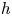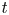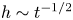95 results
Swelling and Texture of Iron-Bearing Smectites Reduced by Bacteria
-
- Journal:
- Clays and Clay Minerals / Volume 46 / Issue 5 / October 1998
- Published online by Cambridge University Press:
- 28 February 2024, pp. 487-497
-
- Article
- Export citation
Lubrication dynamics of a settling plate
-
- Journal:
- Journal of Fluid Mechanics / Volume 977 / 25 December 2023
- Published online by Cambridge University Press:
- 18 December 2023, A28
-
- Article
-
- You have access
- Open access
- HTML
- Export citation
Debate 30B - Molecular Profiling Should be Done to Guide the Management of Endometrial Cancer?
- from Section IV - Endometrial Cancer
-
-
- Book:
- 50 Big Debates in Gynecologic Oncology
- Published online:
- 20 July 2023
- Print publication:
- 03 August 2023, pp 182-185
-
- Chapter
- Export citation

The Cambridge Handbook on the Material Constitution
-
- Published online:
- 15 January 2023
- Print publication:
- 05 January 2023
1 - The Tradition of the Material Constitution in Western Marxism
- from Part I - History
-
-
- Book:
- The Cambridge Handbook on the Material Constitution
- Published online:
- 15 January 2023
- Print publication:
- 05 January 2023, pp 25-44
-
- Chapter
- Export citation
Part II - Challenges
-
- Book:
- The Cambridge Handbook on the Material Constitution
- Published online:
- 15 January 2023
- Print publication:
- 05 January 2023, pp 169-258
-
- Chapter
- Export citation
Part III - Analyses
-
- Book:
- The Cambridge Handbook on the Material Constitution
- Published online:
- 15 January 2023
- Print publication:
- 05 January 2023, pp 259-366
-
- Chapter
- Export citation
Contents
-
- Book:
- The Cambridge Handbook on the Material Constitution
- Published online:
- 15 January 2023
- Print publication:
- 05 January 2023, pp v-viii
-
- Chapter
- Export citation
Contributors
-
- Book:
- The Cambridge Handbook on the Material Constitution
- Published online:
- 15 January 2023
- Print publication:
- 05 January 2023, pp ix-xii
-
- Chapter
- Export citation
Introduction
-
-
- Book:
- The Cambridge Handbook on the Material Constitution
- Published online:
- 15 January 2023
- Print publication:
- 05 January 2023, pp 1-22
-
- Chapter
- Export citation
Index
-
- Book:
- The Cambridge Handbook on the Material Constitution
- Published online:
- 15 January 2023
- Print publication:
- 05 January 2023, pp 367-382
-
- Chapter
- Export citation
Copyright page
-
- Book:
- The Cambridge Handbook on the Material Constitution
- Published online:
- 15 January 2023
- Print publication:
- 05 January 2023, pp iv-iv
-
- Chapter
- Export citation
Part I - History
-
- Book:
- The Cambridge Handbook on the Material Constitution
- Published online:
- 15 January 2023
- Print publication:
- 05 January 2023, pp 23-168
-
- Chapter
- Export citation
Transatlantic Charismatic Renewal, c. 1950–2000. Edited by Andrew Atherstone, Mark P. Hutchinson, and John Maiden. Global Pentecostal and Charismatic Studies 41. Leiden, Netherlands: Brill, 2021. viii + 260 pp. $63.00 paper.
-
- Journal:
- Church History / Volume 91 / Issue 4 / December 2022
- Published online by Cambridge University Press:
- 03 May 2023, pp. 966-968
- Print publication:
- December 2022
-
- Article
- Export citation
Authoritarian liberalism and the transformation of modern europe: Rejoinder
-
- Journal:
- European Law Open / Volume 1 / Issue 1 / March 2022
- Published online by Cambridge University Press:
- 06 April 2022, pp. 191-208
-
- Article
-
- You have access
- Open access
- HTML
- Export citation
Real-world monitoring progress towards the elimination of hepatitis C virus in Australia using sentinel surveillance of primary care clinics; an ecological study of hepatitis C virus antibody tests from 2009 to 2019 – CORRIGENDUM
-
- Journal:
- Epidemiology & Infection / Volume 150 / 2022
- Published online by Cambridge University Press:
- 04 March 2022, e44
-
- Article
-
- You have access
- Open access
- HTML
- Export citation
Real-world monitoring progress towards the elimination of hepatitis C virus in Australia using sentinel surveillance of primary care clinics; an ecological study of hepatitis C virus antibody tests from 2009 to 2019
-
- Journal:
- Epidemiology & Infection / Volume 150 / 2022
- Published online by Cambridge University Press:
- 06 December 2021, e7
-
- Article
-
- You have access
- Open access
- HTML
- Export citation
A comprehensive evaluation of the quality and complexity of prostate IMRT and VMAT plans generated by an automated inverse planning tool
-
- Journal:
- Journal of Radiotherapy in Practice / Volume 21 / Issue 4 / December 2022
- Published online by Cambridge University Press:
- 27 April 2021, pp. 506-512
-
- Article
- Export citation
Beyond the Post-Sovereign State?: The Past, Present, and Future of Constitutional Pluralism
-
- Journal:
- Cambridge Yearbook of European Legal Studies / Volume 21 / December 2019
- Published online by Cambridge University Press:
- 18 September 2019, pp. 6-23
- Print publication:
- December 2019
-
- Article
- Export citation
5 - Authoritarian Liberalism
- from Part II - The Crisis as a Crisis of the EU’s Political and Democratic Legitimacy
-
-
- Book:
- The Crisis behind the Eurocrisis
- Published online:
- 07 July 2019
- Print publication:
- 25 July 2019, pp 101-121
-
- Chapter
- Export citation