Clostridium difficile infection (CDI) is an enteric disease that is mainly hospital-associated, but also increasing in the community [Reference Hensgens1, Reference Lessa2]. In the Netherlands, in about one-quarter of all diagnosed patients CDI is acquired in the community [Reference Hensgens3]. Over the past decade, a new strain, C. difficile (CD) ribotype 078, has emerged in patients with CDI especially acquired outside the hospital, but has also been found in high numbers in piglets, veal calves, and their immediate environment [Reference Hensgens3, Reference Keel4]. To explain the emergence of community-acquired CDI, the following transmission routes have been suggested: environment-to-person, contact with infected or colonized individuals or animals, and foodborne transmission [Reference Otten5]. A previous study demonstrated that 25% (12/48) of farmers who had daily contact with pigs tested positive for CD, and ribotype 078 was found on 15 of 32 pig farms in both humans and pigs suggesting animal-to-human transmission [Reference Keessen, Harmanus and Dohmen6]. Highest rates were found in pig-breeding farms in piglets, 1–7 days old [Reference Keessen, Harmanus and Dohmen6]. Data on the prevalence of CD in the Dutch general population (i.e. persons not living or working on farms), however, are scarce. In the Netherlands, the locations of pig farms and the area with high occurrence of human CD ribotype 078 infections overlap and CD can be detected in the immediate environment of farms [Reference Goorhuis7], but a direct association between exposure to pigs and high occurrence has not been observed. Our study objective was to assess the prevalence of and risk factors for CD colonization in persons not living or working on a farm in areas with high livestock densities.
This cross-sectional study was part of a larger population-based study on health of residents living in a highly populated rural area with a high density of livestock farms in the Netherlands: the Livestock Farming and Neighbouring Residents’ Health study (Dutch acronym: VGO). The methodology of the VGO study is described in detail by Borlée et al. [Reference Borlée8]. In 2012, a questionnaire survey was conducted among the general population recruited via general practitioners [Reference Borlée9]. Participants were included if they were aged 18–70 years, living in the province of Noord-Brabant or Limburg, and living in municipalities with <30 000 inhabitants. One eligible participant per household was invited. Questionnaire participants who gave consent for contact for a follow-up study, and who were not working or living on livestock farms, were eligible to participate in our cross-sectional population-based study, which was conducted from March 2014 to February 2015. Fecal samples to determine CD colonization (with or without symptoms) were taken once and participants were asked to complete a questionnaire to study risk factors. The distance between participants’ residential address and the nearest farm was calculated [Reference Smit10]. The electronic medical records maintained by general practitioners were used to collect data on comorbidity and exposure to antibiotics [Reference Prins and Verheij11].
CD was cultured by incubating approximately 1 g of feces in 9 ml CDBMN, CD enrichment modified broth (Mediaproducts, 46·1380) with CDMN Selective Supplement (Oxoid, SR0173E), and incubated 10–15 days at 37 °C under anaerobic conditions [Reference Keessen, Harmanus and Dohmen6]. Subcultures were made onto chromID CD agar using direct plating and culture following alcohol shock (Biomérieux, 43871) and suspected isolates were identified as CD by polymerase chain reaction (PCR) to detect glutamate dehydrogenase [Reference Goorhuis7]. Further characterization of CD was performed by determining the presence of toxin genes and by PCR ribotyping [Reference Fawley12, Reference Paltansing13].
Data were analyzed using SPSS version 22.0 (SPSS Inc., Chicago, IL). The Mann–Whitney U-test was used to compare the median distance to farms, the median number of farms within 500 and 1000 m, and the median number of farm animals within 1000 m. Using univariate logistic regression, odds ratios (ORs) with 95% confidence intervals (CIs) were obtained for risk factors. The VGO study was approved by the medical ethical committee of the University Medical Center Utrecht (number 13/533); all participants signed informed consent.
A total of 7180 persons were invited to participate, which resulted in 2494 participants (response rate 34·7%). Of 2494 participants (median age 58 years (range 20–72), 45·3% male), 2432 mailed a fecal sample. The prevalence of CD colonization was 1·23% (30/2432; 95% CI 0·85–1·73). Four CD-positive individuals had a strain belonging to ribotype 078, resulting in a prevalence of 0·16% (95% CI 0·05–0·40). Other toxin producing ribotypes were 014 (n = 7), 002 (n = 2), 265 (n = 2), 001 (n = 1), 005 (n = 1), 012 (n = 1), 013 (n = 1), 017 (n = 1), and 024 (n = 1). Of 30 CD isolates, nine (30%) did not have toxin genes and belonged to PCR ribotypes 039 (n = 4), 010 (n = 1), 085 (n = 1), 071 (n = 1), and an unknown ribotype (n = 2). Of 30 positive samples, 28 were positive both with and without alcohol shock, while two samples were only positive by direct plating without alcohol shock.
Descriptive analyses of the questionnaire data demonstrated that 45% of CD positives had gastroenteritis in the month preceding the study, compared with 20% of negatives (P < 0·01). When including only the toxin-producing CD positives, this difference was respectively 50% vs. 20% (P < 0·01). The frequency of nausea, diarrhea and fever was not significantly different between CD positives compared with negatives (14 vs. 9%, 24 vs. 17%, and 10 vs. 4%, respectively).
The results of risk factor analyses are shown in Table 1. There was no association between CD colonization and distance to the nearest farm, the number of farms within 500 and 1000 m, and the number of farms animals within 1000 m (Table 1). However, CD positives were more likely to have used antibiotics 3 and 6 months before the study compared with negatives (3 months: OR 3·70; 95% CI 1·25–10·95) (6 months: OR 2·64; 95% CI 1·05–6·64) (Table 1). In addition, 11·5% of CD positives who used antibiotics 3 months before the study received penicillins compared with 1·9% of CD negatives (OR 6·76; 95% CI 1·95–23·47). Visiting farms was significantly negative associated with CD colonization (OR 0·42; 95% CI 0·20–0·88).
Table 1. Risk factors for Clostridium difficile (CD) among adults living near livestock farms in the Netherlands
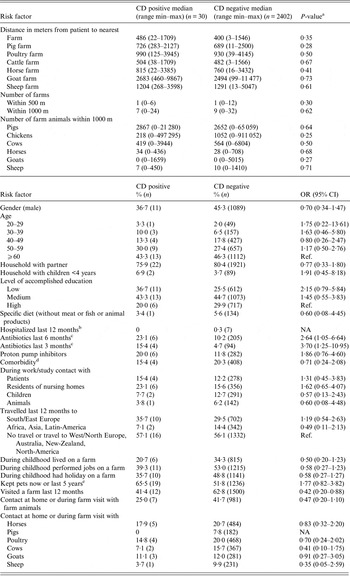
a Mann–Whitney U-test.
b Hospitalized in the Netherlands and/or abroad.
c For 415 persons data on antibiotics and comorbidity was missing.
d Included cerebrovascular disease, chronic cardiovascular disease, liver disease, chronic lung disease, chronic renal disease, auto-immune disease, neurological comorbidity, diabetes, and malignancy.
e Dog, cat, bird, rabbit, guinea pig, hamster, mouse, rat, fish, turtle.
To our knowledge, this is the first study on the prevalence of CD colonization among persons not living or working on farms in areas with a high density of livestock farms. There is limited information on the prevalence of CD in general healthy populations, since most of the studies focused on patients in healthcare facilities [Reference Furuya-Kanamori14]. The prevalence of 1·2% in our study is considerably lower than the 4·7% that was reported among patients at time of admission to three different hospitals in the Netherlands [Reference Terveer15]. However, the investigated groups differed considerable with higher age and more comorbidity in the latter study. Including persons living or working on farms, would probably also have resulted in a higher CD prevalence.
The most frequently found toxigenic PCR ribotypes in our study were ribotypes 014 (n = 7; 23·3%) and 078 (n = 4; 13·3%). Those PCR ribotypes, together with 020, are also the most frequently isolated types in the Netherlands amongst hospitalized patients with CDI [Reference Crobach16]. Ribotype 078 has been found in pigs and humans who have direct contact with pigs (i.e. pig farmers) [Reference Keessen, Harmanus and Dohmen6]. No association was observed between CD colonization and living near (pig) farms, suggesting that there is no significant transmission from the environment around farms to humans. However, it should be noted that farmers were excluded from this study. A limitation of our study is the lack of information on the presence of CD at the farms in the study area, since no microbiological investigations on the farms and their surroundings were performed. An ecological association between the occurrence of human CD type 078 and the location of pig farms in the Netherlands has been reported [Reference Goorhuis7]. Moreover, a previous study reported CD to be present in both humans and animals in about half of pig farms (15/32) [Reference Keessen, Harmanus and Dohmen6]. However, it is unknown whether there are trends in time and what this prevalence is in farms with other animal species than pigs. In our study, we found a significant negative association between CD and visiting farms, for which we have no explanation.
Antibiotic usage is an established risk factor for CDI [Reference Deshpande17]. We also found an association between CD colonization and exposure to antibiotics, especially penicillins. Penicillins have previously been identified as a risk factor for community-associated CDI [Reference Deshpande17]. We did not find an association between CD colonization and other well-known risk factors, such as previous hospitalization, increasing age, and comorbidity [Reference Keessen, Gaastra and Lipman18]. We also did not find an association between proton pump inhibitors and community-acquired CDI. This could be due to the small number of cases and characteristics of this study population.
In conclusion, the prevalence of CD colonization among adults living near livestock farms was 1·2%. In a livestock dense area, CD positive individuals did not live closer to livestock farms than CD negative individuals, indicating that the effect of exposure through the farm environment is limited. Antibiotic exposure was a risk factor for CD colonization.
ACKNOWLEDGEMENTS
The study was funded by the Ministry of Health, Welfare and Sports and the Ministry of Economic Affairs of the Netherlands, and supported by a grant from the Lung Foundation Netherlands (Grant number 3.2.11.022).




