Introduction
Rabbits play an important role in people’s lives. Rabbit meat is of high nutritional value as it is rich in protein, lecithin and lysine, and has low levels of uric acid, calories, fat and cholesterol; it is also easy to digest and absorb (Legendre et al., Reference Legendre, Hoste and Gidenne2017; Mancini et al., Reference Mancini, Preziuso, Fratini, Torracca, Nuvoloni, Dal Bosco and Paci2017). Furthermore, rabbit hair is an important raw material for textiles (Rafat et al., Reference Rafat, de Rochambeau, Brims, Thebault, Deretz, Bonnet and Allain2007). More importantly, however, rabbits are an extremely valuable animal model for studying human disease (Esteves et al., Reference Esteves, Abrantes, Baldauf, BenMohamed, Chen and Christensen2018). Rabbits are widely used in research on hyperlipidemia and atherosclerosis, as well as in fields such as ophthalmology and orthopaedics (Fan et al., Reference Fan, Kitajima, Watanabe, Xu, Zhang, Liu and Chen2015). For example, Watanabe heritable hyperlipidaemic (WHHL) and myocardial infarction-prone WHHL (WHHLMI) rabbits that have been developed through the selection of natural mutations were fed cholesterol to establish a human disease model of atherosclerosis, leading to the development of many effective drugs for curing lipid metabolism disorders and atherosclerosis (Shiomi et al., Reference Shiomi, Koike and Ito2013).
Genome-edited animals are also very important for studying gene functions and pathogenesis (Song et al., Reference Song, Yang, Xu, Zhu, Chen and Zhang2016). Several rabbit genes have been successfully modified using genome editing tools. Somatic cell nuclear transfer (SCNT) is an important technique for producing genome-edited animals; it also has great value in saving endangered species and in clone stem cell therapy, and the rabbit is an important model for studying SCNT (Liu et al., Reference Liu, Wang, Lu, Miao, Cao, Zhang, Wu, Wu, Ding, Wang, Luo, Li and Tan2016; Matoba & Zhang, Reference Matoba and Zhang2018). However, SCNT has a low efficiency (Sugimoto et al., Reference Sugimoto, Kida, Oh, Kitada, Matsumoto, Saeki, Taniguchi and Hosoi2015). Many factors, such as the type and cell cycle of the donor cell, the oocyte status, the fusion method, and the culture medium, affect the success of cloning (Matoba & Zhang, Reference Matoba and Zhang2018). However, abnormal reprogramming is the main reason for the low efficiency of SCNT embryos (Chen et al., Reference Chen, Du, Xu, Chang, Liu, Su, Lin, Ju, Cheng, Wu, Chen and Sung2013). Consequently, as the proteins, RNAs, and other factors in oocytes play a pivotal role in reprogramming and development, the selection of suitable oocytes is crucial to the success of SCNT (Sugimoto et al., Reference Sugimoto, Kida, Oh, Kitada, Matsumoto, Saeki, Taniguchi and Hosoi2015).
The aim of the present study was to clarify whether brilliant cresyl blue (BCB) staining could improve the efficiency of rabbit SCNT. To do this, we collected rabbit ovaries from slaughterhouses, stained the oocytes with BCB, and then matured them in vitro.
Materials and methods
Materials and ethics statement
All chemicals and reagents were obtained from Sigma-Aldrich (MO, USA) unless specifically stated otherwise. Disposable, sterile plasticware was obtained from Corning Incorporated (Corning, NY, USA). The Animal Care and Use Committee of Xi’an Jiaotong University approved all animal use procedures applied in this study.
Collection of ovaries and oocytes
Ovaries were collected from female rabbits in a slaughterhouse within 4 h of being slaughtered. Any excess mesangial cells and fat was cut away from around the ovary and the fallopian tubes and blood stains were removed. The ovaries were then placed in saline containing penicillin and streptomycin and were delivered to the laboratory within 2 h. The ovaries were washed with phosphate-buffered saline (PBS) and the ovarian follicles were aspirated using a 12-gauge needle attached to a 10-ml syringe. The follicular fluid was then poured into a 6-cm dish and cumulus–oocyte complexes (COCs) were picked out under a stereoscope microscope and placed in Dulbecco’s phosphate-buffered saline (DPBS).
BCB staining and in vitro maturation of oocytes
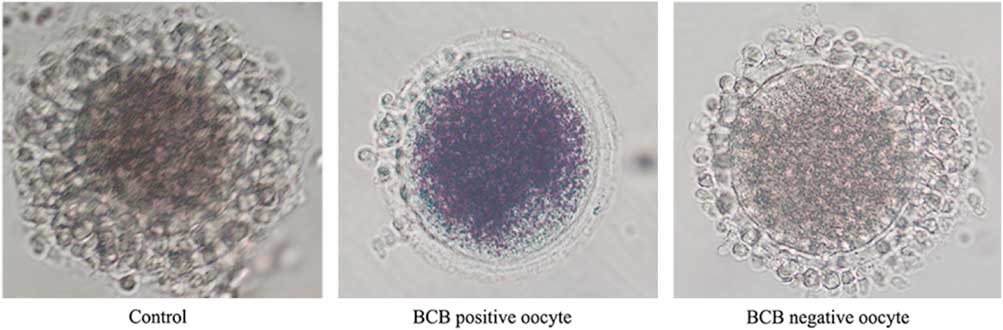
After three washes in DPBS containing 5% (v/v) fetal bovine serum (FBS), the COCs were treated with 26 mM BCB for 90 min in 5% CO2 at 38.5°C, and then washed three times for 5 min in DPBS containing 5% (v/v) FBS. COCs with a blue cytoplasm were separated out into the BCB-positive group, while those with a colourless cytoplasm were placed in the BCB-negative group (Fig. 1). In addition, another set of COCs was cultured without BCB treatment and used as the control group.
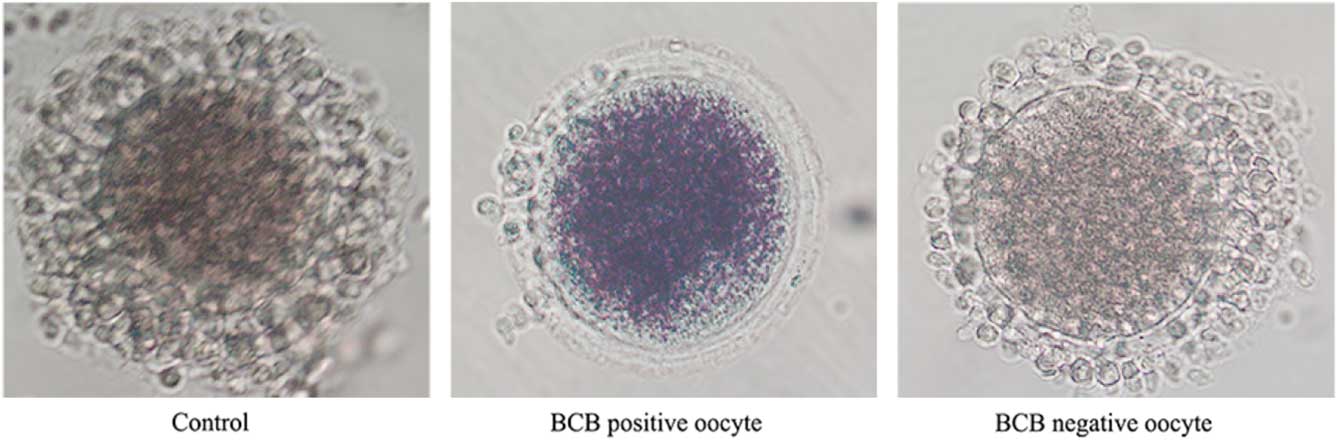
Figure 1 Representative photographs of rabbit COCs after BCB staining. The BCB-positive oocyte stain shows a blue cytoplasm, while oocytes of the BCB-negative and the control groups show a colourless cytoplasm.
The COCs were washed three times for 6 min with DPBS containing 5% (v/v) FBS and then transferred into the oocyte maturation medium, which consisted of tissue culture medium-199 (TCM-199; GIBCO, NY, USA) supplemented with 10% (vol/vol) FBS (FCS; GIBCO, NY, USA), 1 μg/ml 17β-estradiol, 24.2 mg/l sodium pyruvate, 0.05 IU/ml follicle-stimulating hormone (FSH), 0.05 IU/ml luteinizing hormone, and 10 ng/ml epidermal growth factor. The COCs were cultured at 38.5°C with 5% CO2 for 16 h, were then treated with DPBS containing 0.1% hyaluronidase for 5 min and were gently dried for 1 min. Oocytes with polar bodies were considered to be mature and were used for SCNT.
Preparation of donor cells
Fibroblasts were obtained from the ear of a 14-day fetus of New Zealand white rabbit. A small piece of skin (1×1 cm2) was shaved and then washed three times with modified DPBS containing penicillin and streptomycin. The skin was then cut into smaller pieces and cultured for approximately 10 days in Dulbecco’s modified Eagle medium (DMEM) (GIBCO, NY, USA) containing 10% FBS, penicillin, and streptomycin. The fibroblasts were cultured in saturated humidity with 5% CO2 at 38.5°C. When cell confluence reached 80%, the fibroblasts were transferred into a 3.5-cm dish with fresh culture medium and were purified through two passages. Fibroblasts from passages 3 and 4 were then cultured to confluence in 96-well plates and incubated in DMEM supplemented with 0.5% FBS for 3 days. The cells were digested with DPBS containing 0.25% (w/v) trypsin and 1 mM ethylenediaminetetraacetic acid (EDTA) for 1 min, after which digestion was terminated by adding DMEM containing 10% FBS. The sample was then centrifuged at 350 g for 10 min to obtain a cell pellet. The pellet was diluted in DMEM containing 7.5 mg/ml cytochalasin B and 10% FBS for SCNT.
Somatic cell nuclear transfer
Oocytes were transferred into the micromanipulation medium, which was composed of M199 containing 7.5 μg/ml cytochalasin B. Each oocyte was held with a micropipette and rotated using another micropipette to orientate the polar body in the 2 o’clock direction. The micropipette was then inserted in the 3 o’clock direction and the polar body and surrounding cytoplasm were removed. A donor cell was injected into the perivitelline space of the enucleated oocyte, and the donor cell and enucleated oocyte complex (DOC) were transferred into fusion medium, which consisted of 0.25 M sorbitol in water supplemented with 0.5 mM HEPES, 0.1 mM Ca(CH3COO)2, 0.5 mM Mg(CH3COO)2, and 1 mg/ml bovine serum albumin. The DOC was fused in the fusion medium by applying three DC pulses of 3.2 kV/cm for 20 μs each. Once fused, the DOC was washed three times for 6 min and allowed to recover for 1 h at 38.5°C and 5% CO2.
Embryo activation and culture
The fused embryos were activated by electrical pulses, as described above, and then washed three times with DPBS for 3 min. They were then treated with 2 mM 6-dimethylaminopurine (6-DMAP) and 5 μg/ml cycloheximide (CHX) in Earle’s balanced salt solution (EBSS)-complete medium for 3 h, after which they were washed three times with DPBS for 3 min. The cloned embryos were subsequently cultured in EBSS-complete medium in saturated humidity with 5% CO2 at 38.5°C.
Nuclear staining
Nuclear staining was performed as described previously (Qu et al., Reference Qu, Zhao, Wang, Zhang, Li, Fan and Liu2018). The embryos were washed three times in 0.1% PBS/polyvinyl alcohol for 5 min per wash, after which 4′,6-diamidino-2-phenylindole hydrochloride (DAPI; Vysis Inc., Downers Grove, USA) was applied for 3 min. The stained embryos were then washed and mounted on slides. The slides were examined using epifluorescence microscopy and a Nikon Eclipse Ti-S microscope (Nikon, Tokyo, Japan). All images were captured with a Nikon DS-Ri1 digital camera and saved in .TIFF format. The nuclei were identified by their blue fluorescence.
Apoptosis analysis
Apoptosis analysis of blastocysts was performed using the dead end fluorometric terminal deoxynucleotidyl transferase dUTP nick end labelling (TUNEL) kit. Embryos were washed with 0.2% PVA–PBS and then fixed with 4% paraformaldehyde at 37°C for 30 min. Unless otherwise stated, the following steps were carried out at room temperature. Embryos were washed, then permeabilized for 20 min with 0.2% Triton X-100. After washing, the embryos were equilibrated in E-buffer for 5–8 min. The embryos were then transferred to the apoptosis dye solution at 37°C in the dark for 1 h. The 51-μl dye solution contained 45 μl E-buffer, 5 μl Nucleotide Mix and 1 μl rTDT. After this step, all subsequent steps were performed in the dark. SSC (2×) was used to terminate the reaction for 15 min. Embryos were washed and stained with DAPI for 3 min. Finally, the film was sealed with immunofluorescence staining and photographed. It was then mounted on slides and examined by epifluorescence microscopy using a Nikon Eclipse Ti-S microscope (Nikon, Tokyo, Japan). All images were captured with a Nikon DS-Ri1 digital camera and saved in .TIFF format.
Immunofluorescence staining
Embryos were washed three times with PBS containing 0.2% PVA, and then fixed with 4% paraformaldehyde at 37°C for 30 min. They were then washed three times for 5 min, and permeabilized with 0.2% Triton X-100 for 30 min. Next, they were treated with sealing solution for 2 h at room temperature, incubated with anti-CDX2 antibody overnight at 4°C, washed three times for 5 min and incubated with secondary antibody labelled with Alexa Fluor 488 for 2 h at room temperature. Then, the embryos were incubated with DAPI for 3 min, washed three times for 5 min, mounted on slides and examined by epifluorescence microscopy using a Nikon Eclipse Ti-S microscope (Nikon, Tokyo, Japan). All images were captured with a Nikon DS-Ri1 digital camera and saved in .TIFF format. CDX2, a specific marker of trophoblast cells (TE) in the blastocyst, was used to define trophoblast cells by immunofluorescence staining with anti-CDX2 antibody (An et al., Reference An, Peng, Cheng, Lu, Zhou, Zhang and Su2019). The inner cell mass (ICM) was negative for CDX2, and all nuclei in the blastocyst were stained by DAPI.
Statistical analysis
Statistical analysis was performed as described previously (Qu et al., Reference Qu, Zhao, Wang, Zhang, Li, Fan and Liu2018). Each experiment was repeated at least three times. The maturation rate was the number of oocytes with polar bodies divided by the total number of oocytes with or without polar bodies (Fig. 2). Cytokinesis proceeds during cleavage of the SCNT embryo, and the SCNT blastocyst has a cavity structure (Fig. 3). The cleavage rate was the number of cleavage embryos divided by the total number of fused embryos, while the blastocyst rate was the number of blastocysts divided by the total number of fused embryos. Differences in the maturation rate, fusion rate, cleavage rate, and blastocyst rate among treatment groups were analyzed using the χ2 test. The total cell number, ICM/TE staining, and apoptosis staining were determined by randomly selecting blastocysts from each treatment group, using approximately 20 embryos per replicate. The differences among treatment groups were analyzed by one-way analysis of variance (ANOVA). All statistical analyses were undertaken using SPSS software with a significance level of P<0.05. Data were expressed as the mean±standard error of the mean (SEM).
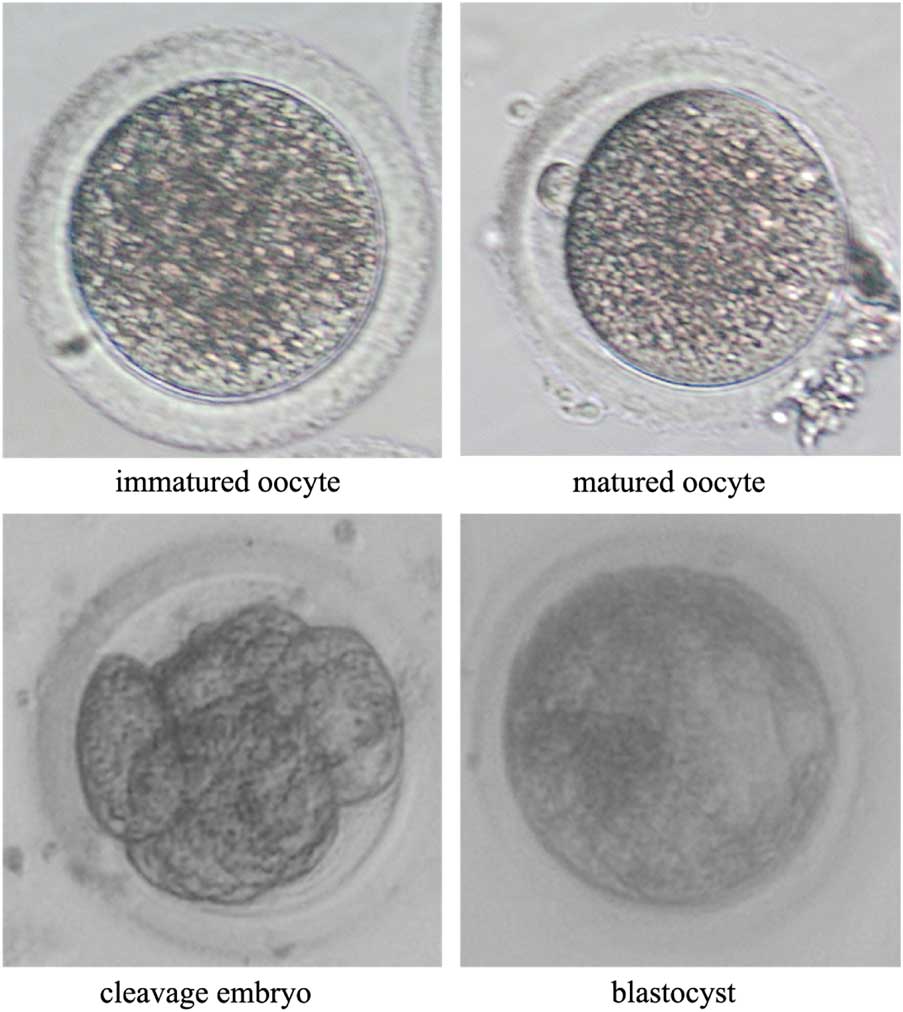
Figure 2 Representative photographs of rabbit oocytes after in vitro maturation and of embryos after in vitro culture. The immature oocyte has no polar body, while the mature oocyte has a polar body in the perovenal space. Cytokinesis proceeds during cleavage of the SCNT embryo, and the SCNT blastocyst has a cavity structure.

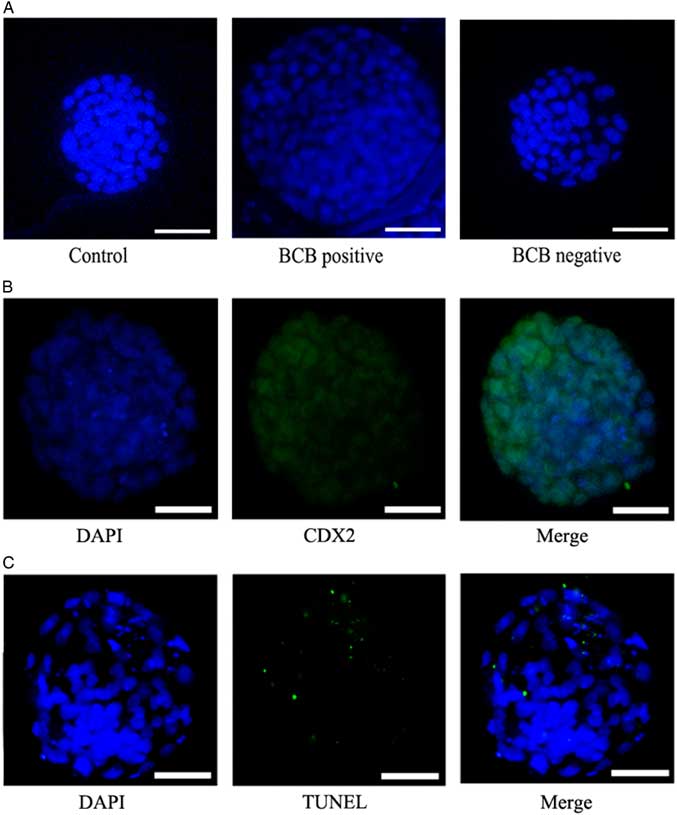
Figure 3 Representative photographs of the total number of blastocysts in each group. (A) SCNT blastocysts developed from the control group, BCB-positive group, and BCB-negative group were stained with DAPI. Bars represent 40 μm. (B) Trophectoderm (TE) was detected by Cdx2 antibody, a special marker of TE (green), DNA was stained by DAPI (blue) to visualize all blastomeres. Bars represent 40 μm. (C) The apoptotic blastomeres were detected by TUNEL (green). DNA was stained by DAPI (blue) to visualize all blastomeres. Bars represent 40 μm.
Results
BCB-positive oocytes have higher rates of in vitro maturation
COCs in the control group, which were not stained with BCB, had a maturation rate of 65.3% in vitro. By contrast, COCs in the BCB-negative group had a maturation rate of 48.9%, which was significantly lower than the control group (P<0.05), while those in the BCB-positive group had a maturation rate of 81.4%, which was significantly higher than the BCB-negative group and control group (P<0.05) (Table 1). These results indicated that BCB staining is a reliable method for selecting oocytes with a high maturation ability.
Table 1 Effect of oocyte selection by BCB staining on the maturation of rabbit oocytes in vitro

a,b,c Different superscripts within same column indicate significant difference (P<0.05).
BCB-positive oocytes have a higher developmental competence in rabbit cloned embryos in vitro
Matured oocytes from the three groups were used for SCNT. There was no significant difference in fusion rate between the three groups (P>0.05). However, the BCB-positive group had a significantly higher cleavage rate (86.6%) and blastocyst rate (30.5%) than the BCB-negative group (P<0.05). Furthermore, the total number of blastocysts was also significantly higher in the BCB-positive group (90.0±7.5) than in the BCB-negative group (65.3±6.3) and control group (67.5±5.7) (P<0.05) (Table 2). The ICM/TE index in the BCB-positive group (42.3±4.2) was significantly higher than that of the BCB-negative group (30.2±2.1) and the control group (33.9±5.1) (P<0.05). The apoptosis index in the BCB-positive group (2.1±0.6) was significantly lower than that of the BCB-negative group (8.2±0.9) and the control group (6.7±1.1) (P<0.05). These results indicated that BCB-positive oocytes had a higher developmental competence in rabbit SCNT embryos in vitro and that BCB staining is a reliable method for selecting oocytes to enhance the efficiency of SCNT.
Table 2 Effect of oocyte selection by BCB staining on the developmental competence of rabbit cloned embryos in vitro

a,b,c Different superscripts within same column indicate significant difference (P<0.05).
Discussion
Gene knock-in and knock-out experiments are very useful for studying gene function and pathogenesis. Several rabbit genes have been successfully modified via gene editing tools such as zinc finger nuclease, transcription activator-like effector nuclease, clustered regularly interspaced short palindromic repeat (CRISPR) and CRISPR-associated protein 9 (CRISPR/Cas9) (Honda & Ogura, Reference Honda and Ogura2017; Liu et al., Reference Liu, Sui, Liu, Liu, Chen, Deng, Xu and Li2018). Cytoplasmic injection, pronuclear injection, and SCNT are important steps in the production of gene-modified animals (Honda & Ogura, Reference Honda and Ogura2017). SCNT involves the injection of a somatic cell into a denucleated oocyte, followed by activation, culturing, and transfer to the oviduct or uterus, where the reconstructed embryo can develop into a new individual. SCNT allows gene editing to be performed in donor cells and allows the accuracy and safety of gene modification to be assessed before micromanipulation of the embryo and embryo transfer (Yin et al., Reference Yin, Jiang, Fang, Kong, Xing, Li, Chen and Li2015). Consequently, SCNT is the main method that is used for the production of transgenic cattle, sheep, and pigs (Holm et al., Reference Holm, Alstrup and Luo2016; Tan et al., Reference Tan, Proudfoot, Lillico and Whitelaw2016). However, this technique still has a very low success rate, which limits its application (Qu et al., Reference Qu, Qing, Liu, Qin, Wang, Qiao, Ge, Liu, Zhang, Cui and Wang2017). The live fetus birth rate in rabbits is no more than 5% (Meng et al., Reference Meng, Polgar, Liu and Dinnyes2009). Therefore, the optimization of SCNT is extremely important.
The oocyte is the most important factor that affects the success of SCNT (Gonzalez-Munoz & Cibelli, Reference Gonzalez-Munoz and Cibelli2018). The low efficiency of SCNT is mainly caused by the donor nucleus not being completely reprogrammed by the cytoplasm of the recipient oocyte (Caixeta et al., Reference Caixeta, Sousa, Guimaraes, Leme, Spricigo, Netto, Pivato and Dode2017). The reprogramming ability of oocytes is influenced by several factors, such as the age of the female and the drug treatment used (Cervera & Garcia-Ximenez, Reference Cervera and Garcia-Ximenez2003). As the use of oocytes with a low reprogramming ability will result in failure, the denucleation, fusion, in vitro culture, and embryo transfer steps in this very long procedure become meaningless if high-quality oocytes are not used as nuclear receptors in the first step (Bai et al., Reference Bai, Song, Zhang, Huang, Huang, Sun and Lei2018; Li et al., Reference Li, Guo, Zhu, Jin, Zhang, Zhang, Xing, Xuan, Luo, Luo, Wang, Cui, Li, Cui, Yin and Kang2017).
Oocytes are mainly obtained using in vivo or in vitro methods (Sugimoto et al., Reference Sugimoto, Kida, Oh, Kitada, Matsumoto, Saeki, Taniguchi and Hosoi2015; Yin et al., Reference Yin, Jiang, Fang, Kong, Xing, Li, Chen and Li2015). The in vivo method involves female rabbits being treated with a hormone and the oocytes being allowed to mature in vivo. Once the oocytes have matured, the rabbits are euthanized and the oocytes are collected from the oviducts or ovaries. The in vitro method involves ovaries being collected from dead rabbits (primarily from slaughterhouses) and immature oocytes being collected from the ovaries (Liu et al., Reference Liu, Wang, Lu, Miao, Cao, Zhang, Wu, Wu, Ding, Wang, Luo, Li and Tan2016). This method allows large numbers of ovaries and oocytes to be obtained at low cost in a convenient way and has animal welfare benefits.
Oocyte quality has recently been assessed using morphological methods, including assessing the number and density of cumulus cells encapsulated outside the oocyte, and the cytoplasmic colour and uniformity of the oocyte (Wang & Sun, Reference Wang and Sun2007). However, it is difficult to distinguish the developmental potential of oocytes using these ambiguous criteria. Therefore, in the present study, we tested the use of BCB staining and found that this was a reliable method for selecting rabbit oocytes with a high maturation ability.
BCB staining has previously been used to screen the oocytes of pigs, sheep, mice, dogs, and cattle (Kumar et al., Reference Kumar, Faraji, Sarwalia, Kumar, Gohain, De, Kumar and Datta2018; Salviano et al., Reference Salviano, Collares, Becker, Rodrigues and Rodrigues2016; Su et al., Reference Su, Wang, Li, Peng, Hua, Li, Quan, Guo and Zhang2012). However, there has been no report on its application in rabbits. Glucose-6-phosphate dehydrogenase (G6PDH) has a high activity in growing oocytes, resulting in a low developmental potential (Pereira et al., Reference Pereira, Lorenzo, Carneiro, Bilodeau-Goeseels, Kastelic, Esteller-Vico, Lopez-Bejar and Liu2014). The activity of G6PDH decreases once oocytes have finished growing, resulting in a high developmental potential (Alm et al., Reference Alm, Torner, Lohrke, Viergutz, Ghoneim and Kanitz2005). G6PDH with high activity turns BCB from blue to colourless. However, oocytes with high developmental potential and thus lower activity of G6PDH retain the blue colour. Therefore, BCB staining is an effective method for selecting high-quality bovine oocytes for SCNT. Bovine embryos derived from BCB-positive oocytes had a low apoptosis index and exhibited histone modifications and gene expressions that better facilitated reprogramming compared with those derived from BCB-negative oocytes (Su et al., Reference Su, Wang, Li, Peng, Hua, Li, Quan, Guo and Zhang2012). In the present study, we found that BCB-positive oocytes had higher cleavage and blastocyst rates than BCB-negative oocytes. Furthermore, the BCB-positive oocytes also had a higher total number of blastocysts, higher ICM/TE index and a lower apoptosis index than the BCB-negative and control groups, which is another important index of the developmental competence of the embryo.
In conclusion, our results demonstrate for the first time that BCB staining can be used to select rabbit oocytes with a high maturation ability. Furthermore, BCB-positive oocytes support a higher developmental competence of rabbit SCNT embryos in vitro. These findings indicate that BCB staining is a reliable method for selecting oocytes with high efficiency for SCNT.
Conflict of interest
The authors declare no competing or financial interests.
Financial support
This work was supported by the Natural Science Foundation of Shaanxi Province under grant no. 2014FWPT017; Wuzhong Innovation and Entrepreneurship Talent Project of Suzhou under grant no. WC201526; and the National Natural Science Foundation of China under grant no. 81270348.
Ethical standards
Not applicable.