INTRODUCTION
Community-acquired pneumonia (CAP) causes significant morbidity and mortality worldwide, particularly in children aged <5 years [Reference Zar1, Reference Walker2] and the elderly [Reference Amodio3, Reference Janssens and Krause4]. Streptococcus pneumoniae (Spn) is the leading aetiological agent identified in CAP, causing ~25% of CAP in adults and 8% in young children [Reference Said5–Reference O'Brien7]. However, there is currently no adequate gold standard for aetiological diagnosis of CAP: even with the best clinical and laboratory tools, 30–60% of cases have no confirmed aetiology [Reference Ewig8, Reference Ruiz9].
Numerous tests exist to identify the bacterial aetiology of CAP, but they all suffer limitations. Lung aspirates and pleural fluid cultures are sensitive and specific for bacterial identification, but their collection is invasive and rarely indicated. Sputum and nasopharyngeal swab testing cannot differentiate between pathogens colonizing the upper respiratory tract and those causing lower respiratory tract infections [Reference Cherian10]. Blood cultures have a sensitivity of 0–14% [Reference Song, Eun and Nahm11–Reference Afshar13] because of infrequent bacteraemia [Reference Said5], prior antibiotic use, and inadequate sample volumes, especially in children [Reference Afshar13]. For pneumococcal detection, urine antigen testing is more sensitive than blood culture, but false-positive results arise in populations with high nasopharyngeal carriage such as children [Reference Azzari14, Reference Charkaluk15] or patients with recent pneumococcal infection [Reference Song, Eun and Nahm11]. Polymerase chain reaction (PCR) on blood specimens has been suggested as a potential diagnostic tool with superior sensitivity over blood culture combined with high specificity [Reference Avni16–Reference Toikka20] but its validity remains uncertain. The World Health Organization (WHO) developed radiological criteria to standardize the interpretation of paediatric chest radiography (CXR) in clinical trials of Haemophilus influenzae type b (Hib) vaccine and pneumococcal conjugate vaccine (PCV). For these trials, primary endpoint pneumonia (PEP) was defined as alveolar consolidation or pleural effusion on CXR, as agreed upon by two of three independent readers [Reference Cherian21]. Compared to all severe or hospitalized pneumonia, this WHO-defined outcome is more likely associated with pneumococcus or Hib. For example, a trial in The Gambia found that 9-valent PCV prevented 12% of all severe pneumonia and 37% of PEP [Reference Cutts22]. This approach also has limitations: it has been evaluated primarily in children aged <2 years [23] and its sensitivity and specificity are poorly defined. In sum, the lack of a gold standard to diagnose pneumococcal pneumonia makes the evaluation of new laboratory tools for aetiological confirmation challenging.
The PneumoTone study aimed to gather baseline data on pneumococcal meningitis and pneumonia epidemiology in northern Togo, located in the African meningitis belt, to assess the impact of PCV introduction on Spn disease burden across all age groups. In this sub-study, we used latent class analysis (LCA) to assess the diagnostic value of routine tests such as semi-automated blood culture (SABC), CXR, and serum C-reactive protein (CRP), as well as newer tests such as the lytA real-time PCR (rt-PCR) on whole blood in diagnosing Spn in CAP patients in northern Togo. We then applied these findings to estimate the proportion of CAP attributable to Spn in our study population.
METHODS
Study population
We included all patients residing in Tône or Cinkassé districts, northern Togo, who presented with clinical signs or symptoms of pneumonia, were hospitalized for at least one night at one of the five study sites during 1 May 2010 to 31 October 2013 and provided informed consent to participate.
Togo introduced Hib conjugate vaccine into its routine immunization programme during 2008. Vaccine coverage with three doses of pentavalent vaccine was 81·6% in children aged <24 months at the national level during 2013 [Reference Smith, Diggle and Clarke24].
Data collection
We defined clinical pneumonia, severe pneumonia, PEP, suspected pneumococcal pneumonia, and confirmed pneumococcal pneumonia using PneumoADIP criteria [Reference Deloria Knoll25]. Following medical examination, data were collected on medical history, including HIV status (self-reported), rhinopharyngitis or pneumonia in the last 2 weeks, and symptoms at admission. We defined sepsis based on clinical signs as in Goldstein et al. for children [Reference Goldstein26] and Bone et al. for adults [Reference Bone27]. The following events were considered acute complications: appearance or worsening of cardio-respiratory distress, appearance or worsening of severity criteria, secondary infection, sepsis, organ failure, and death.
Radiological and laboratory tests
All patients had an antero-posterior CXR obtained on admission, which was digitalized and read off-site by a US paediatrician experienced in reading adult and paediatric CXRs as well as a Togolese general radiologist; a second US paediatrician who participated in the original development of the WHO radiological criteria [Reference Cherian21]arbitrated cases where the primary readers had discordant findings. All readers were trained on a standard set of films and used WHO criteria for identifying PEP including presence or absence of alveolar consolidation or pleural effusion and the size of any consolidation. Laboratory tests included two sets of two blood cultures collected 30–90 min apart and inoculated into aerobic and anaerobic BacT/ALERT® bottles (bioMérieux, France) for adults and two blood cultures collected 30–90 min apart cultured in paediatric bottles for children aged <15 years, serum CRP dosage (SECOMAM® spectrophotometer; Secomam, France), and rt-PCR on whole blood to identify Spn (lytA gene), Hib (bexA gene) and Staphylococcus aureus (etvick gene) with a Ct cut-off of 35 for positivity [Reference Pai, Gertz and Beall28]. A subset of patients had a nasopharyngeal aspirate collected and tested using the Fast Track Diagnostics Respiratory 21+ multiplex PCR platform (Fast Track Diagnostics, Luxembourg) for viral and bacterial detection.
Statistical analysis
We used LCA [Reference Formann and Kohlmann29] to construct eight models enabling us to categorize patients into two classes interpreted as likely pneumococcal CAP or non-pneumococcal CAP. The models all included laboratory tests (SABC, serum CRP levels, lytA rt-PCR) and the presence or absence of PEP on CXR; various combinations of clinical criteria were also included in some models. Local dependence between blood culture and lytA rt-PCR was evaluated using a likelihood ratio test comparing models with independent tests to models with tests combined into a single variable. Goodness-of-fit was assessed by parametric bootstrap (2000 simulations). A total of four models with adequate fit were found, and all of them incorporated local dependence between blood culture and lytA rt-PCR. The final models included PEP on CXR, blood culture, lytA rt-PCR and CRP level, the latter categorized in quartiles or dichotomized with a cut-off of 40 mg/l [Reference Cherian10]. Two models also included the occurrence of acute complications. The proportion of CAP attributable to Spn, the sensitivity and specificity of each test, and the receiver operating characteristic (ROC) curve for serum CRP dosage were estimated based on the models’ categorization of likely pneumococcal CAP or non-pneumococcal CAP. The corresponding 95% confidence intervals were assessed by non-parametric bootstrap (2000 simulations). We performed a descriptive analysis of both identified latent classes for the model with good fit that used the smallest number of variables. We also conducted a sensitivity analysis to confirm our a priori hypothesis of two latent classes and not more, and to check the respective weight of each of the laboratory and radiological tests in the categorization of cases by the model. Finally, we performed a subgroup analysis using a similar approach on patients with nasopharyngeal aspirate testing done, including an indicator variable for Spn colonization in the LCA models.
Statistical analyses were performed using Stata v. 12 (StataCorp LP, USA), and R v. 3.0.2 (R Foundation for Statistical Computing, Austria) with poLCA and OptimalCutpoints packages.
Ethics statement
The study followed the ethical principles of the Declaration of Helsinki, recommendations of the French Speaking Epidemiologists Association (ADELF), the International Conference on Harmonization, and the Council for International Organizations of Medical Sciences. The surveillance protocol was approved by National Ethical Committee of Togo and a French ethical review committee. All patients enrolled in surveillance (or their guardians) provided written informed consent.
RESULTS
Of the 1684 patients with CAP enrolled at one of the study sites from May 2010 to October 2013, 1501 (89·1%) had all tests done and were included in the analysis (Table 1). Of these, 596 (39·7%) had a CXR with PEP, 66 (4·4%) had a blood culture positive for Spn, 113 (7·5%) were positive by lytA rt-PCR, and 886 (59·0%) had CRP ⩾40 mg/l. Thirty-two Spn were identified by both blood culture and lytA rt-PCR, 81 by PCR only, and 34 by blood culture only. Nasopharyngeal aspirate testing was performed on 838 patients.
Table 1. Characteristics of patients hospitalized for community-acquired pneumonia included in the analysis in Tône and Cinkassé districts, northern Togo, 2010–2013

HIV, Human immunodefficieny virus; PEP, primary endpoint pneumonia; CXR, chest radiography; Spn, Streptococcus pneumoniae; PCR, polymerase chain reaction; CRP, C-reactive protein.
* Recent rhinopharyngitis is defined by a rhinopharyngitis episode in the last 2 weeks.
† Recent influenza is defined by an influenza episode in the last 2 weeks.
P values <0·05 are shown in bold.
The four final LCA models produced highly consistent results, with kappas for categorizing cases as likely Spn or likely non-Spn CAP from 0·928 to 0·999, so only the model including CXR and the three laboratory tests (with dichotomized CRP) is presented below. The characteristics of the two categories defined by the LCA model pointed to one of them as likely pneumococcal CAP (Table 2). In this category 99·2% of the patients had PEP on CXR, 12·8% had a positive blood culture, 16·1% had a positive lytA rt-PCR and 99·6% had CRP>40 mg/l compared to 8·6%, 0%, 3·0% and 37·8%, respectively in the other category (P < 0·001 for each comparison). In the 515 likely pneumococcal CAP, four patients had no PEP on CXR but had a positive blood culture or lytA rt-PCR and CRP >40 mg/l. Of the 986 likely non-pneumococcal CAP, 85 had PEP on CXR, but 81 of these had negative blood culture and lytA rt-PCR and CRP <40 mg/l. Based on the model, Spn was a common aetiology of CAP, causing an estimated 515 cases (34·3%, 95% CI 32·1–37·0), and was more frequent with increasing age (P < 0·001) and in male patients (P = 0·004), particularly in the 20–40 years age group (Fig. 1). Cases identified as likely pneumococcal CAP were more often severe than likely non-pneumococcal cases with a higher frequency of lethargy, hypoxia, clinical signs of sepsis, and acute complications, and a higher case-fatality ratio (4·5% vs. 0·7%, P < 0·001). Most of these differences were also observed in children aged <5 years.

Fig. 1. Estimated proportion of community-acquired pneumonia attributable to Streptococcus pneumoniae (Spn) by age and by sex in Tône and Cinkassé districts, northern Togo, 2010–2013.
Table 2. Characteristics of patients with community-acquired pneumonia attributable to Streptococcus pneumoniae and attributable to another cause based on the latent classes in Tône and Cinkassé districts, northern Togo, 2010–2013

HIV, Human immunodefficieny virus; PEP, primary endpoint pneumonia; CXR, chest radiography; Spn, Streptococcus pneumoniae; PCR, polymerase chain reaction; CRP, C-reactive protein.
* Recent rhinopharyngitis is defined by a rhinopharyngitis episode in the last 2 weeks.
† Recent influenza is defined by an influenza episode in the last 2 weeks.
P values <0·05 are shown in bold.
Table 3 presents the sensitivity and specificity of PEP, blood culture, and lytA rt-PCR estimated by the LCA model. Based on these data, PEP had the highest sensitivity yet remained reasonably specific for detecting likely Spn aetiology. lytA rt-PCR had higher sensitivity than SABC (P = 0·006) and the combination of lytA rt-PCR plus SABC had higher sensitivity than lytA rt-PCR alone (P = 0·027). The serum CRP cut-off optimizing both sensitivity and specificity based on the analysis of the area under the ROC curve was 71·2 mg/l (95% CI 67·6–75·1). Comparison of the diagnostic values of each test after estimates done separately in children aged <5 years vs. older patients found that the sensitivity and specificity of PEP and lytA rt-PCR did not differ significantly across the two age groups, whereas blood culture was less sensitive in young children (4·4% vs. 13·6%, P = 0·031) but similarly specific (100% for both groups).
Table 3. Diagnostic values of blood culture, lytA rt-PCR, CRP level, and chest radiography to identify likely pneumococcal pneumonia as categorized by the latent class analysis
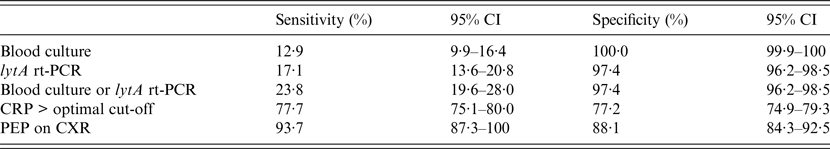
CI, Confidence interval; PCR, polymerase chain reaction; CRP, C-reactive protein; PEP, primary endpoint pneumonia; CXR, chest radiography.
Of the 838 patients with a nasopharyngeal aspirate done, 230 (27·5%) had Spn carriage identified. Pneumococcal nasopharyngeal carriage was more frequent in children age <5 years compared to older persons (50·5% vs. 24·3%, P < 0·001), in patients with PEP compared to no PEP (38·9% vs. 20·6%, P < 0·001), with a positive compared to a negative blood culture for Spn (71·0% vs. 25·8%, P < 0·001), with a positive compared to a negative lytA rt-PCR (47·6% vs. 26·4%, P = 0·004), and with CRP ⩾40 mg/l compared to lower CRP levels (35·6% vs. 19·2%, P < 0·001). None of the LCA models including nasopharyngeal aspirate results had good statistical fit. Using the LCA-defined categories, Spn nasopharyngeal carriage was more frequent in likely pneumococcal CAP (255 patients) than non-pneumococcal CAP (583 patients) (42·4% vs. 20·9%, P < 0·001) and had a positive predictive value (PPV) of 47% (108/230) and a negative predictive value (NPV) of 76% (461/608).
DISCUSSION
In this study, we used an innovative mathematical approach to investigate the diagnostic value of radiological and laboratory findings to identify Spn as the causal agent for CAP, and then estimate its contribution to CAP burden in the population of northern Togo. We calculated that in all age groups and in the context of Hib conjugate and whole-cell pertussis vaccine use in national infant immunization schedules, 34·3% of hospitalized CAP cases were likely attributable to pneumococcus. This proportion was higher in men than women and increased with age. Cases categorized as likely pneumococcal CAP were more severe, were associated with more complications, and had a higher case-fatality ratio than cases categorized as non-pneumococcal CAP, consistent with what has been described elsewhere [Reference Ishiguro30, Reference Bordon31]. PEP on CXR was the most sensitive test for identifying likely Spn cases. Specificity of PEP was lower than that of microbiological tools, but was still very high at nearly 90%. LytA rt-PCR on whole blood was superior to blood culture in terms of sensitivity and both were highly specific.
Our estimate of the proportion of CAP likely attributable to Spn was consistent with the literature. Spn was estimated to cause 25% of CAP in a meta-analysis of studies of European adults [Reference Said5]. In Kenyan adults, 46% of CAP was attributable to Spn in a prospective study that also used LCA [Reference Jokinen and Scott32], compared to 38·0% in patients aged ⩾18 years in our study. The increasing prevalence of likely Spn with age is also compatible with the fractions attributable to Spn observed in the literature including 8% in children [Reference O'Brien7], 25% in European adults [Reference Said5], and close to 50% in the elderly [Reference Zalacain33].
Our estimate of the diagnostic value of PEP did not differ significantly between children aged <5 years and patients aged ⩾5 years, despite WHO CXR interpretation guidelines having been developed and validated mainly for children aged <2 years [Reference Deloria Knoll25]. For comparative purposes, we calculated the proportion of Spn in CXR-confirmed CAP cases (equivalent to the positive PPV of PEP on CXR) in young children in a trial of 9-valent PCV from The Gambia [Reference Cutts22]. This was done by dividing the vaccine efficacy (VE) of PCV against PEP by the VE against all invasive pneumococcal disease, yielding a PPV estimate of 74%. The NPV of PEP was 90% based on a similar calculation. These estimates are consistent with our findings of a PPV and NPV of 85·7% and 99·6%, respectively, and support the importance of PEP on CXR for identifying likely Spn pneumonia cases in all age groups, although further data are needed to strengthen the methodology for older children and adults. It is important to note that these measures of diagnostic value refer to the WHO standard methodology of CXR interpretation, and are therefore unlikely to be applicable to routine clinical care in a resource-poor setting.
lytA rt-PCR on whole blood was superior to blood culture in terms of sensitivity. Previous studies in mice [Reference Rouphael17, Reference Ng18] and humans [Reference Resti34, Reference Werno, Anderson and Murdoch35] have shown PCR on whole blood to have a higher sensitivity, although comparing earlier results to our findings is difficult because of different study designs, analytical approaches, blood fraction used for rt-PCR, and small sample sizes. The sensitivity we found for blood culture was low and consistent with other studies [Reference Afshar13]. A significantly lower sensitivity was found in children aged <5 years, whereas lytA rt-PCR sensitivity did not vary significantly between age groups. Although lytA rt-PCR was more sensitive than blood culture, using a combination of both tests significantly improved the overall sensitivity, illustrating their complementary value for aetiological confirmation of cases. lytA rt-PCR specificity was high but not 100%, which could be explained by cross-reactivity with other bacteria such as Streptococcus mitis [Reference Whatmore36].
The lower frequency of recent rhinopharyngitis in likely pneumococcal CAP and the similar frequency of recent influenza in both groups is counter-intuitive, as viral infections (particularly influenza) are known to be associated with pneumococcal pneumonia [Reference Simonsen37]. However, recent rhinopharyngitis and influenza were both self-reported and may therefore be subject to misclassification. In addition, the role of influenza as a precursor of pneumococcal pneumonia has been shown in the case of severe pandemic influenza [Reference Morens, Taubenberger and Fauci38] but is less well established in inter-pandemic periods and for mild disease. Finally, our analysis of nasopharyngeal aspirate data found no association between viral infection in general (or influenza infection in particular) and pneumococcal colonization and a negative association between viral infection and pneumococcal bacteraemia, suggesting that these interactions are not straightforward [Reference Moïsi39].
Pneumococcal nasopharyngeal carriage could not be included in the LCA models. The subgroup analysis revealed that colonization has poor predictive value for likely Spn CAP. This is consistent with the observation that Spn carriage does not necessarily predict Spn CAP [Reference Weinberger40].
Our study had several limitations. The final LCA models included only four tests and two of them were not independent. Sensitivity analysis revealed that the LCA models relied heavily on PEP for categorization of patients; the other tests identified only a small number of additional likely Spn cases in patients who were PEP-negative. Pneumococcal nasopharyngeal colonization findings were not included in the LCA models because they led to poor fit. However, the addition of other diagnostic tools such as urine antigen detection tests for pneumococcal capsular polysaccharide, which were not done in our study, might have further strengthened our analysis.
The LCA approach offers the opportunity to limit the bias inherent to the use of imperfect tests in diagnostic evaluations. Together, the large and representative sample of cases, the similarity of findings in the four final LCA models, and the consistency with the literature all support the validity of our results. We estimated that Spn may cause about one third of CAP cases in our population and plays an important role in both childhood and adult disease. We provide data that PEP on CXR may have a role in monitoring PCV impact in older populations. We have continued to monitor hospitalized pneumonia trends in this meningitis belt population after the 2014 introduction of PCV13 into the routine immunization programme and plan to use LCA to measure the proportion of CAP likely due to Spn in the post-vaccine introduction period and improve estimations of vaccine impact.
ACKNOWLEDGEMENTS
This work was supported by an Investigator-Initiated Research grant from Pfizer (IIR WS951939).
DECLARATION OF INTEREST
None.







