani, Bassett, Fox, Navarrete, de Sutter, Heindryckx, Fissore and Parrington2017), suggesting compensation in the knock-out mouse and possible synergy between the two gene products in wild type animals.
With regard to bull fertility, individual sperm levels of WBP2NL vary between sires and within individual ejaculates. Excessive amounts of WBP2NL were found in macrocephalic spermatozoa with abnormally large sperm head and PAS, while certain morphologically deviant spermatozoa, particularly the ones with abnormally shaped sperm heads and those positive for ubiquitin, in this case applied as a sperm quality biomarker (Sutovsky et al., Reference Sutovsky, Simerly, Hewitson and Schatten2015), lacked WBP2NL altogether (Kennedy et al., Reference Kennedy, Krieger, Sutovsky, Xu, Vargovic, Didion, Ellersieck, Hennessy, Verstegen, Oko and Sutovsky2014). High abundance of Plcz1 mRNA in spermatozoa was also associated with fertility of Holstein bulls (Kasimanickam et al., Reference Kasimanickam, Kasimanickam, Arangasamy, Saberivand, Stevenson and Kastelic2012), a breed in which Plcz1 gene polymorphism also appears to be a contributing factor (Pan et al., Reference Ostermeier, Miller, Huntriss, Diamond and Krawetz2013). In conclusion, while the identity of the sperm-borne oocyte-activating factor is still being researched, there appears to be a consensus that it is a partially soluble protein (or several proteins) in the sperm head PT.
Zygotic centrosome assembly and pronuclear apposition
Somatic cell centrosome is a cellular microtubule organizing center composed of two perpendicularly oriented, barrel-shaped (pinwheel-like on cross-section) centrioles and pericentriolar material (PCM) with the ability to nucleate tubulins to polymerize into microtubules. In bovine gametes, the centrosome is reduced in a complementary fashion, with oocytes retaining PCM but devoid of centrioles, and spermatozoa carrying a single, duplication-competent centriole (or no centrioles, as in rodents) and little, if any PCM (Sathananthan et al., Reference Sasanami, Sugiura, Tokumoto, Yoshizaki, Dohra, Nishio, Mizushima, Hiyama and Matsuda1997; Manandhar et al., Reference Manandhar, Schatten and Sutovsky2005). As in other non-rodent Eutherian mammals, bull sperm centrosome is reduced to a single centriole with little or no PCM during spermiogenesis/spermatid elongation in the testis. This centriole is housed within the complex structure of the sperm tail connecting piece and thus termed the proximal centriole (Sutovsky et al., Reference Sutovsky, Aarabi, Miranda-Vizuete and Oko1996; Simerly et al., Reference Shin and Kim1999). The distal centriole is degraded during spermiogenesis, and only some remnants of centrosomal material may be found within the distal centriolar vault (Manandhar et al., Reference Manandhar, Schatten and Sutovsky2005). Failure of proper centrosome remodeling during spermiogenesis has been associated with the tail stump defect in infertile bulls (Blom and Birch-Andersen, Reference Blom and Birch-Andersen1980).
At fertilization, soon after sperm head and sperm tail midpiece incorporation, the striated columns caging the centriole break up, a disassembly process required for the formation of zygotic centrosome and leading up to the organization of microtubule-made sperm aster responsible for the apposition of paternal and maternal pronuclei (Sutovsky et al., Reference Sutovsky, Aarabi, Miranda-Vizuete and Oko1996; Sutovsky and Schatten, Reference Sutovsky1997; Sutovsky, Reference Sutovsky2010) (Figure 2). The gamma-tubulin (TUBG) containing, microtubule-nucleating PCM is recruited from bovine oocyte cytoplasm by the liberated sperm centriole (Shin and Kim, Reference Sharma, Conine, Shea, Boskovic, Derr, Bing, Belleannee, Kucukural, Serra, Sun, Song, Carone, Ricci, Li, Fauquier, Moore, Sullivan, Mello, Garber and Rando2003). The ability of bull spermatozoa to form sperm aster after IVF has been correlated with the developmental potential of bovine IVF embryos in artificial insemination (AI) bulls (Navara et al., Reference Nakamura, Terada, Horiuchi, Emuta, Murakami, Yaegashi and Okamura1996; Hara et al., Reference Hara, Tagiri, Hirabayashi and Hochi2015; Hochi, Reference Hochi2016). Inspired by those findings, bovine oocytes have been injected with human spermatozoa as a way of testing for human male infertility due to sperm centrosomal insufficiency (Nakamura et al., Reference Mtango, Sutovsky, Vandevoort, Latham and Sutovsky2001). It was noted that in both human and bovine zygote, proteasomal activity is required for the formation of sperm aster, possibly because proteasomes are present in the sperm tail connecting piece and necessary for its disassembly that liberates the sperm centriole and makes it competent to recruit centrosomal proteins from ooplasm (Rawe et al., Reference Rassoulzadegan, Grandjean, Gounon, Vincent, Gillot and Cuzin2008). Altogether, it is well documented that bull spermatozoa contribute crucial components of the zygotic centrosome at fertilization, and such a mode of centrosomal inheritance is similar to primates and other ungulates, but different from rodents.
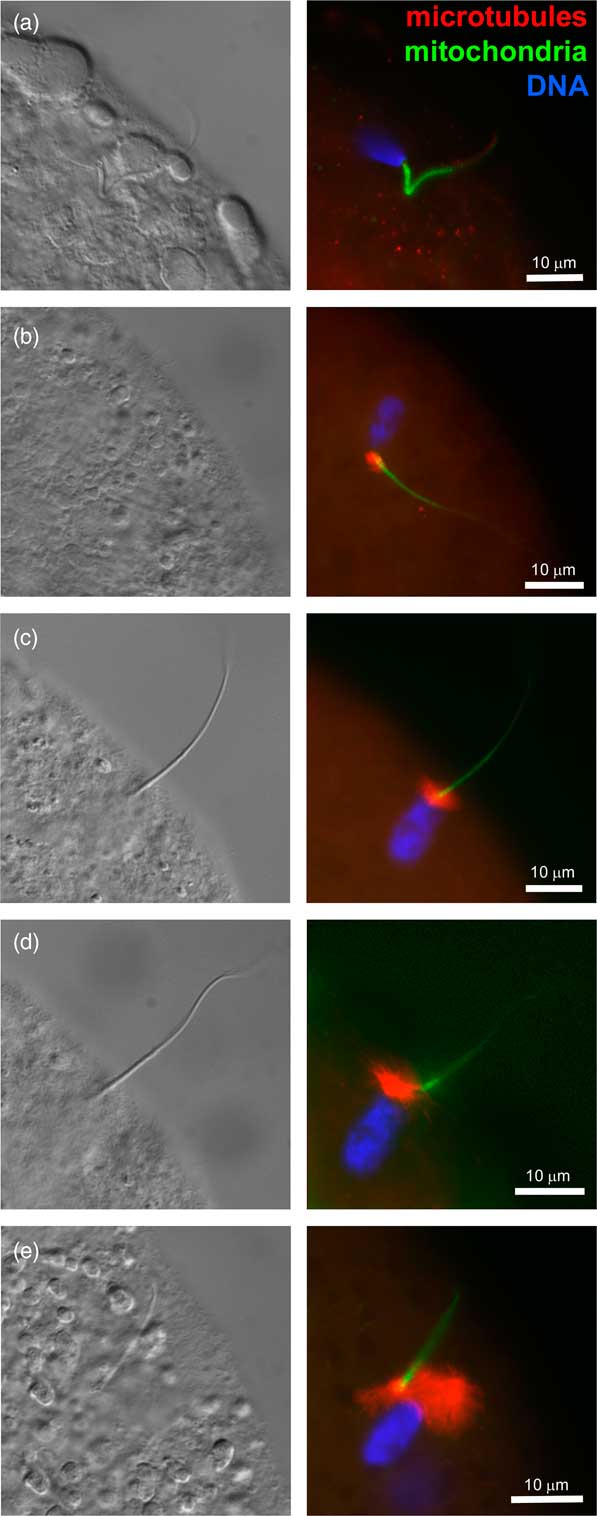

Figure 2 (Colour online) Progression of sperm incorporation, sperm nucleus decondensation and sperm aster formation during bovine fertilization in vitro. In all panels, sperm aster microtubules were labeled with monoclonal anti-tubulin antibody E7 (red), sperm mitochondria were pre-labeled with vital stain MitoTracker Green FM (green) before IVF and DNA in the sperm nucleus (blue) was counterstained with 2-(4-amidinophenyl)-1H-indole-6-carboxamidine. Left columns show parfocal bright-field images acquired with differential interference contrast optics (DIC). The samples were examined and photographed under a Nikon Eclipse 800 epifluorescence microscope (Nikon Instruments Inc., Melville, NY, USA) with Cool Snap camera (Roper Scientific, Tucson, AZ, USA) and MetaMorph software (Universal Imaging Corp., Downingtown, PA, USA). Images were edited and contrast balanced by Adobe Photoshop CS5 (Adobe Systems Inc., San Jose, CA, USA).
Pronuclear development
Before sperm–oolemma fusion, bull sperm head found within oocyte periviteline space is composed of a nucleus covered with a reduced nuclear envelope and PT all around, and IAM in the acrosomal region and plasma membrane over the equatorial segment and post-acrosomal region. With the exception of a small contingent of somatic and sperm-specific histones in the nucleus and PT (Aul and Oko, Reference Aul and Oko2002; Tovich and Oko, Reference Thompson, Ramalho-Santos and Sutovsky2003) sperm DNA is packaged with protamines, mainly protamine 1 (PRM1) (Balhorn, Reference Balhorn2007). Proper sperm protamination correlates with bull fertility in AI service (Dogan et al., Reference Dogan, Vargovic, Oliveira, Belser, Kaya, Moura, Sutovsky, Parrish, Topper and Memili2015).
The sperm nucleus is demembranated in a stepwise fashion at sperm–oolemma fusion and entry in the ooplasm, when the sperm plasma membrane comingles with oolemma, PT, first in the post-acrosomal and then in the subacrosomal region spreads across the ooplasm, and the sperm nuclear envelope, devoid of nuclear pore complexes and nuclear lamina, dissolves (Sutovsky et al., Reference Sutovsky, Neuber and Schatten1997; Sutovsky et al., Reference Sutovsky, Oko, Hewitson and Schatten1998). Next, protamines, inserted during sperm nucleus compaction at spermiogenesis, are removed and replaced with oocyte-specific histones, which themselves will later be replaced with somatic cell type histones (Gao et al., Reference Gao, Chung, Parseghian, King, Adashi and Latham2004). At the time of protamine–histone exchange reversal, the nuclear envelope is reconstituted by membrane vesicle fusion and nuclear lamina formation on both maternal and paternal chromatin, and nuclear pores are reinserted, possibly from a premade stock of ooplasmic annulate lamellae (Sutovsky et al., Reference Sutovsky, Oko, Hewitson and Schatten1998).
As the maternal and paternal pronuclei grow, maternally stored nucleolar proteins re-enter the nuclear compartment to form nucleolus precursor bodies that will be remodeled into active ribosomal RNA producing nucleoli at the time of major zygotic genome activation (four to eight cells in bovine) (Maddox-Hyttel et al., Reference Liu, Pang, Chiu, Wong, Lao, Lee and Yeung2007; Ogushi et al., Reference Odhiambo, Sutovsky, DeJarnette, Marshall and Sutovsky2008). The zygote-driven remodeling of bovine sperm nucleus is facilitated by multiple ooplasmic factors, including the disulfide bond reducing peptide glutathione, histones, lamins, nucleoporins and nucleoplasmin (Sutovsky and Schatten, Reference Sutovsky1997; Burns et al., Reference Burns, Viveiros, Ren, Wang, DeMayo, Frail, Eppig and Matzuk2003). Such an extensive restructuring is aided by oocyte proteasomes that accumulate in both pronuclei from the onset of pronuclear development (Figure 3). In addition to pronuclear accumulation, proteasomes are abundant in mammalian zygote cytoplasm, as shown in the zygotes of a transgenic pig expressing proteasomes tagged with green fluorescent protein (Miles et al., Reference Massicotte, Coenen, Mourot and Sirard2013).
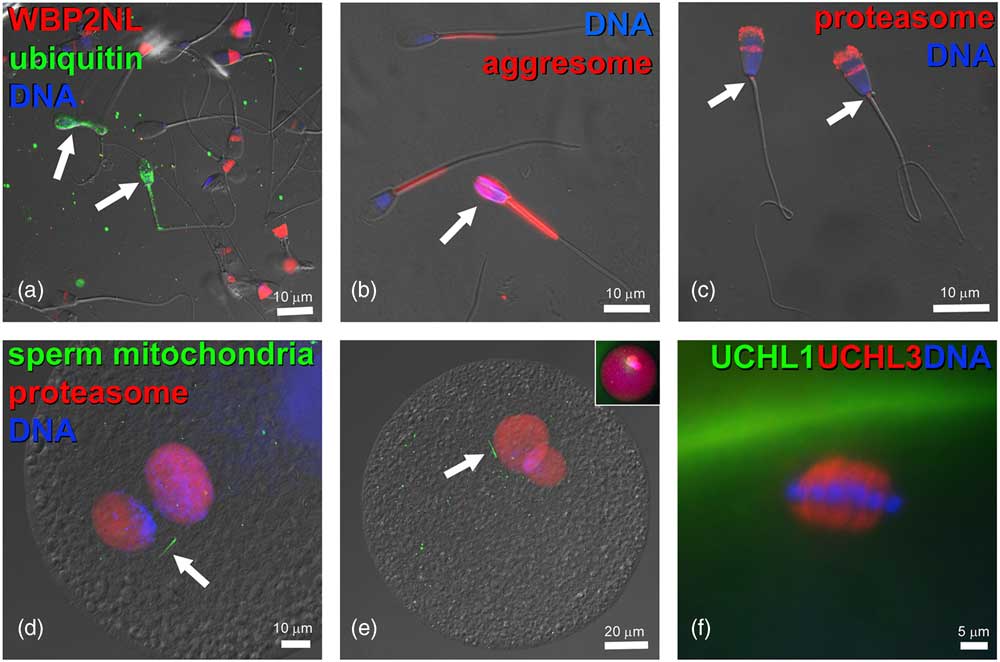
Figure 3 (Colour online) Components of ubiquitin–proteasome system in bovine gametes and embryos. (a) Bull spermatozoa (arrows) with abnormal phenotypes are ubiquitinated on their surface (green) and lack the WBP2NL/PAWP protein (red) present at varied levels in the post-acrosomal sheaths of surrounding morphologically normal spermatozoa. (b) Aggresome-binding ProteoStat probe (red) binds exclusively to mitochondrial sheaths of normal spermatozoa but detects aggresomes in the deformed sperm head of an abnormal spermatozoon (arrow). (c) Proteasomes (red) detected in bull sperm head acrosome and sperm tail connecting piece (arrows) by polyclonal antibody recognizing the 20S proteasomal core subunit PSMB10 (PW8150, Enzo Lifesciences, Ann Arbor, MI, USA). (d,e) Proteasomes (red) in the apposing (d), and apposed (e) maternal and paternal pronuclei of bovine zygotes, detected by a polyclonal antibody recognizing multiple 20S proteasomal core subunits (PW8155, Enzo Lifesciences). Inset in (e) shows the same zygote from which the differential interference contrast layer has been subtracted and red fluorescence brightened to reveal cytoplasmic proteasome labeling. Bovine oocytes were fertilized by spermatozoa pre-labeled with MitoTracker Green FM (green) to detect sperm tail mitochondrial sheaths (arrows). (f) Detection of deubiquitinases ubiquitin C-terminal hydrolase L1 (UCHL1) (green) and UCHL3 (red) in the bovine oocyte cortex and meiotic spindle, respectively. DNA in all panels was counterstained with 2-(4-amidinophenyl)-1H-indole-6-carboxamidine. The samples were examined and photographed under a Nikon Eclipse 800 epifluorescence microscope with Cool Snap camera and MetaMorph software. Images were edited and contrast balanced by Adobe Photoshop CS5.
Proteasomal inhibitors alter male pronuclear chromatin (Huo et al., Reference Huo, Fan, Liang, Yu, Zhong, Chen and Sun2004; Rawe et al., Reference Rassoulzadegan, Grandjean, Gounon, Vincent, Gillot and Cuzin2008) possibly interfering with protamine removal and/or oocyte–somatic histone swap. The ensemble of epigenetic histone marks, the zygotic histone code, which is reset at the onset of embryo development, is also influenced by UPS (Nevoral and Sutovsky, Reference Navara, First and Schatten2017), possibly through UPS regulation of sirtuin 1 (SIRT1), a histone deacetylase (Adamkova et al., Reference Adamkova, Yi, Petr, Zalmanova, Hoskova, Jelinkova, Moravec, Kralickova, Sutovsky, Sutovsky and Nevoral2017). The involvement of UPS extends beyond the zygote. Genes encoding UPS components such as proteasomal subunits, ubiquitin-conjugating enzymes and ubiquitin ligases likely regulate later events of bovine preimplantation development, including the major zygotic genome activation and blastocyst differentiation (Massicotte et al., Reference Manandhar and Toshimori2006; Adjaye et al., Reference Adjaye, Herwig, Brink, Herrmann, Greber, Sudheer, Groth, Carnwath, Lehrach and Niemann2007). It can be concluded that the UPS regulates multiple facets of early zygotic development in bovines and other mammals.
Mitochondrial inheritance and the fate of sperm tail accessory structures
At fertilization, bull spermatozoa carry into ooplasm several dozen mitochondria organized into a helical structure of the mitochondrial sheath in the sperm tail midpiece. Early studies demonstrated that bull sperm mitochondria are sought out and targeted for degradation by the fertilized oocyte, typically lasting only up to 4-cell stage (Sutovsky et al., Reference Sutovsky, Aarabi, Miranda-Vizuete and Oko1996). Such targeted mitophagy, not observed in interspecies zygotes of domestic cattle oocytes and gaur (Bos gaurus) spermatozoa (Sutovsky et al., Reference Sutovsky, Aarabi, Miranda-Vizuete and Oko1996) may be developmentally advantageous due to the high potential for sperm mitochondrial DNA (mtDNA) damage, mutation and deletion during spermatogenesis and fertilization (reviewed in Song et al., Reference Simerly, Zoran, Payne, Dominko, Sutovsky, Navara, Salisbury and Schatten2014). Failure of this mechanism in cattle, causing heteroplasmy by leakage of paternal mtDNA, could affect offspring health and production traits influenced by mitochondrial fitness, such as milk protein content and meat quality (Gibson et al., Reference Gibson, Freeman and Boettcher1997).
Bovine post-fertilization mitophagy depends on the progression of the zygotic cell cycle, and it is not a coincidence that both processes are co-regulated by UPS. Consequently, sperm mitochondria become ubiquitinated inside the oocyte (Sutovsky et al., Reference Sutovsky, Manandhar, Wu and Oko1999) and targeted for degradation by proteasomal proteolysis and lysosomal/autophagic machinery (Sutovsky et al., Reference Sutovsky, Moreno, Ramalho-Santos, Dominko, Thompson and Schatten2000). Recent studies in domestic pig zygotes indicate that the central role in sperm mitophagy is played by the ubiquitin-binding autophagy receptor SQSTM1 and the proteasome-serving ubiquitinated protein dislocase valosin containing protein (VCP) (Song et al., Reference Song, Ballard, Yi and Sutovsky2016). The participation of VCP explains why proteasomal inhibitors block sperm mitophagy, but proteasomes seldom associate directly with the sperm mitochondrial sheath inside the zygote. Some mitochondrial membrane and matrix proteins, such as prohibitin (Thompson et al., Reference Taylor, Schnabel and Sutovsky2003), are already tagged with ubiquitin in sperm mitochondria in bull spermatozoa, and the properties of the heavily disulfide bond cross-linked, ubiquitinated sperm mitochondrial sheath are similar to those of an aggresome, a ubiquitinated protein aggregate subject to autophagy in somatic cells (Thompson et al., Reference Taylor, Schnabel and Sutovsky2003; Kennedy et al., Reference Kennedy, Krieger, Sutovsky, Xu, Vargovic, Didion, Ellersieck, Hennessy, Verstegen, Oko and Sutovsky2014).
Attention has also been given to sperm tail accessory structures including the fibrous sheath covering the principal piece and the axonemal outer dense fibers running the whole length of the sperm tail except the connecting piece, where they are transformed into striated columns. While the fibrous sheath appears to dissolve very quickly within the first few hours of bull sperm incorporation, the outer dense fibers and microtubules may persist past one cell stage (Sutovsky et al., Reference Sutovsky, Aarabi, Miranda-Vizuete and Oko1996). Given the minute size of bovine sperm tail, its resident proteins are not likely to be the source of nutrition for the embryo, as has been proposed in Drosophila, a species endowed with an enormous sperm tail. However, the densely amalgamated tail accessory structures could harbor paternal RNAs that affect embryo development, as will be discussed below.
Research on sperm ubiquitin has been extended beyond sperm mitochondrion ubiquitination and led to the discovery of sperm surface ubiquitination in defective bull spermatozoa (Sutovsky et al., Reference Sutovsky, Moreno, Ramalho-Santos, Dominko, Simerly and Schatten2001; Baska et al., Reference Baska, Manandhar, Feng, Agca, Tengowski, Sutovsky, Yi and Sutovsky2008), and later on to identification of aggresomes outside bull sperm mitochondrial sheath (Kennedy et al., Reference Kennedy, Krieger, Sutovsky, Xu, Vargovic, Didion, Ellersieck, Hennessy, Verstegen, Oko and Sutovsky2014). Sperm surface ubiquitination has been validated as a sperm quality biomarker in bulls and other species (Sutovsky et al., Reference Sutovsky, Simerly, Hewitson and Schatten2015) and correlated with sperm acrosome (Odhiambo et al., Reference Nevoral and Sutovsky2011) and DNA integrity (Sutovsky et al., Reference Sutovsky, Navara and Schatten2002), expression of fertility-related sperm proteins (platelet activating factor receptor) (Sutovsky et al., Reference Sutovsky, Moreno, Ramalho-Santos, Dominko, Simerly and Schatten2007) and PAWP/WBP2NL, which also correlated with sperm aggresome content in AI bulls (Kennedy et al., Reference Kennedy, Krieger, Sutovsky, Xu, Vargovic, Didion, Ellersieck, Hennessy, Verstegen, Oko and Sutovsky2014). Identification of sperm surface ligands unique to defective spermatozoa enabled the development of nanopurification for the improvement of bull AI semen (Odhiambo et al., Reference Odhiambo, DeJarnette, Geary, Kennedy, Suarez, Sutovsky and Sutovsky2014). At present, there is already enough evidence to conclude that the clonal maternal inheritance of bovine mitochondrial genes is regulated by UPS through proteasomal and autophagosomal protein/organelle degradation.
Regulatory molecules in bull spermatozoa that affect early embryo development
Mammalian spermatozoa carry their own transcriptome (Ostermeier et al., Reference Ostermeier, Dix, Miller, Khatri and Krawetz2004), which is likely reflective of genome activity before germ cell meiosis and particularly early after meiosis, when round spermatids remain transcriptionally active until the process of histone–protamine exchange is initiated. Approximately 3500 species of mRNA alone are found in human spermatozoa (Ostermeier et al., Reference Oren-Benaroya, Orvieto, Gakamsky, Pinchasov and Eisenbach2002), among which the genes within UPS appear to be dysregulated in teratospermic infertile men (Platts et al., Reference Pizarro, Pasten, Kong and Morales2007). Recent sequencing effort revealed transcripts of over 13 000 genes in bull spermatozoa (Selvaraju et al., Reference Saunders, Larman, Parrington, Cox, Royse, Blayney, Swann and Lai2017). Beside mRNA, non-coding RNAs such as micro-RNA (miR), long Piwi-interacting RNAs and small nucleolar RNAs could be carried by bull spermatozoa and influence embryo development and inheritance (Hossain et al., Reference Hossain, Sohel, Schellander and Tesfaye2012). The miR-196b, targeting homeobox genes, appears to be the most abundant miR in bull spermatozoa (Selvaraju et al., Reference Saunders, Larman, Parrington, Cox, Royse, Blayney, Swann and Lai2017). The miR content may vary between a high- and low-motility spermatozoa (Capra et al., Reference Capra, Turri, Lazzari, Cremonesi, Gliozzi, Fojadelli, Stella and Pizzi2017), as well as between bulls of varied fertility (Govindaraju et al., Reference Govindaraju, Uzun, Robertson, Atli, Kaya, Topper, Crate, Padbury, Perkins and Memili2012; Fagerlind et al., Reference Fagerlind, Stalhammar, Olsson and Klinga-Levan2015).
Protamine Prm1 is among the most abundant bull sperm mRNAs (Card et al., Reference Card, Anderson, Zamberlan, Krieger, Kaproth and Sartini2013), probably reflecting its abundance in spermatids, wherein its product is required for the proper packaging of sperm DNA. Sperm levels of several other transcripts, including ubiquitin-conjugating enzyme gene Ube2e3 correlate with bull AI fertility (Parthipan et al., Reference Parrish2017). This and other UPS-related sperm-borne mRNAs could be translated by the zygote, for example, to assist the post-fertilization clean-up of maternally stored proteins.
Some studies suggest the lone presence of spermatozoa and seminal plasma can stimulate specific transcriptional response in the female reproductive system that may be favorable for the establishment of pregnancy. Seminal plasma may influence gene expression in uterine epithelia directly and indirectly regulate embryo development through uterine secretion of embryotrophic growth factors (Bromfield et al., Reference Bromfield, Schjenken, Chin, Care, Jasper and Robertson2014). Such findings warrant further inquiry into the transcriptome of bovine seminal plasma, from which some RNA containing protein complexes could be adsorbed on the bull sperm surface and carried past cervix during mating, or be introduced directly in uterus with extended semen during AI.
Mouse studies provide additional insight into potential roles of sperm RNAs in zygotic development and inheritance. Some of the sperm mRNA contributed to zygote could be translated to regulate early development and even to promote the RNA-mediated non-Mendelian inheritance, as suggested for the mouse sperm transmitted mutated Kit allele dominating the mRNA produced from wild type allele (Rassoulzadegan et al., Reference Primakoff and Myles2006). Another example, miR-34c is present in spermatozoa but absent from oocytes and appears to be required for embryo cleavage (Liu et al., Reference Spallanzani2011). Pronuclear injection of miR-24 caused paramutation in the mouse zygote coinciding with heritable histone modification associated with promoter region of transcription factor Sox9 gene (Grandjean et al., Reference Grandjean, Gounon, Wagner, Martin, Wagner, Bernex, Cuzin and Rassoulzadegan2009). Pertinent to the regulation of zygotic development by UPS, injection of eight different sperm derived miR in mouse zygotes caused degradation of maternal, oocyte stored mRNA that, among other effects on gene expression, affected the genes encoding aforementioned, UPS-regulated SIRT1 protein and ubiquitin ligase UBE3A, both involved in histone code modification (Rodgers et al., Reference Redgrove, Anderson, Dun, McLaughlin, O’Bryan, Aitken and Nixon2016). Similarly, sperm-borne transfer RNA fragments, acquired during epididymal passage seem to repress expression of specific genes in the murine embryo and may mediate epigenetic influences between father and offspring (Sharma et al., Reference Sharma and Kinsey2017). Altogether, new evidence indicates that RNAs with both the protein coding and regulatory functions are present in bull spermatozoa and contributed to the zygote at fertilization.
Effects of paternal genome on conception rate and offspring fertility
Gene polymorphisms and epigenetic marks transmitted to offspring by the father can directly influence offspring fertility and have an effect on the development of their own offspring. Efforts are under way to identify genes in bulls with extreme high v. low fertility that bear on sperm phenotype and sperm production traits (Taylor et al., Reference Swann2018). There are already some examples of gene polymorphisms that affect bull sperm phenotype. Genes encoding for sperm head proteins, integrin β 5 (ITGB5) and collagen type I α 2 (COL1A1), have been associated with bull fertility (Feugang et al., Reference Feugang, Kaya, Page, Chen, Mehta, Hirani, Nazareth, Topper, Gibbs and Memili2009). Other polymorphisms associated with bull sperm phenotypes include genes encoding for sperm surface CD9, which may influence sperm motility (Kumar et al., Reference Kumar, Singh, Deb, Tyagi, Mandal, Kumar, Sengar, Sharma and Singh2015) and TEX11 (testis expressed 11) associated with scrotal circumference (de Camargo et al., Reference de Camargo, Porto-Neto, Kelly, Bunch, McWilliam, Tonhati, Lehnert, Fortes and Moore2015). Holstein bull fertility may be influenced by single-nucleotide polymorphisms in the miR target sites of gene encoding spermatid transitional protein 1 (TNP1) (Zhang et al., Reference Yue, Chang, DeJarnette, Marshall, Lei and Liu2015) and by polymorphism-driven alternative splicing of the sperm flagella 2 (SPEF2) gene (Guo et al., Reference Guo, Yang, Ju, Wang, Qi, Zhang, Wang, Liu, Feng, Chen, Xu, Zhong and Huang2013). The copy number in spermatogenesis associated multi-copy genes such as preferentially expressed antigen in melanoma may also result in varied bull fertility (Yue et al., Reference Yonezawa2013). More attention should be directed to epigenetic marks on sperm-borne genome and chromatin that are generally thought to be erased in the zygote and re-established in the early embryo. Both sperm histone H3 modifications (Kutchy et al., Reference Kutchy, Menezes, Chiappetta, Tan, Wills, Kaya, Topper, Moura, Perkins and Memili2017) and sperm DNA methylation (Kropp et al., Reference Kropp, Carrillo, Namous, Daniels, Salih, Song and Khatib2017) have been associated with bull fertility. Finally, genes that are transcribed from paternal allele during early development and may support developmental competence and pregnancy establishment, or guard against pregnancy loss will be examined for potentially detrimental polymorphisms (Cochran et al., Reference Cochran, Cole, Null and Hansen2013). Ongoing genomic studies will likely identify a number of gene mutations that may contribute to fertility or convey subfertility to bulls used in AI service.
Summary
Gamete interactions during bovine fertilization represent an intricate mesh of remodeling, recycling and signaling that results in transformation of gamete-specific structures into zygotic components. Thus, the sperm acrosome is remodeled during capacitation causing the exposure of zona-binding ligands on the OAM, and is partially lost to acrosomal exocytosis induced by the oocyte ZP. The sperm head PT dissolves in ooplasm to release oocyte-activating factors, and the denuded sperm nucleus is turned into a zygotic paternal pronucleus. The reduced sperm centrosome contributes the formation of zygotic centrosomes and initiates nucleation of microtubule sperm aster necessary for maternal and paternal pronucleus apposition. Various sperm-borne RNAs may regulate early embryo development and even impose epigenetic marks. Sperm contributed paternal genome affects developmental competence of embryos and ultimately imposes heritable traits on offspring.
While some sperm accessory structures are simply recycled after fertilization as they become obsolete, the fate of sperm mitochondria is of particular interest as these have to be eliminated to promote clonal, maternal inheritance of mitochondrial genes and avoid heteroplasmy. Similarly, the regulatory influence of sperm-borne RNAs on zygotic and embryonic development warrants further investigation. Altogether, the ability of some sperm contributed structures and molecules to develop into the functional zygotic components and properly induce embryonic development may vary between bulls, bearing on their reproductive performance, and on the fitness, health, fertility and production traits of their offspring.
Acknowledgments
The author wishes to thank Kathryn Craighead for editorial and clerical assistance. Work in the author’s laboratory pertinent to this review article is currently funded by Agriculture and Food Research Initiative Competitive Grant No. 2015-67015-23231 from the USDA National Institute of Food and Agriculture, Grant No. 5 R01 HD084353-02 from NIH National Institute of Child and Human Development, and by seed funding from the Food for The 21st Century Program of the University of Missouri.
Declaration of interest
There is no conflicts of interest to declare.
Ethics statement
This is a review article; consequently, there are no applicable ethics/compliance committee approvals to be listed.
Software and data repository resources
There are no deposited data or models associated with this article.







